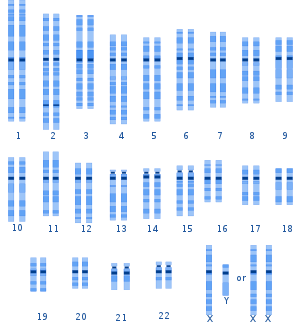Human evolutionary genetics studies how one human genome
differs from another human genome, the evolutionary past that gave rise
to the human genome, and its current effects. Differences between
genomes have anthropological, medical, historical and forensic implications and applications. Genetic data can provide important insights into human evolution.
Origin of apes
-10 —
–
-9 —
–
-8 —
–
-7 —
–
-6 —
–
-5 —
–
-4 —
–
-3 —
–
-2 —
–
-1 —
–
0 —
Biologists classify humans, along with only a few other species, as great apes (species in the family Hominidae). The living Hominidae include two distinct species of chimpanzee (the bonobo, Pan paniscus, and the common chimpanzee, Pan troglodytes), two species of gorilla (the western gorilla, Gorilla gorilla, and the eastern gorilla, Gorilla graueri), and two species of orangutan (the Bornean orangutan, Pongo pygmaeus, and the Sumatran orangutan, Pongo abelii). The great apes with the family Hylobatidae of gibbons form the superfamily Hominoidea of apes.
Apes, in turn, belong to the primate order (>400 species), along with the Old World monkeys, the New World monkeys, and others. Data from both mitochondrial DNA (mtDNA) and nuclear DNA (nDNA) indicate that primates belong to the group of Euarchontoglires, together with Rodentia, Lagomorpha, Dermoptera, and Scandentia. This is further supported by Alu-like short interspersed nuclear elements (SINEs) which have been found only in members of the Euarchontoglires.
Phylogenetics
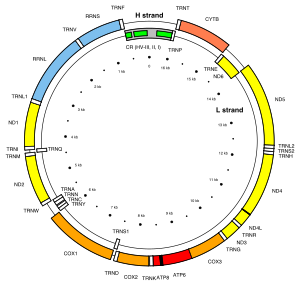
A phylogenetic tree is usually derived from DNA or protein sequences from populations. Often, mitochondrial DNA or Y chromosome sequences are used to study ancient human demographics. These single-locus sources of DNA do not recombine and are almost always inherited from a single parent, with only one known exception in mtDNA.
Individuals from closer geographic regions generally tend to be more
similar than individuals from regions farther away. Distance on a
phylogenetic tree can be used approximately to indicate:
- Genetic distance. The genetic difference between humans and chimpanzees is less than 2%, or three times larger than the variation among modern humans (estimated at 0.6%).
- Temporal remoteness of the most recent common ancestor. The mitochondrial most recent common ancestor of modern humans is estimated to have lived roughly 160,000 years ago, the latest common ancestors of humans and chimpanzees roughly 5 to 6 million years ago.
Speciation of humans and the African apes
The
separation of humans from their closest relatives, the non-human apes
(chimpanzees and gorillas), has been studied extensively for more than a
century. Five major questions have been addressed:
- Which apes are our closest ancestors?
- When did the separations occur?
- What was the effective population size of the common ancestor before the split?
- Are there traces of population structure (subpopulations) preceding the speciation or partial admixture succeeding it?
- What were the specific events (including fusion of chromosomes 2a and 2b) prior to and subsequent to the separation?
General observations
As discussed before, different parts of the genome show different sequence divergence between different hominoids.
It has also been shown that the sequence divergence between DNA from
humans and chimpanzees varies greatly. For example, the sequence
divergence varies between 0% to 2.66% between non-coding, non-repetitive
genomic regions of humans and chimpanzees.
The percentage of nucleotides in the human genome (hg38) that had
one-to-one exact matches in the chimpanzee genome (pantro6) was 84.38%.
Additionally gene trees, generated by comparative analysis of DNA
segments, do not always fit the species tree. Summing up:
- The sequence divergence varies significantly between humans, chimpanzees and gorillas.
- For most DNA sequences, humans and chimpanzees appear to be most closely related, but some point to a human-gorilla or chimpanzee-gorilla clade.
- The human genome has been sequenced, as well as the chimpanzee genome. Humans have 23 pairs of chromosomes, while chimpanzees, gorillas, and orangutans have 24. Human chromosome 2 is a fusion of two chromosomes 2a and 2b that remained separate in the other primates.
Divergence times
The
divergence time of humans from other apes is of great interest. One of
the first molecular studies, published in 1967 measured immunological
distances (IDs) between different primates. Basically the study measured the strength of immunological response that an antigen from one species (human albumin) induces in the immune system of another species (human, chimpanzee, gorilla and Old World monkeys).
Closely related species should have similar antigens and therefore
weaker immunological response to each other's antigens. The
immunological response of a species to its own antigens (e.g. human to
human) was set to be 1.
The ID between humans and gorillas was determined to be 1.09,
that between humans and chimpanzees was determined as 1.14. However the
distance to six different Old World monkeys was on average 2.46,
indicating that the African apes are more closely related to humans than
to monkeys. The authors consider the divergence time between Old World
monkeys and hominoids to be 30 million years ago (MYA), based on fossil
data, and the immunological distance was considered to grow at a
constant rate. They concluded that divergence time of humans and the
African apes to be roughly ~5 MYA. That was a surprising result. Most
scientists at that time thought that humans and great apes diverged much
earlier (>15 MYA).
The gorilla was, in ID terms, closer to human than to chimpanzees; however, the difference was so slight that the trichotomy
could not be resolved with certainty. Later studies based on molecular
genetics were able to resolve the trichotomy: chimpanzees are phylogenetically
closer to humans than to gorillas. However, some divergence times
estimated later (using much more sophisticated methods in molecular
genetics) do not substantially differ from the very first estimate in
1967, but a recent paper puts it at 11–14 MYA.
Divergence times and ancestral effective population size
The
sequences of the DNA segments diverge earlier than the species. A large
effective population size in the ancestral population (left) preserves
different variants of the DNA segments (=alleles) for a longer period of
time. Therefore, on average, the gene divergence times (tA for DNA segment A; tB for DNA segment B) will deviate more from the time the species diverge (tS) compared to a small ancestral effective population size (right).
Current methods to determine divergence times use DNA sequence alignments and molecular clocks.
Usually the molecular clock is calibrated assuming that the orangutan
split from the African apes (including humans) 12-16 MYA. Some studies
also include some old world monkeys and set the divergence time of them
from hominoids to 25-30 MYA. Both calibration points are based on very
little fossil data and have been criticized.
If these dates are revised, the divergence times estimated from
molecular data will change as well. However, the relative divergence
times are unlikely to change. Even if we can't tell absolute divergence
times exactly, we can be pretty sure that the divergence time between
chimpanzees and humans is about sixfold shorter than between chimpanzees
(or humans) and monkeys.
One study (Takahata et al., 1995) used 15 DNA sequences
from different regions of the genome from human and chimpanzee and 7 DNA
sequences from human, chimpanzee and gorilla.
They determined that chimpanzees are more closely related to humans
than gorillas. Using various statistical methods, they estimated the
divergence time human-chimp to be 4.7 MYA and the divergence time
between gorillas and humans (and chimps) to be 7.2 MYA.
Additionally they estimated the effective population size
of the common ancestor of humans and chimpanzees to be ~100,000. This
was somewhat surprising since the present day effective population size
of humans is estimated to be only ~10,000. If true that means that the
human lineage would have experienced an immense decrease of its
effective population size (and thus genetic diversity) in its evolution.
A and B are two different loci.
In the upper figure they fit to the species tree. The DNA that is
present in today's gorillas diverged earlier from the DNA that is
present in today's humans and chimps. Thus both loci should be more
similar between human and chimp than between gorilla and chimp or
gorilla and human. In the lower graph, locus A has a more recent common
ancestor in human and gorilla compared to the chimp sequence. Whereas
chimp and gorilla have a more recent common ancestor for locus B. Here
the gene trees are incongruent to the species tree.
Another study (Chen & Li, 2001) sequenced 53 non-repetitive,
intergenic DNA segments from a human, a chimpanzee, a gorilla, and
orangutan. When the DNA sequences were concatenated to a single long sequence, the generated neighbor-joining tree supported the Homo-Pan
clade with 100% bootstrap (that is that humans and chimpanzees are the
closest related species of the four). When three species are fairly
closely related to each other (like human, chimpanzee and gorilla), the
trees obtained from DNA sequence data may not be congruent with the tree
that represents the speciation (species tree).
The shorter internodal time span (TIN) the more common are incongruent gene trees. The effective population size (Ne)
of the internodal population determines how long genetic lineages are
preserved in the population. A higher effective population size causes
more incongruent gene trees. Therefore, if the internodal time span is
known, the ancestral effective population size of the common ancestor of
humans and chimpanzees can be calculated.
When each segment was analyzed individually, 31 supported the Homo-Pan clade, 10 supported the Homo-Gorilla clade, and 12 supported the Pan-Gorilla
clade. Using the molecular clock the authors estimated that gorillas
split up first 6.2-8.4 MYA and chimpanzees and humans split up 1.6-2.2
million years later (internodal time span) 4.6-6.2 MYA. The internodal
time span is useful to estimate the ancestral effective population size
of the common ancestor of humans and chimpanzees.
A parsimonious analysis revealed that 24 loci supported the Homo-Pan clade, 7 supported the Homo-Gorilla clade, 2 supported the Pan-Gorilla clade and 20 gave no resolution. Additionally they took 35 protein coding loci from databases. Of these 12 supported the Homo-Pan clade, 3 the Homo-Gorilla clade, 4 the Pan-Gorilla
clade and 16 gave no resolution. Therefore, only ~70% of the 52 loci
that gave a resolution (33 intergenic, 19 protein coding) support the
'correct' species tree. From the fraction of loci which did not support
the species tree and the internodal time span they estimated previously,
the effective population of the common ancestor of humans and
chimpanzees was estimated to be ~52 000 to 96 000. This value is not as
high as that from the first study (Takahata), but still much higher than
present day effective population size of humans.
A third study (Yang, 2002) used the same dataset that Chen and Li
used but estimated the ancestral effective population of 'only' ~12,000
to 21,000, using a different statistical method.
Genetic differences between humans and other great apes
The
alignable sequences within genomes of humans and chimpanzees differ by
about 35 million single-nucleotide substitutions. Additionally about 3%
of the complete genomes differ by deletions, insertions and
duplications.
Since mutation rate is relatively constant, roughly one half of
these changes occurred in the human lineage. Only a very tiny fraction
of those fixed differences gave rise to the different phenotypes of
humans and chimpanzees and finding those is a great challenge. The vast
majority of the differences are neutral and do not affect the phenotype.
Molecular evolution may act in different ways, through protein
evolution, gene loss, differential gene regulation and RNA evolution.
All are thought to have played some part in human evolution.
Gene loss
Many
different mutations can inactivate a gene, but few will change its
function in a specific way. Inactivation mutations will therefore be
readily available for selection to act on. Gene loss could thus be a
common mechanism of evolutionary adaptation (the "less-is-more"
hypothesis).
80 genes were lost in the human lineage after separation from the
last common ancestor with the chimpanzee. 36 of those were for olfactory receptors. Genes involved in chemoreception and immune response are overrepresented. Another study estimated that 86 genes had been lost.
Hair keratin gene KRTHAP1
A gene for type I hair keratin
was lost in the human lineage. Keratins are a major component of hairs.
Humans still have nine functional type I hair keratin genes, but the
loss of that particular gene may have caused the thinning of human body
hair. Based on the assumption of a constant molecular clock, the study
predicts the gene loss occurred relatively recently in human
evolution—less than 240 000 years ago, but both the Vindija Neandertal
and the high-coverage Denisovan sequence contain the same premature stop
codons as modern humans and hence dating should be greater than 750 000
years ago.
Myosin gene MYH16
Stedman et al. (2004) stated that the loss of the sarcomeric myosin gene MYH16 in the human lineage led to smaller masticatory muscles.
They estimated that the mutation that led to the inactivation (a two
base pair deletion) occurred 2.4 million years ago, predating the
appearance of Homo ergaster/erectus in Africa. The period that followed was marked by a strong increase in cranial capacity, promoting speculation that the loss of the gene may have removed an evolutionary constraint on brain size in the genus Homo.
Another estimate for the loss of the MYH16 gene is 5.3 million years ago, long before Homo appeared.
Other
- CASPASE12, a cysteinyl aspartate proteinase. The loss of this gene is speculated to have reduced the lethality of bacterial infection in humans.
Gene addition
Segmental duplications (SDs or LCRs) have had roles in creating new primate genes and shaping human genetic variation.
Human-specific DNA insertions
When
the human genome was compared to the genomes of five comparison primate
species, including the chimpanzee, gorilla, orangutan, gibbon, and
macaque, it was found that there are approximately 20,000 human-specific
insertions believed to be regulatory. While most insertions appear to
be fitness neutral, a small amount have been identified in positively
selected genes showing associations to neural phenotypes and some
relating to dental and sensory perception-related phenotypes. These
findings hint at the seemingly important role of human-specific
insertions in the recent evolution of humans.
Selection pressures
Human accelerated regions
are areas of the genome that differ between humans and chimpanzees to a
greater extent than can be explained by genetic drift over the time
since the two species shared a common ancestor. These regions show signs
of being subject to natural selection, leading to the evolution of
distinctly human traits. Two examples are HAR1F, which is believed to be related to brain development and HAR2 (a.k.a. HACNS1) that may have played a role in the development of the opposable thumb.
It has also been hypothesized that much of the difference between humans and chimpanzees is attributable to the regulation of gene expression rather than differences in the genes themselves. Analyses of conserved non-coding sequences, which often contain functional and thus positively selected regulatory regions, address this possibility.
Sequence divergence between humans and apes
When the draft sequence of the common chimpanzee (Pan troglodytes)
genome was published in the summer 2005, 2400 million bases (of ~3160
million bases) were sequenced and assembled well enough to be compared
to the human genome.
1.23% of this sequenced differed by single-base substitutions. Of this,
1.06% or less was thought to represent fixed differences between the
species, with the rest being variant sites in humans or chimpanzees.
Another type of difference, called indels
(insertions/deletions) accounted for many fewer differences (15% as
many), but contributed ~1.5% of unique sequence to each genome, since
each insertion or deletion can involve anywhere from one base to
millions of bases.
A companion paper examined segmental duplications in the two genomes,
whose insertion and deletion into the genome account for much of the
indel sequence. They found that a total of 2.7% of euchromatic sequence
had been differentially duplicated in one or the other lineage.
| Locus | Human-Chimp | Human-Gorilla | Human-Orangutan |
|---|---|---|---|
| Alu elements | 2 | - | - |
| Non-coding (Chr. Y) | 1.68 ± 0.19 | 2.33 ± 0.2 | 5.63 ± 0.35 |
| Pseudogenes (autosomal) | 1.64 ± 0.10 | 1.87 ± 0.11 | - |
| Pseudogenes (Chr. X) | 1.47 ± 0.17 | - | - |
| Noncoding (autosomal) | 1.24 ± 0.07 | 1.62 ± 0.08 | 3.08 ± 0.11 |
| Genes (Ks) | 1.11 | 1.48 | 2.98 |
| Introns | 0.93 ± 0.08 | 1.23 ± 0.09 | - |
| Xq13.3 | 0.92 ± 0.10 | 1.42 ± 0.12 | 3.00 ± 0.18 |
| Subtotal for X chromosome | 1.16 ± 0.07 | 1.47 ± 0.08 | - |
| Genes (Ka) | 0.8 | 0.93 | 1.96 |
The sequence divergence has generally the following pattern:
Human-Chimp < Human-Gorilla << Human-Orangutan, highlighting
the close kinship between humans and the African apes. Alu elements diverge quickly due to their high frequency of CpG
dinucleotides which mutate roughly 10 times more often than the average
nucleotide in the genome. The mutation rate is higher in the male germ line, therefore the divergence in the Y chromosome—which is inherited solely from the father—is higher than in autosomes. The X chromosome
is inherited twice as often through the female germ line as through the
male germ line and therefore shows slightly lower sequence divergence.
The sequence divergence of the Xq13.3 region is surprisingly low between
humans and chimpanzees.
Mutations altering the amino acid sequence of proteins (Ka)
are the least common. In fact ~29% of all orthologous proteins are
identical between human and chimpanzee. The typical protein differs by
only two amino acids.
The measures of sequence divergence shown in the table only take the substitutional differences, for example from an A (adenine) to a G (guanine), into account. DNA sequences may however also differ by insertions and deletions (indels) of bases. These are usually stripped from the alignments before the calculation of sequence divergence is performed.
Genetic differences between modern humans and Neanderthals
An international group of scientists completed a draft sequence of the Neanderthal genome in May 2010. The results indicate some breeding between modern humans (Homo sapiens) and Neanderthals (Homo neanderthalensis),
as the genomes of non-African humans have 1–4% more in common with
Neanderthals than do the genomes of subsaharan Africans. Neanderthals
and most modern humans share a lactose-intolerant variant of the lactase
gene that encodes an enzyme that is unable to break down lactose in
milk after weaning. Modern humans and Neanderthals also share the FOXP2
gene variant associated with brain development and with speech in
modern humans, indicating that Neanderthals may have been able to speak.
Chimps have two amino acid differences in FOXP2 compared with human and
Neanderthal FOXP2.
Genetic differences among modern humans
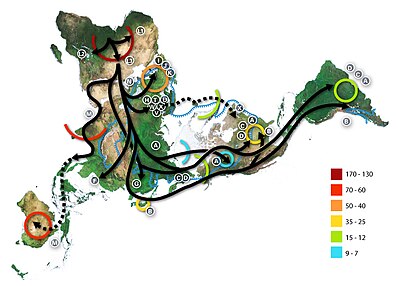
H. sapiens is thought to have emerged about 300,000 years ago. It dispersed throughout Africa, and after 70,000 years ago throughout Eurasia and Oceania.
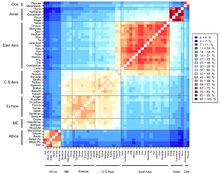
A 2009 study identified 14 "ancestral population clusters", the most remote being the San people of Southern Africa.
With their rapid expansion throughout different climate zones,
and especially with the availability of new food sources with the domestication of cattle and the development of agriculture, human populations have been exposed to significant selective pressures since their dispersal.
For example, East Asians have been found to be separated from Europids by a number of concentrated alleles suggestive of selection pressures, including variants of the EDAR, ADH1B, ABCC1, and ALDH2genes.
The East Asian types of ADH1B in particular are associated with rice domestication and would thus have arisen after the development of rice cultivation roughly 10,000 years ago. Several phenotypical traits of characteristic of East Asians are due to a single mutation of the EDAR gene, dated to c. 35,000 years ago.
As of 2017, the Single Nucleotide Polymorphism Database (dbSNP), which lists SNP and other variants, listed a total of 324 million variants found in sequenced human genomes.
Nucleotide diversity,
the average proportion of nucleotides that differ between two
individuals, is estimated at between 0.1% and 0.4% for contemporary
humans (compared to 2% between humans and chimpanzees).
This corresponds to genome differences at a few million sites; the 1000 Genomes Project
similarly found
that "a typical [individual] genome differs from the reference human
genome at 4.1 million to 5.0 million sites … affecting 20 million bases
of sequence."
In February 2019, scientists discovered evidence, based on genetics studies using artificial intelligence (AI), that suggest the existence of an unknown human ancestor species, not Neanderthal, Denisovan or human hybrid (like Denny (hybrid hominin)), in the genome of modern humans.