From Wikipedia, the free encyclopedia
https://en.wikipedia.org/wiki/Enzyme_inhibitor
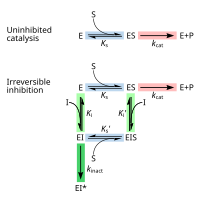
Top:
enzyme (E) accelerates conversion of
substrates (S) to
products
(P). Bottom: by binding to enzyme, inhibitor (I) blocks binding of
substrate. Binding site shown in blue checkerboard, substrate as black
rectangle, and inhibitor as green rounded rectangle.
An enzyme inhibitor is a molecule that binds to an enzyme and blocks its activity. Enzymes are proteins that speed up chemical reactions necessary for life, in which substrate molecules are converted into products. An enzyme facilitates a specific chemical reaction by binding the substrate to its active site, a specialized area on the enzyme that accelerates the most difficult step of the reaction.
An enzyme inhibitor hinders ("inhibits") this process, either by
binding to the enzyme's active site (thus preventing the substrate
itself from binding) or by binding to another site on the enzyme such
that the enzyme's catalysis of the reaction is blocked. Enzyme inhibitors may bind reversibly or irreversibly. Irreversible inhibitors form a chemical bond with the enzyme such that the enzyme is inhibited until the chemical bond is broken. By contrast, reversible inhibitors bind non-covalently
and may spontaneously leave the enzyme, allowing the enzyme to resume
its function. Reversible inhibitors produce different types of
inhibition depending on whether they bind to the enzyme, the
enzyme-substrate complex, or both.
Enzyme inhibitors play an important role in all cells, since they
are generally specific to one enzyme and serve to control that enzyme's
activity. For example, enzymes in a metabolic pathway
may be inhibited by molecules produced later in the pathway, thus
curtailing the production of molecules that are no longer needed. This type of negative feedback is an important way to maintain balance in a cell. Enzyme inhibitors also control essential enzymes such as proteases or nucleases that, if left unchecked, may damage a cell. Many poisons produced by animals or plants are enzyme inhibitors that block the activity of crucial enzymes in prey or predators.
Many drug molecules are enzyme inhibitors that inhibit an aberrant human enzyme or an enzyme critical for the survival of a pathogen such as a virus, bacterium or parasite. Examples include methotrexate (used in chemotherapy and in treating rheumatic arthritis) and the protease inhibitors used to treat HIV/AIDS. Since anti-pathogen inhibitors generally target only one enzyme, such drugs are highly specific and generally produce few side effects in humans, provided that no analogous enzyme is found in humans. (This is often the case, since such pathogens and humans are genetically distant.) Medicinal enzyme inhibitors often have low dissociation constants,
meaning that only a minute amount of the inhibitor is required to
inhibit the enzyme. A low concentration of the enzyme inhibitor reduces
the risk for liver and kidney damage and other adverse drug reactions in humans. Hence the discovery and refinement of enzyme inhibitors is an active area of research in biochemistry and pharmacology.
Structural classes
Enzyme inhibitors are a chemically diverse set of substances that range in size from organic small molecules to macromolecular proteins.
Small molecule inhibitors include essential primary metabolites
that inhibit upstream enzymes that produce those metabolites. This
provides a negative feedback loop that prevents over production of
metabolites and thus maintains cellular homeostasis (steady internal conditions). Small molecule enzyme inhibitors also include secondary metabolites,
which are not essential to the organism that produces them, but provide
the organism with a evolutionary advantage, in that they can be used to
repel predators or competing organisms or immobilize prey. In addition, many drugs are small molecule enzyme inhibitors that target either disease-modifying enzymes in the patient or enzymes in pathogens which are required for the growth and reproduction of the pathogen.
In addition to small molecules, some proteins act as enzyme inhibitors. The most prominent example are serpins (serine protease inhibitors) which are produced by animals to protect against inappropriate enzyme activation and by plants to prevent predation. Another class of inhibitor proteins is the ribonuclease inhibitors, which bind to ribonucleases in one of the tightest known protein–protein interactions. A special case of protein enzyme inhibitors are zymogens that contain an autoinhibitory N-terminal peptide that binds to the active site of enzyme that intramolecularly
blocks its activity as a protective mechanism against uncontrolled
catalysis. The N-terminal peptide is cleaved (split) from the zymogen
enzyme precursor by another enzyme to release an active enzyme.
The binding site of inhibitors on enzymes is most commonly the same site that binds the substrate of the enzyme. These active site inhibitors are known as orthosteric ("regular" orientation) inhibitors.
The mechanism of orthosteric inhibition is simply to prevent substrate
binding to the enzyme through direct competition which in turn prevents
the enzyme from catalysing the conversion of substrates into products.
Alternatively, the inhibitor can bind to a site remote from the enzyme
active site. These are known as allosteric ("alternative" orientation) inhibitors. The mechanism of allosteric inhibition are varied and include changing the conformation (shape) of the enzyme such that it can no longer bind substrate (kinetically indistinguishable from competitive orthosteric inhibition)
or alternatively stabilise binding of substrate to the enzyme but lock
the enzyme in a conformation which is no longer catalytically active.
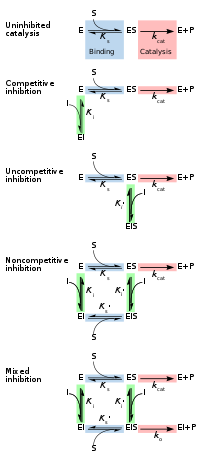
Reversible inhibitors
Kinetic
mechanisms for reversible inhibition. Substrate (S) binding to enzyme
(E) in blue, catalysis releasing product (P) in red, inhibitor (I)
binding to enzyme in green.
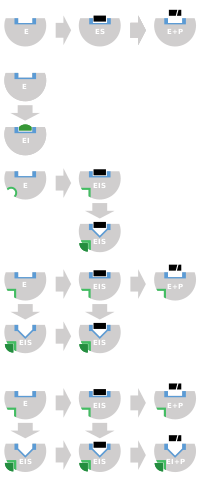
Schematics
for reversible inhibition. Binding site in blue, substrate in black,
inhibitor in green, and allosteric site in light green.
Competitive
inhibitors usually bind to the active site. Non-competitive bind to a
remote (allosteric) site. Uncompetitive inhibitors only bind once the
substrate is bound, fully disrupting catalysis, and mixed inhibition is
similar but with only partial disruption of catalysis.
Reversible inhibitors attach to enzymes with non-covalent interactions such as hydrogen bonds, hydrophobic interactions and ionic bonds. Multiple weak bonds between the inhibitor and the enzyme active site combine to produce strong and specific binding.
In contrast to irreversible inhibitors, reversible inhibitors
generally do not undergo chemical reactions when bound to the enzyme and
can be easily removed by dilution or dialysis. A special case are covalent reversible inhibitors that form a chemical bond with the enzyme, but the bond can be cleaved so the inhibition is fully reversible.
Reversible inhibitors are generally categorized into four types, as introduced by Cleland in 1963. They are classified according to the effect of the inhibitor on the Vmax (maximum reaction rate catalysed by the enzyme) and Km
(the concentration of substrate resulting in half maximal enzyme
activity) as the concentration of the enzyme's substrate is varied.
Competitive
In competitive inhibition, the substrate and inhibitor cannot bind to the enzyme at the same time. This usually results from the inhibitor having an affinity for the
active site of an enzyme where the substrate also binds; the substrate
and inhibitor compete for access to the enzyme's active site.
This type of inhibition can be overcome by sufficiently high
concentrations of substrate (Vmax remains constant), i.e., by out-competing the inhibitor. However, the apparent Km will increase as it takes a higher concentration of the substrate to reach the Km point, or half the Vmax.
Competitive inhibitors are often similar in structure to the real
substrate (see for example the "methotrexate versus folate" figure in
the "Drugs" section).
Uncompetitive
In uncompetitive inhibition, the inhibitor binds only to the enzyme-substrate complex. This type of inhibition causes Vmax to decrease (maximum velocity decreases as a result of removing activated complex) and Km to decrease (due to better binding efficiency as a result of Le Chatelier's principle and the effective elimination of the ES complex thus decreasing the Km which indicates a higher binding affinity). Uncompetitive inhibition is rare.
Non-competitive
In non-competitive inhibition, the binding of the inhibitor to the enzyme reduces its activity but does not affect the binding of substrate. As a result, the extent of inhibition depends only on the concentration of the inhibitor. Vmax will decrease due to the inability for the reaction to proceed as efficiently, but Km will remain the same as the actual binding of the substrate, by definition, will still function properly.
Mixed
In mixed inhibition,
the inhibitor may bind to the enzyme whether or not the substrate has
already bound. Hence mixed inhibition is a combination of competitive
and noncompetitive inhibition. Furthermore, the affinity of the inhibitor for the free enzyme and the enzyme-substrate complex may differ.
By increasing concentrations of substrate [S], this type of inhibition
can be reduced (due to the competitive contribution), but not entirely
overcome (due to the noncompetitive component).
Although it is possible for mixed-type inhibitors to bind in the active
site, this type of inhibition generally results from an allosteric effect where the inhibitor binds to a different site on an enzyme. Inhibitor binding to this allosteric site changes the conformation (that is, the tertiary structure or three-dimensional shape) of the enzyme so that the affinity of the substrate for the active site is reduced.
These four types of inhibition can also be distinguished by the
effect of increasing the substrate concentration [S] on the degree of
inhibition caused by a given amount of inhibitor. For competitive
inhibition the degree of inhibition is reduced by increasing [S], for
noncompetitive inhibition the degree of inhibition is unchanged, and for
uncompetitive (also called anticompetitive) inhibition the degree of
inhibition increases with [S].
Quantitative description
Reversible inhibition can be described quantitatively in terms of the inhibitor's binding to the enzyme and to the enzyme-substrate complex, and its effects on the kinetic constants of the enzyme. In the classic Michaelis-Menten
scheme (shown in the "inhibition mechanism schematic" diagram), an
enzyme (E) binds to its substrate (S) to form the enzyme–substrate
complex ES. Upon catalysis, this complex breaks down to release product P
and free enzyme. The inhibitor (I) can bind to either E or ES with the dissociation constants Ki or Ki', respectively.
- Competitive inhibitors can bind to E, but not to ES. Competitive inhibition increases Km (i.e., the inhibitor interferes with substrate binding), but does not affect Vmax (the inhibitor does not hamper catalysis in ES because it cannot bind to ES).
- Uncompetitive inhibitors bind to ES. Uncompetitive inhibition decreases both Km and Vmax. The inhibitor affects substrate binding by increasing the enzyme's affinity for the substrate (decreasing Km) as well as hampering catalysis (decreases Vmax).
- Non-competitive inhibitors have identical affinities for E and ES (Ki = Ki'). Non-competitive inhibition does not change Km (i.e., it does not affect substrate binding) but decreases Vmax (i.e., inhibitor binding hampers catalysis).
- Mixed-type inhibitors bind to both E and ES, but their affinities for these two forms of the enzyme are different (Ki ≠ Ki'). Thus, mixed-type inhibitors affect substrate binding (increase or decrease Km) and hamper catalysis in the ES complex (decrease Vmax).
When an enzyme has multiple substrates, inhibitors can show different
types of inhibition depending on which substrate is considered. This
results from the active site containing two different binding sites
within the active site, one for each substrate. For example, an
inhibitor might compete with substrate A for the first binding site, but
be a non-competitive inhibitor with respect to substrate B in the
second binding site.
Traditionally reversible enzyme inhibitors have been classified
as competitive, uncompetitive, or non-competitive, according to their
effects on Km and Vmax.
These three types of inhibition result respectively from the inhibitor
binding only to the enzyme E in the absence of substrate S, to the
enzyme–substrate complex ES, or to both. The division of these classes
arises from a problem in their derivation and results in the need to use
two different binding constants for one binding event.
It is further assumed that binding of the inhibitor to the enzyme
results in 100% inhibition and fails to consider the possibility of
partial inhibition.
The common form of the inhibitory term also obscures the relationship
between the inhibitor binding to the enzyme and its relationship to any
other binding term be it the Michaelis–Menten equation or a dose
response curve associated with ligand receptor binding. To demonstrate
the relationship the following rearrangement can be made:
![{\displaystyle {\begin{aligned}{\cfrac {V_{\max }}{1+{\cfrac {\ce {[I]}}{K_{i}}}}}&={V_{\max }}\left({\cfrac {K_{i}}{K_{i}+[{\ce {I}}]}}\right)&&{\text{multiply by }}{\cfrac {K_{i}}{K_{i}}}=1\\&={V_{\max }}\left({\cfrac {K_{i}+[{\ce {I}}]-[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{add }}[{\ce {I}}]-[{\ce {I}}]=0{\text{ to numerator}}\\&={V_{\max }}\left(1-{\cfrac {[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{simplify }}{\cfrac {K_{i}+[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}=1\\&=V_{\max }-V_{\max }{\cfrac {\ce {[I]}}{K_{i}+[{\ce {I}}]}}&&{\text{multiply out by }}V_{\max }\end{aligned}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/37eda4dec307f8acfca89b2d8f4811474ea764ec)
This rearrangement demonstrates that similar to the Michaelis–Menten
equation, the maximal rate of reaction depends on the proportion of the
enzyme population interacting with its substrate.
fraction of the enzyme population bound by substrate
![{\displaystyle {\cfrac {\ce {[S]}}{[{\ce {S}}]+K_{m}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/eb08dd139085a394e6e7370f47ebfa255f1ad685)
fraction of the enzyme population bound by inhibitor
![{\displaystyle {\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/9ed50a1f7a5f2c52f406b52263916ab48b268e07)
the effect of the inhibitor is a result of the percent of the enzyme
population interacting with inhibitor. The only problem with this
equation in its present form is that it assumes absolute inhibition of
the enzyme with inhibitor binding, when in fact there can be a wide
range of effects anywhere from 100% inhibition of substrate turn over to
no inhibition. To account for this the equation can be easily modified
to allow for different degrees of inhibition by including a delta Vmax term.
![{\displaystyle V_{\max }-\Delta V_{\max }{\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/7dff424ec79284c3a1cea14f0f82b0eaace53c69)
or
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/f3874623edd9524ba2741fe448927bf5cf0ab257)
This term can then define the residual enzymatic activity present
when the inhibitor is interacting with individual enzymes in the
population. However the inclusion of this term has the added value of
allowing for the possibility of activation if the secondary Vmax
term turns out to be higher than the initial term. To account for the
possibly of activation as well the notation can then be rewritten
replacing the inhibitor "I" with a modifier term (stimulator or
inhibitor) denoted here as "X".
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/306d44733a89308883053e3b8372a8cf9ce0239b)
While this terminology results in a simplified way of dealing with
kinetic effects relating to the maximum velocity of the Michaelis–Menten
equation, it highlights potential problems with the term used to
describe effects relating to the Km. The Km
relating to the affinity of the enzyme for the substrate should in most
cases relate to potential changes in the binding site of the enzyme
which would directly result from enzyme inhibitor interactions. As such
a term similar to the delta Vmax term proposed above to modulate Vmax should be appropriate in most situations:
![{\displaystyle K_{m1}-(K_{m1}-K_{m2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/cb4e0de216e1e625bb803ee725bf85c9989a15f5)
Dissociation constants
Lineweaver–Burk
diagrams of different types of reversible enzyme inhibitors. The arrow
shows the effect of increasing concentrations of inhibitor.
An enzyme inhibitor is characterised by its dissociation constant Ki,
the concentration at which the inhibitor half occupies the enzyme. In
non-competitive inhibition, the inhibitor can also bind to the
enzyme-substrate complex, and the presence of bound substrate can change
the affinity of the inhibitor for the enzyme, resulting in a second
dissociation constant Ki'. Hence Ki and Ki' are the dissociation constants of the inhibitor for the enzyme and to the enzyme-substrate complex, respectively. The enzyme-inhibitor constant Ki can be measured directly by various methods; one especially accurate method is isothermal titration calorimetry, in which the inhibitor is titrated into a solution of enzyme and the heat released or absorbed is measured. However, the other dissociation constant Ki'
is difficult to measure directly, since the enzyme-substrate complex is
short-lived and undergoing a chemical reaction to form the product.
Hence, Ki' is usually measured indirectly, by observing the enzyme activity under various substrate and inhibitor concentrations, and fitting the data via nonlinear regression to a modified Michaelis–Menten equation.
![V={\frac {V_{max}[S]}{\alpha K_{m}+\alpha ^{\prime }[S]}}={\frac {(1/\alpha ^{\prime })V_{max}[S]}{(\alpha /\alpha ^{\prime })K_{m}+[S]}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4a8f0a9dda1d308de7f090f99c2833f944f11a09)
where the modifying factors α and α' are defined by the inhibitor concentration and its two dissociation constants
![\alpha =1+{\frac {[I]}{K_{i}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/57fcf54938a9784f9313437681b220079ff43ee5)
![\alpha ^{\prime }=1+{\frac {[I]}{K_{i}^{\prime }}}.](https://wikimedia.org/api/rest_v1/media/math/render/svg/65bf16742482cae7b0743781f47c327ddcf537e3)
Thus, in the presence of the inhibitor, the enzyme's effective Km and Vmax become (α/α')Km and (1/α')Vmax,
respectively. However, the modified Michaelis-Menten equation assumes
that binding of the inhibitor to the enzyme has reached equilibrium,
which may be a very slow process for inhibitors with sub-nanomolar
dissociation constants. In these cases, the inhibition becomes
effectively irreversible, hence it is more practical to treat such
tight-binding inhibitors as irreversible (see below).
The effects of different types of reversible enzyme inhibitors on
enzymatic activity can be visualised using graphical representations of
the Michaelis–Menten equation, such as Lineweaver–Burk, Eadie-Hofstee or Hanes-Woolf plots. An illustration is provided by the three Lineweaver–Burk plots depicted in the Lineweaver–Burk diagrams figure. In the top diagram, the competitive inhibition lines intersect on the y-axis, illustrating that such inhibitors do not affect Vmax. In the bottom diagram, the non-competitive inhibition lines intersect on the x-axis, showing these inhibitors do not affect Km. However, since it can be difficult to estimate Ki and Ki' accurately from such plots, it is advisable to estimate these constants using more reliable nonlinear regression methods.
Special cases
Partially competitive
The
mechanism of partially competitive inhibition is similar to that of
non-competitive, except that the EIS complex has catalytic activity,
which may be lower or even higher (partially competitive activation)
than that of the enzyme–substrate (ES) complex. This inhibition
typically displays a lower Vmax, but an unaffected Km value.
Substrate or product
Substrate
or product inhibition is where either an enzymes substrate or product
also act as an inhibitor. This inhibition may follow the competitive,
uncompetitive or mixed patterns. In substrate inhibition there is a
progressive decrease in activity at high substrate concentrations,
potentially from an enzyme having two competing substrate-binding sites. At low substrate, the high-affinity site is occupied and normal kinetics are followed. However, at higher concentrations, the second inhibitory site becomes occupied, inhibiting the enzyme. Product inhibition (either the enzyme's own product, or a product to an enzyme downstream in its metabolic pathway) is often a regulatory feature in metabolism and can be a form of negative feedback.
Slow-tight
Slow-tight inhibition occurs when the initial enzyme–inhibitor complex EI undergoes conformational isomerism
(a change in shape) to a second more tightly held complex, EI*, but the
overall inhibition process is reversible. This manifests itself as
slowly increasing enzyme inhibition. Under these conditions, traditional
Michaelis–Menten kinetics give a false value for Ki, which is time–dependent. The true value of Ki can be obtained through more complex analysis of the on (kon) and off (koff) rate constants for inhibitor association with kinetics similar to irreversible inhibition.
Multi-substrate analogues
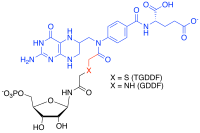
TGDDF/GDDF
multi-substrate adduct inhibitor. Substrate analogue in black, cofactor
analogue in blue, non-cleavable linker in red.
Peptide-based HIV-1 protease inhibitor
ritonavir with substrate binding sites located in enzyme labelled as S2, S1, S1', and S2'.
Multi-substrate analogue inhibitors are high affinity selective
inhibitors that can be prepared for enzymes that catalyse reactions with
more than one substrate by capturing the binding energy of each of
those substrate into one molecule. For example, in the formyl transfer reactions of purine biosynthesis, a potent Multi-substrate Adduct Inhibitor (MAI) to glycinamide ribonucleotide (GAR) TFase was prepared synthetically by linking analogues of the GAR substrate and the N-10-formyl tetrahydrofolate cofactor together to produce thioglycinamide ribonucleotide dideazafolate (TGDDF), or enzymatically from the natural GAR substrate to yield GDDF. Here the subnanomolar dissociation constant (KD) of TGDDF was greater than predicted presumably due to entropic
advantages gained and/or positive interactions acquired through the
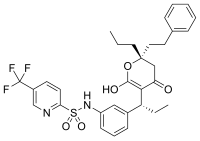
atoms linking the components. MAIs have also been observed to be
produced in cells by reactions of pro-drugs such as isoniazid or enzyme inhibitor ligands (for example, PTC124) with cellular cofactors such as nicotinamide adenine dinucleotide (NADH) and adenosine triphosphate (ATP) respectively.
Examples
As
enzymes have evolved to bind their substrates tightly, and most
reversible inhibitors bind in the active site of enzymes, it is
unsurprising that some of these inhibitors are strikingly similar in
structure to the substrates of their targets. Inhibitors of dihydrofolate reductase (DHFR) are prominent examples.[46] Other examples of these substrate mimics are the protease inhibitors, a therapeutically effective class of antiretroviral drugs used to treat HIV/AIDS. The structure of ritonavir, a peptidomimetic (peptide mimic) protease inhibitor containing three peptide bonds,
as shown in the "competitive inhibitor examples" figure. As this drug
resembles the peptide that is the substrate of the HIV protease, it
competes with the substrate in the enzyme's active site.
Enzyme inhibitors are often designed to mimic the transition state or intermediate of an enzyme-catalysed reaction.
This ensures that the inhibitor exploits the transition state
stabilising effect of the enzyme, resulting in a better binding affinity
(lower Ki) than substrate-based designs. An example of such a transition state inhibitor is the antiviral drug oseltamivir; this drug mimics the planar nature of the ring oxonium ion in the reaction of the viral enzyme neuraminidase.
However, not all inhibitors are based on the structures of
substrates. For example, the structure of another HIV protease inhibitor
tipranavir
is not based on a peptide and has no obvious structural similarity to a
protein substrate. These non-peptide inhibitors can be more stable than
inhibitors containing peptide bonds, because they will not be
substrates for peptidases and are less likely to be degraded.
In drug design it is important to consider the concentrations of
substrates to which the target enzymes are exposed. For example, some protein kinase inhibitors have chemical structures that are similar to ATP, one of the substrates of these enzymes.
However, drugs that are simple competitive inhibitors will have to
compete with the high concentrations of ATP in the cell. Protein kinases
can also be inhibited by competition at the binding sites where the
kinases interact with their substrate proteins, and most proteins are
present inside cells at concentrations much lower than the concentration
of ATP. As a consequence, if two protein kinase inhibitors both bind in
the active site with similar affinity, but only one has to compete with
ATP, then the competitive inhibitor at the protein-binding site will
inhibit the enzyme more effectively.
Irreversible inhibitors
Types
Irreversible
inhibitors bind to the enzyme's binding site then undergo a chemical
reaction to form a covalent enzyme-inhibitor complex (EI*). Binding site
in blue, inhibitor in green.
Irreversible inhibitors covalently bind to an enzyme, and this type of inhibition can therefore not be readily reversed. Irreversible inhibitors often contain reactive functional groups such as nitrogen mustards, aldehydes, haloalkanes, alkenes, Michael acceptors, phenyl sulfonates, or fluorophosphonates. These electrophilic groups react with amino acid side chains to form covalent adducts. The residues modified are those with side chains containing nucleophiles such as hydroxyl or sulfhydryl groups; these include the amino acids serine (that reacts with DFP, see the "DFP reaction" diagram), and also cysteine, threonine, or tyrosine.
Irreversible inhibition is different from irreversible enzyme inactivation.
Irreversible inhibitors are generally specific for one class of enzyme
and do not inactivate all proteins; they do not function by destroying protein structure but by specifically altering the active site of their target. For example, extremes of pH or temperature usually cause denaturation
of all protein structure, but this is a non-specific effect. Similarly,
some non-specific chemical treatments destroy protein structure: for
example, heating in concentrated hydrochloric acid will hydrolyse the peptide bonds holding proteins together, releasing free amino acids.
Irreversible inhibitors display time-dependent inhibition and their potency therefore cannot be characterised by an IC50 value.
This is because the amount of active enzyme at a given concentration of
irreversible inhibitor will be different depending on how long the
inhibitor is pre-incubated with the enzyme. Instead, kobs/[I] values are used, where kobs is the observed pseudo-first order rate of inactivation (obtained by plotting the log of % activity versus time) and [I] is the concentration of inhibitor. The kobs/[I] parameter is valid as long as the inhibitor does not saturate binding with the enzyme (in which case kobs = kinact) where kinact is the rate of inactivation.
Measuring
Kinetic
mechanism for irreversible inhibition. Substrate binding in blue,
catalysis in red, inhibitor binding in green, inactivation reaction in
dark green.
Irreversible inhibitors first form a reversible non-covalent complex
with the enzyme (EI or ESI). Subsequently a chemical reaction occurs
between the enzyme and inhibitor to produce the covalently modified
"dead-end complex" EI* (an irreversible covalent complex). The rate at
which EI* is formed is called the inactivation rate or kinact.
Since formation of EI may compete with ES, binding of irreversible
inhibitors can be prevented by competition either with substrate or with
a second, reversible inhibitor. This protection effect is good evidence
of a specific reaction of the irreversible inhibitor with the active
site.
The binding and inactivation steps of this reaction are
investigated by incubating the enzyme with inhibitor and assaying the
amount of activity remaining over time. The activity will be decreased
in a time-dependent manner, usually following exponential decay. Fitting these data to a rate equation
gives the rate of inactivation at this concentration of inhibitor. This
is done at several different concentrations of inhibitor. If a
reversible EI complex is involved the inactivation rate will be
saturable and fitting this curve will give kinact and Ki.
Another method that is widely used in these analyses is mass spectrometry.
Here, accurate measurement of the mass of the unmodified native enzyme
and the inactivated enzyme gives the increase in mass caused by reaction
with the inhibitor and shows the stoichiometry of the reaction. This is usually done using a MALDI-TOF mass spectrometer. In a complementary technique, peptide mass fingerprinting involves digestion of the native and modified protein with a protease such as trypsin. This will produce a set of peptides
that can be analysed using a mass spectrometer. The peptide that
changes in mass after reaction with the inhibitor will be the one that
contains the site of modification.
Slow binding
Chemical
mechanism for irreversible inhibition of ornithine decarboxylase by
DFMO. Pyridoxal 5'-phosphate (Py) and enzyme (E) are not shown. Adapted
from Poulin et al, 1992.
Not all irreversible inhibitors form covalent adducts with their
enzyme targets. Some reversible inhibitors bind so tightly to their
target enzyme that they are essentially irreversible. These
tight-binding inhibitors may show kinetics similar to covalent
irreversible inhibitors. In these cases, some of these inhibitors
rapidly bind to the enzyme in a low-affinity EI complex and this then
undergoes a slower rearrangement to a very tightly bound EI* complex
(see the "irreversible inhibition mechanism" diagram). This kinetic
behaviour is called slow-binding. This slow rearrangement after binding often involves a conformational change
as the enzyme "clamps down" around the inhibitor molecule. Examples of
slow-binding inhibitors include some important drugs, such methotrexate, allopurinol, and the activated form of acyclovir.
Some examples
Trypanothione reductase with the lower molecule of an inhibitor bound irreversibly and the upper one reversibly. Created from Bond,
et al, 2004. (
PDB: 1GXF)
Diisopropylfluorophosphate (DFP) is an example of an irreversible protease inhibitor
(see the "DFP reaction" diagram). The enzyme hydrolyses the
phosphorus–fluorine bond, but the phosphate residue remains bound to the
serine in the active site, deactivating it. Similarly, DFP also reacts with the active site of acetylcholine esterase in the synapses of neurons, and consequently is a potent neurotoxin, with a lethal dose of less than 100 mg.
Suicide inhibition is an unusual type of irreversible inhibition where the enzyme converts the inhibitor into a reactive form in its active site. An example is the inhibitor of polyamine biosynthesis, α-difluoromethylornithine (DFMO), which is an analogue of the amino acid ornithine, and is used to treat African trypanosomiasis (sleeping sickness). Ornithine decarboxylase
can catalyse the decarboxylation of DFMO instead of ornithine (see the
"DFMO inhibitor mechanism" diagram). However, this decarboxylation
reaction is followed by the elimination of a fluorine atom, which
converts this catalytic intermediate into a conjugated imine,
a highly electrophilic species. This reactive form of DFMO then reacts
with either a cysteine or lysine residue in the active site to
irreversibly inactivate the enzyme.
Since irreversible inhibition often involves the initial formation of a non-covalent enzyme inhibitor (EI) complex, it is sometimes possible for an inhibitor to bind to an enzyme in more than one way. For example, in the figure showing trypanothione reductase from the human protozoan parasite Trypanosoma cruzi, two molecules of an inhibitor called quinacrine mustard
are bound in its active site. The top molecule is bound reversibly, but
the lower one is bound covalently as it has reacted with an amino acid
residue through its nitrogen mustard group.
Applications
Enzyme inhibitors are found in nature and also produced artificially in the laboratory. Naturally occurring enzyme inhibitors regulate many metabolic processes and are essential for life. In addition, naturally produced poisons are often enzyme inhibitors that have evolved for use as toxic agents against predators, prey, and competing organisms. These natural toxins include some of the most poisonous substances known. Artificial inhibitors are often used as drugs, but can also be insecticides such as malathion, herbicides such as glyphosate, or disinfectants such as triclosan. Other artificial enzyme inhibitors block acetylcholinesterase, an enzyme which breaks down acetylcholine, and are used as nerve agents in chemical warfare.
Metabolic regulation
Enzyme inhibition is a common feature of metabolic pathway control in cells. Metabolic flux through a pathway is often regulated by a pathway's metabolites acting as inhibitors and enhancers for the enzymes in that same pathway. The glycolytic pathway is a classic example. This catabolic pathway consumes glucose and produces ATP, NADH and pyruvate. A key step for the regulation of glycolysis is an early reaction in the pathway catalysed by phosphofructokinase-1
(PFK1). When ATP levels rise, ATP binds an allosteric site in PFK1 to
decrease the rate of the enzyme reaction; glycolysis is inhibited and
ATP production falls. This negative feedback
control helps maintain a steady concentration of ATP in the cell.
However, metabolic pathways are not just regulated through inhibition
since enzyme activation is equally important. With respect to PFK1, fructose 2,6-bisphosphate and ADP are examples of metabolites that are allosteric activators.
Physiological enzyme inhibition can also be produced by specific protein inhibitors. This mechanism occurs in the pancreas, which synthesises many digestive precursor enzymes known as zymogens. Many of these are activated by the trypsin
protease, so it is important to inhibit the activity of trypsin in the
pancreas to prevent the organ from digesting itself. One way in which
the activity of trypsin is controlled is the production of a specific
and potent trypsin inhibitor
protein in the pancreas. This inhibitor binds tightly to trypsin,
preventing the trypsin activity that would otherwise be detrimental to
the organ.
Although the trypsin inhibitor is a protein, it avoids being hydrolysed
as a substrate by the protease by excluding water from trypsin's active
site and destabilising the transition state. Other examples of physiological enzyme inhibitor proteins include the barstar inhibitor of the bacterial ribonuclease barnase.
Natural poisons
Animals and plants have evolved to synthesise a vast array of poisonous products including secondary metabolites, peptides and proteins that can act as inhibitors. Natural toxins are usually small organic molecules and are so diverse that there are probably natural inhibitors for most metabolic processes.
The metabolic processes targeted by natural poisons encompass more than
enzymes in metabolic pathways and can also include the inhibition of
receptor, channel and structural protein functions in a cell. For
example, paclitaxel (taxol), an organic molecule found in the Pacific yew tree, binds tightly to tubulin dimers and inhibits their assembly into microtubules in the cytoskeleton.
Many natural poisons act as neurotoxins that can cause paralysis leading to death and function for defence against predators or in hunting and capturing prey. Some of these natural inhibitors, despite their toxic attributes, are valuable for therapeutic uses at lower doses. An example of a neurotoxin are the glycoalkaloids, from the plant species in the family Solanaceae (includes potato, tomato and eggplant), that are acetylcholinesterase inhibitors.
Inhibition of this enzyme causes an uncontrolled increase in the
acetylcholine neurotransmitter, muscular paralysis and then death.
Neurotoxicity can also result from the inhibition of receptors; for
example, atropine from deadly nightshade (Atropa belladonna) that functions as a competitive antagonist of the muscarinic acetylcholine receptors.
Although many natural toxins are secondary metabolites, these
poisons also include peptides and proteins. An example of a toxic
peptide is alpha-amanitin, which is found in relatives of the death cap mushroom. This is a potent enzyme inhibitor, in this case preventing the RNA polymerase II enzyme from transcribing DNA. The algal toxin microcystin is also a peptide and is an inhibitor of protein phosphatases. This toxin can contaminate water supplies after algal blooms and is a known carcinogen that can also cause acute liver haemorrhage and death at higher doses.
Proteins can also be natural poisons or antinutrients, such as the trypsin inhibitors (discussed in the "metabolic regulation" section above) that are found in some legumes.
A less common class of toxins are toxic enzymes: these act as
irreversible inhibitors of their target enzymes and work by chemically
modifying their substrate enzymes. An example is ricin, an extremely potent protein toxin found in castor oil beans. This enzyme is a glycosidase that inactivates ribosomes. Since ricin is a catalytic irreversible inhibitor, this allows just a single molecule of ricin to kill a cell.
Drugs
The coenzyme folic acid (top) compared to the anti-cancer drug methotrexate (bottom)
The most common uses for enzyme inhibitors are as drugs to treat
disease. Many of these inhibitors target a human enzyme and aim to
correct a pathological condition. For instance, aspirin is a widely used drug that acts as a suicide inhibitor of the cyclooxygenase enzyme. This inhibition in turn suppresses the production of proinflammatory prostaglandins and thus aspirin may be used to reduce pain, fever, and inflammation.
As of 2017, an estimated 29% of approved drugs are enzyme inhibitors of which approximately one-fifth are kinase inhibitors. A notable class of kinase drug targets is the receptor tyrosine kinases which are essential enzymes that regulate cell growth; their over-activation may result in cancer. Hence kinase inhibitors such as imatinib are frequently used to treat malignancies. Janus kinases are another notable example of drug enzyme targets. Inhibitors of Janus kinases block the production of inflammatory cytokines and hence these inhibitors are used to treat a variety of inflammatory diseases in including arthritis, asthma, and Crohn's disease.
An example of the structural similarity of some inhibitors to the
substrates of the enzymes they target is seen in the figure comparing
the drug methotrexate to folic acid. Folic acid is the oxidised form of the substrate of dihydrofolate reductase,
an enzyme that is potently inhibited by methotrexate. Methotrexate
blocks the action of dihydrofolate reductase and thereby halts thymidine biosynthesis. This block of nucleotide biosynthesis is selectively toxic to rapidly growing cells, therefore methotrexate is often used in cancer chemotherapy.
A common treatment for erectile dysfunction is sildenafil (Viagra). This compound is a potent inhibitor of cGMP specific phosphodiesterase type 5, the enzyme that degrades the signalling molecule cyclic guanosine monophosphate. This signalling molecule triggers smooth muscle relaxation and allows blood flow into the corpus cavernosum,
which causes an erection. Since the drug decreases the activity of the
enzyme that halts the signal, it makes this signal last for a longer
period of time.
Antibiotics
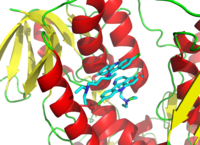
The structure of a complex between penicillin G and the
Streptomyces transpeptidase. (
PDB: 1PWC)
Drugs are also used to inhibit enzymes needed for the survival of pathogens. For example, bacteria are surrounded by a thick cell wall made of a net-like polymer called peptidoglycan. Many antibiotics such as penicillin and vancomycin inhibit the enzymes that produce and then cross-link the strands of this polymer together.
This causes the cell wall to lose strength and the bacteria to burst.
In the figure, a molecule of penicillin (shown in a ball-and-stick form)
is shown bound to its target, the transpeptidase from the bacteria Streptomyces R61 (the protein is shown as a ribbon diagram).
Antibiotic drug design is facilitated when an enzyme that is essential to the pathogen's survival is absent or very different in humans. Humans do not make peptidoglycan, therefore antibiotics that inhibit this process are selectively toxic to bacteria. Selective toxicity is also produced in antibiotics by exploiting differences in the structure of the ribosomes in bacteria, or how they make fatty acids.
Antivirals
Drugs that inhibit enzymes needed for the replication of viruses are effective in treating viral infections. Antiviral drugs include protease inhibitors used to treat HIV/AIDS and Hepatitis C, reverse-transcriptase inhibitors targeting HIV/AIDS, neuraminidase inhibitors targeting influenza, and terminase inhibitors targeting human cytomegalovirus.
Pesticides
Many pesticides are enzyme inhibitors. Acetylcholinesterase
(AChE) is an enzyme found in animals, from insects to humans. It is
essential to nerve cell function through its mechanism of breaking down
the neurotransmitter acetylcholine into its constituents, acetate and choline. This is somewhat unusual among neurotransmitters as most, including serotonin, dopamine, and norepinephrine, are absorbed from the synaptic cleft rather than cleaved. A large number of AChE inhibitors are used in both medicine and agriculture. Reversible competitive inhibitors, such as edrophonium, physostigmine, and neostigmine, are used in the treatment of myasthenia gravis and in anaesthesia to reverse muscle blockade. The carbamate pesticides are also examples of reversible AChE inhibitors. The organophosphate pesticides such as malathion, parathion, and chlorpyrifos irreversibly inhibit acetylcholinesterase.
Herbicides
The herbicide glyphosate is an inhibitor of 3-phosphoshikimate 1-carboxyvinyltransferase, other herbicides, such as the sulfonylureas inhibit the enzyme acetolactate synthase. Both enzymes are needed for plants to make branched-chain amino acids. Many other enzymes are inhibited by herbicides, including enzymes needed for the biosynthesis of lipids and carotenoids and the processes of photosynthesis and oxidative phosphorylation.
Discovery and design of inhibitors
Robots used for the high-throughput screening of chemical libraries to discover new enzyme inhibitors
New drugs are the products of a long drug development process, the first step of which is often the discovery of a new enzyme inhibitor. There are two principle approaches of discovering these inhibitors.
The first general method is rational drug design based on mimicking the transition state of the chemical reaction catalysed by the enzyme.
The designed inhibitor often closely resembles the substrate, except
that the portion of the substrate that undergoes chemical reaction is
replaced a chemically stable functional group that resembles the transition state. Since the enzyme has evolved to stabilise the transition state, transition state analogues generally possess higher affinity for the enzyme compared to the substrate, and therefore are effective inhibitors.
The second way of discovering new enzyme inhibitors is high-throughput screening
of large libraries of structurally diverse compounds to identify hit
molecules that bind to the enzyme. This method has been extended to
include virtual screening of databases of diverse molecules using computers, which are then followed by experimental confirmation of binding of the virtual screening hits. Complementary approaches that can provide new starting points for inhibitors include fragment-based lead discovery and screening of DNA-encoded chemical libraries.
Hits from any of the above approaches can be optimised to high affinity binders that efficiently inhibit the enzyme. Computer-based methods for predicting the binding orientation and affinity of an inhibitor for an enzyme such as molecular docking and molecular mechanics can be used to assist in the optimisation process. New inhibitors are used to obtain crystallographic structures
of the enzyme in an inhibitor/enzyme complex to show how the molecule
is binding to the active site, allowing changes to be made to the
inhibitor to optimise binding in a process known as structure-based drug design. This test and improve cycle is repeated until a sufficiently potent inhibitor is produced.



![{\displaystyle {\begin{aligned}{\cfrac {V_{\max }}{1+{\cfrac {\ce {[I]}}{K_{i}}}}}&={V_{\max }}\left({\cfrac {K_{i}}{K_{i}+[{\ce {I}}]}}\right)&&{\text{multiply by }}{\cfrac {K_{i}}{K_{i}}}=1\\&={V_{\max }}\left({\cfrac {K_{i}+[{\ce {I}}]-[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{add }}[{\ce {I}}]-[{\ce {I}}]=0{\text{ to numerator}}\\&={V_{\max }}\left(1-{\cfrac {[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}\right)&&{\text{simplify }}{\cfrac {K_{i}+[{\ce {I}}]}{K_{i}+[{\ce {I}}]}}=1\\&=V_{\max }-V_{\max }{\cfrac {\ce {[I]}}{K_{i}+[{\ce {I}}]}}&&{\text{multiply out by }}V_{\max }\end{aligned}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/37eda4dec307f8acfca89b2d8f4811474ea764ec)
![{\displaystyle {\cfrac {\ce {[S]}}{[{\ce {S}}]+K_{m}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/eb08dd139085a394e6e7370f47ebfa255f1ad685)
![{\displaystyle {\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/9ed50a1f7a5f2c52f406b52263916ab48b268e07)
![{\displaystyle V_{\max }-\Delta V_{\max }{\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/7dff424ec79284c3a1cea14f0f82b0eaace53c69)
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[I]}}{[{\ce {I}}]+K_{i}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/f3874623edd9524ba2741fe448927bf5cf0ab257)
![{\displaystyle V_{\max 1}-(V_{\max 1}-V_{\max 2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/306d44733a89308883053e3b8372a8cf9ce0239b)
![{\displaystyle K_{m1}-(K_{m1}-K_{m2}){\cfrac {\ce {[X]}}{[{\ce {X}}]+K_{x}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/cb4e0de216e1e625bb803ee725bf85c9989a15f5)
![2D plots of 1/[S] concentration (x-axis) and 1/V (y-axis) demonstrating that as inhibitor concentration is changed, competitive inhibitor lines intersect at a single point on the y-axis, non-competitive inhibitors intersect at the x-axis, and mixed inhibitors intersect a point that is on neither axis.](https://upload.wikimedia.org/wikipedia/commons/thumb/b/ba/Inhibition_diagrams-1-.png/220px-Inhibition_diagrams-1-.png)
![V={\frac {V_{max}[S]}{\alpha K_{m}+\alpha ^{\prime }[S]}}={\frac {(1/\alpha ^{\prime })V_{max}[S]}{(\alpha /\alpha ^{\prime })K_{m}+[S]}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/4a8f0a9dda1d308de7f090f99c2833f944f11a09)
![\alpha =1+{\frac {[I]}{K_{i}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/57fcf54938a9784f9313437681b220079ff43ee5)
![\alpha ^{\prime }=1+{\frac {[I]}{K_{i}^{\prime }}}.](https://wikimedia.org/api/rest_v1/media/math/render/svg/65bf16742482cae7b0743781f47c327ddcf537e3)