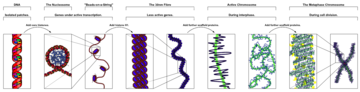
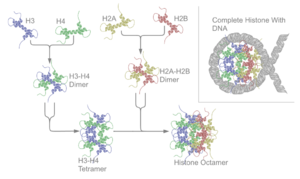
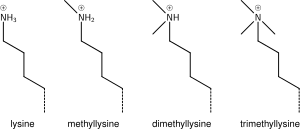
Human cancer cells with nuclei (specifically the DNA) stained blue. The central and rightmost cell are in interphase, so the entire nuclei are labeled. The cell on the left is going through mitosis and its DNA has condensed.
In biology, cell theory is the historic scientific theory, now universally accepted, that living organisms are made up of cells, that they are the basic structural/organizational unit of all organisms, and that all cells come from pre-existing cells. Cells are the basic unit of structure in all organisms and also the basic unit of reproduction. With continual improvements made to microscopes over time, magnification technology advanced enough to discover cells in the 17th century. This discovery is largely attributed to Robert Hooke, and began the scientific study of cells, also known as cell biology. Over a century later, many debates about cells began amongst scientists. Most of these debates involved the nature of cellular regeneration, and the idea of cells as a fundamental unit of life. Cell theory was eventually formulated in 1839. This is usually credited to Matthias Schleiden and Theodor Schwann. However, many other scientists like Rudolf Virchow contributed to the theory. It was an important step in the movement away from spontaneous generation.
The three tenets to the cell theory are as described below:
- All living organisms are composed of one or more cells.
- The cell is the basic unit of structure and organization in organisms.
- Cells arise from pre-existing cells.
Microscopes
Anton van Leeuwenhoek's microscope from the 17th century with a magnification of 300x.
Robert Hooke's microscope
The discovery of the cell was made possible through the invention of the microscope. In the first century BC, Romans were able to make glass, discovering that objects appeared to be larger under the glass. In Italy during the 12th century, Salvino D’Armate made a piece of glass fit over one eye, allowing for a magnification effect to that eye. The expanded use of lenses in eyeglasses in the 13th century probably led to wider spread use of simple microscopes (magnifying glasses) with limited magnification. Compound microscope, which combine an objective lens with an eyepiece to view a real image achieving much higher magnification, first appeared in Europe around 1620 In 1665, Robert Hooke used a microscope about six inches long with two convex lenses inside and examined specimens under reflected light for the observations in his book Micrographia. Hooke also used a simpler microscope with a single lens for examining specimens with directly transmitted light, because this allowed for a clearer image.
Extensive microscopic study was done by Anton van Leeuwenhoek, a draper who took the interest in microscopes after seeing one while on an apprenticeship in Amsterdam in 1648. At some point in his life before 1668, he was able to learn how to grind lenses. This eventually led to Leeuwenhoek making his own unique microscope. His were a single lens simple microscope, rather than a compound microscope. This was because he was able to use a single lens that was a small glass sphere but allowed for a magnification of 270x. This was a large progression since the magnification before was only a maximum of 50x. After Leeuwenhoek, there was not much progress for the microscopes until the 1850s, two hundred years later. Carl Zeiss, a German engineer who manufactured microscopes, began to make changes to the lenses used. But the optical quality did not improve until the 1880s when he hired Otto Schott and eventually Ernst Abbe.
Optical microscopes can focus on objects the size of a wavelength or larger, giving restrictions still to advancement in discoveries with objects smaller than the wavelengths of visible light. Later in the 1920s, the electron microscope was developed, making it possible to view objects that are smaller than optical wavelengths, once again, changing the possibilities in science.
Discovery of cells
The cell was first discovered by Robert Hooke in 1665, which can be found to be described in his book Micrographia. In this book, he gave 60 ‘observations’ in detail of various objects under a coarse, compound microscope. One observation was from very thin slices of bottle cork. Hooke discovered a multitude of tiny pores that he named "cells". This came from the Latin word Cella, meaning ‘a small room’ like monks lived in and also Cellulae, which meant the six sided cell of a honeycomb. However, Hooke did not know their real structure or function. What Hooke had thought were cells, were actually empty cell walls of plant tissues. With microscopes during this time having a low magnification, Hooke was unable to see that there were other internal components to the cells he was observing. Therefore, he did not think the "cellulae" were alive. His cell observations gave no indication of the nucleus and other organelles found in most living cells. In Micrographia, Hooke also observed mould, bluish in color, found on leather. After studying it under his microscope, he was unable to observe “seeds” that would have indicated how the mould was multiplying in quantity. This led to Hooke suggesting that spontaneous generation, from either natural or artificial heat, was the cause. Since this was an old Aristotelian theory still accepted at the time, others did not reject it and was not disproved until Leeuwenhoek later discovers generation is achieved otherwise.
Anton van Leeuwenhoek is another scientist who saw these cells soon after Hooke did. He made use of a microscope containing improved lenses that could magnify objects almost 300-fold, or 270x. Under these microscopes, Leeuwenhoek found motile objects. In a letter to The Royal Society on October 9, 1676, he states that motility is a quality of life therefore these were living organisms. Over time, he wrote many more papers in which described many specific forms of microorganisms. Leeuwenhoek named these “animalcules,” which included protozoa and other unicellular organisms, like bacteria. Though he did not have much formal education, he was able to identify the first accurate description of red blood cells and discovered bacteria after gaining interest in the sense of taste that resulted in Leeuwenhoek to observe the tongue of an ox, then leading him to study "pepper water" in 1676. He also found for the first time the sperm cells of animals and humans. Once discovering these types of cells, Leeuwenhoek saw that the fertilization process requires the sperm cell to enter the egg cell. This put an end to the previous theory of spontaneous generation. After reading letters by Leeuwenhoek, Hooke was the first to confirm his observations that were thought to be unlikely by other contemporaries.
The cells in animal tissues were observed after plants were because the tissues were so fragile and susceptible to tearing, it was difficult for such thin slices to be prepared for studying. Biologists believed that there was a fundamental unit to life, but were unsure what this was. It would not be until over a hundred years later that this fundamental unit was connected to cellular structure and existence of cells in animals or plants. This conclusion was not made until Henri Dutrochet. Besides stating “the cell is the fundamental element of organization”, Dutrochet also claimed that cells were not just a structural unit, but also a physiological unit.
In 1804, Karl Rudolphi and J.H.F. Link were awarded the prize for "solving the problem of the nature of cells", meaning they were the first to prove that cells had independent cell walls by the Königliche Societät der Wissenschaft (Royal Society of Science), Göttingen. Before, it had been thought that cells shared walls and the fluid passed between them this way.
Cell theory
Matthias Jakob Schleiden (1804–1881)
Theodor Schwann (1810–1882)
Credit for developing cell theory is usually given to two scientists: Theodor Schwann and Matthias Jakob Schleiden. While Rudolf Virchow contributed to the theory, he is not as credited for his attributions toward it. In 1839, Schleiden suggested that every structural part of a plant was made up of cells or the result of cells. He also suggested that cells were made by a crystallization process either within other cells or from the outside. However, this was not an original idea of Schleiden. He claimed this theory as his own, though Barthelemy Dumortier had stated it years before him. This crystallization process is no longer accepted with modern cell theory. In 1839, Theodor Schwann states that along with plants, animals are composed of cells or the product of cells in their structures. This was a major advancement in the field of biology since little was known about animal structure up to this point compared to plants. From these conclusions about plants and animals, two of the three tenets of cell theory were postulated.
1. All living organisms are composed of one or more cells
2. The cell is the most basic unit of life
Schleiden's theory of free cell formation through crystallization was refuted in the 1850s by Robert Remak, Rudolf Virchow, and Albert Kolliker. In 1855, Rudolf Virchow added the third tenet to cell theory. In Latin, this tenet states Omnis cellula e cellula. This translated to:
3. All cells arise only from pre-existing cells
However, the idea that all cells come from pre-existing cells had in fact already been proposed by Robert Remak; it has been suggested that Virchow plagiarized Remak and did not give him credit. Remak published observations in 1852 on cell division, claiming Schleiden and Schawnn were incorrect about generation schemes. He instead said that binary fission, which was first introduced by Dumortier, was how reproduction of new animal cells were made. Once this tenet was added, the classical cell theory was complete.
Modern interpretation
The generally accepted parts of modern cell theory include:- All known living things are made up of one or more cells
- All living cells arise from pre-existing cells by division.
- The cell is the fundamental unit of structure and function in all living organisms.
- The activity of an organism depends on the total activity of independent cells.
- Energy flow (metabolism and biochemistry) occurs within cells.
- Cells contain DNA which is found specifically in the chromosome and RNA found in the cell nucleus and cytoplasm.
- All cells are basically the same in chemical composition in organisms of similar species.
Modern version
The modern version of the cell theory includes the ideas that:- Energy flow occurs within cells.
- Heredity information (DNA) is passed on from cell to cell.
- All cells have the same basic chemical composition.
Opposing concepts in cell theory: history and background
The cell was first discovered by Robert Hooke in 1665 using a microscope. The first cell theory is credited to the work of Theodor Schwann and Matthias Jakob Schleiden in the 1830s. In this theory the internal contents of cells were called protoplasm and described as a jelly-like substance, sometimes called living jelly. At about the same time, colloidal chemistry began its development, and the concepts of bound water emerged. A colloid being something between a solution and a suspension, where Brownian motion is sufficient to prevent sedimentation. The idea of a semipermeable membrane, a barrier that is permeable to solvent but impermeable to solute molecules was developed at about the same time. The term osmosis originated in 1827 and its importance to physiological phenomena realized, but it wasn’t until 1877, when the botanist Pfeffer proposed the membrane theory of cell physiology. In this view, the cell was seen to be enclosed by a thin surface, the plasma membrane, and cell water and solutes such as a potassium ion existed in a physical state like that of a dilute solution. In 1889 Hamburger used hemolysis of erythrocytes to determine the permeability of various solutes. By measuring the time required for the cells to swell past their elastic limit, the rate at which solutes entered the cells could be estimated by the accompanying change in cell volume. He also found that there was an apparent nonsolvent volume of about 50% in red blood cells and later showed that this includes water of hydration in addition to the protein and other nonsolvent components of the cells.Evolution of the membrane and bulk phase theories
Two opposing concepts developed within the context of studies on osmosis, permeability, and electrical properties of cells. The first held that these properties all belonged to the plasma membrane whereas the other predominant view was that the protoplasm was responsible for these properties. The membrane theory developed as a succession of ad-hoc additions and changes to the theory to overcome experimental hurdles. Overton (a distant cousin of Charles Darwin) first proposed the concept of a lipid (oil) plasma membrane in 1899. The major weakness of the lipid membrane was the lack of an explanation of the high permeability to water, so Nathansohn (1904) proposed the mosaic theory. In this view, the membrane is not a pure lipid layer, but a mosaic of areas with lipid and areas with semipermeable gel. Ruhland refined the mosaic theory to include pores to allow additional passage of small molecules. Since membranes are generally less permeable to anions, Leonor Michaelis concluded that ions are adsorbed to the walls of the pores, changing the permeability of the pores to ions by electrostatic repulsion. Michaelis demonstrated the membrane potential (1926) and proposed that it was related to the distribution of ions across the membrane. Harvey and Danielli (1939) proposed a lipid bilayer membrane covered on each side with a layer of protein to account for measurements of surface tension. In 1941 Boyle & Conway showed that the membrane of frog muscle was permeable to both K+ and Cl−, but apparently not to Na+, so the idea of electrical charges in the pores was unnecessary since a single critical pore size would explain the permeability to K+, H+, and Cl− as well as the impermeability to Na+, Ca+, and Mg2+. Over the same time period, it was shown (Procter & Wilson, 1916) that gels, which do not have a semipermeable membrane, would swell in dilute solutions. Loeb (1920) also studied gelatin extensively, with and without a membrane, showing that more of the properties attributed to the plasma membrane could be duplicated in gels without a membrane. In particular, he found that an electrical potential difference between the gelatin and the outside medium could be developed, based on the H+ concentration. Some criticisms of the membrane theory developed in the 1930s, based on observations such as the ability of some cells to swell and increase their surface area by a factor of 1000. A lipid layer cannot stretch to that extent without becoming a patchwork (thereby losing its barrier properties. Such criticisms stimulated continued studies on protoplasm as the principal agent determining cell permeability properties. In 1938, Fischer and Suer proposed that water in the protoplasm is not free but in a chemically combined form—the protoplasm represents a combination of protein, salt and water—and demonstrated the basic similarity between swelling in living tissues and the swelling of gelatin and fibrin gels. Dimitri Nasonov (1944) viewed proteins as the central components responsible for many properties of the cell, including electrical properties. By the 1940s, the bulk phase theories were not as well developed as the membrane theories. In 1941, Brooks & Brooks published a monograph, "The Permeability of Living Cells", which rejects the bulk phase theories.Emergence of the steady-state membrane pump concept
With the development of radioactive tracers, it was shown that cells are not impermeable to Na+. This was difficult to explain with the membrane barrier theory, so the sodium pump was proposed to continually remove Na+ as it permeates cells. This drove the concept that cells are in a state of dynamic equilibrium, constantly using energy to maintain ion gradients. In 1935, Karl Lohmann discovered ATP and its role as a source of energy for cells, so the concept of a metabolically-driven sodium pump was proposed. The tremendous success of Hodgkin, Huxley, and Katz in the development of the membrane theory of cellular membrane potentials, with differential equations that modeled the phenomena correctly, provided even more support for the membrane pump hypothesis.The modern view of the plasma membrane is of a fluid lipid bilayer that has protein components embedded within it. The structure of the membrane is now known in great detail, including 3D models of many of the hundreds of different proteins that are bound to the membrane. These major developments in cell physiology placed the membrane theory in a position of dominance and stimulated the imagination of most physiologists, who now apparently accept the theory as fact—there are, however, a few dissenters.
The reemergence of the bulk phase theories
In 1956, Afanasy S. Troshin published a book, The Problems of Cell Permeability, in Russian (1958 in German, 1961 in Chinese, 1966 in English) in which he found that permeability was of secondary importance in determination of the patterns of equilibrium between the cell and its environment. Troshin showed that cell water decreased in solutions of galactose or urea although these compounds did slowly permeate cells. Since the membrane theory requires an impermanent solute to sustain cell shrinkage, these experiments cast doubt on the theory. Others questioned whether the cell has enough energy to sustain the sodium/potassium pump. Such questions became even more urgent as dozens of new metabolic pumps were added as new chemical gradients were discovered.In 1962, Gilbert Ling became the champion of the bulk phase theories and proposed his association-induction hypothesis of living cells.
Types of cells
Prokaryote cell.
Eukaryote cell.
Cells can be subdivided into the following subcategories:
- Prokaryotes: Prokaryotes are relatively small cells surrounded by the plasma membrane, with a characteristic cell wall that may differ in composition depending on the particular organism. Prokaryotes lack a nucleus (although they do have circular or linear DNA) and other membrane-bound organelles (though they do contain ribosomes). The protoplasm of a prokaryote contains the chromosomal region that appears as fibrous deposits under the microscope, and the cytoplasm. Bacteria and Archaea are the two domains of prokaryotes.
- Eukaryotes: Eukaryotic cells are also surrounded by the plasma membrane, but on the other hand, they have distinct nuclei bound by a nuclear membrane or envelope. Eukaryotic cells also contain membrane-bound organelles, such as (mitochondria, chloroplasts, lysosomes, rough and smooth endoplasmic reticulum, vacuoles). In addition, they possess organized chromosomes which store genetic material.