Successive dispersals of Homo erectus (yellow), Homo neanderthalensis (ochre) and Homo sapiens (red).
Expansion of early modern humans from Africa through the Near East
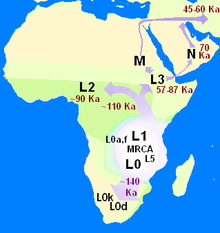
Map of the migration of modern humans out of Africa, based on mitochondrial DNA. Colored rings indicate thousand years before present.
In paleoanthropology, the recent African origin of modern humans, also called the "Out of Africa" theory (OOA), recent single-origin hypothesis (RSOH), replacement hypothesis, or recent African origin model (RAO), is the dominant model of the geographic origin and early migration of anatomically modern humans (Homo sapiens). It follows the early expansions of hominins out of Africa, accomplished by Homo erectus and then Homo neanderthalensis.
The model proposes a "single origin" of Homo sapiens in the taxonomic sense, precluding
parallel evolution of traits considered anatomically modern in other regions, but not precluding multiple admixture between H. sapiens and archaic humans in Europe and Asia.
H. sapiens most likely developed in the Horn of Africa between 300,000 and 200,000 years ago.
The "recent African origin" model proposes that all modern non-African
populations are substantially descended from populations of H. sapiens that left Africa after that time.
There were at least several "out-of-Africa" dispersals of modern
humans, possibly beginning as early as 270,000 years ago, including
215,000 years ago to at least Greece, and certainly via northern Africa about 130,000 to 115,000 years ago.
These early waves appear to have mostly died out or retreated by 80,000 years ago.
The most significant "recent" wave took place about 60,000–70,000 years ago, via the so-called "Southern Route", spreading rapidly along the coast of Asia and reaching Australia by around 65,000–50,000 years ago, while Europe was populated by an early offshoot which settled the Near East and Europe less than 55,000 years ago.
In the 2010s, studies in population genetics have uncovered evidence of interbreeding between H. sapiens and archaic humans in Eurasia and Oceania but not in Africa, which means that all non-African modern population groups, while mostly derived from early H. sapiens, are to a lesser extent also descended from regional variants of archaic humans.
Proposed waves
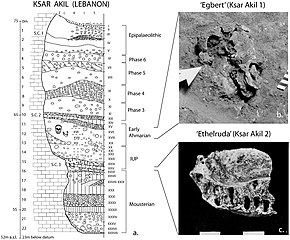
Layer sequence at Ksar Akil in the Levantine corridor, and discovery of two fossils of Homo sapiens, dated to 40,800 to 39,200 years BP for "Egbert",and 42,400–41,700 BP for "Ethelruda"
"Recent African origin," or Out of Africa II, refers to the migration of anatomically modern humans (Homo sapiens) out of Africa after their emergence at c. 300,000 to 200,000 years ago, in contrast to "Out of Africa I", which refers to the migration of archaic humans from Africa to Eurasia roughly 1.8 to 0.5 million years ago.
Since the beginning of the 21st century, the picture of "recent
single-origin" migrations has become significantly more complex, not
only due to the discovery of modern-archaic admixture but also due to
the increasing evidence that the "recent out-of-Africa" migration took
place in a number of waves spread over a long time period.
As of 2010, there were two main accepted dispersal routes for the
out-of-Africa migration of early anatomically modern humans: via the
"Northern Route" (via Nile Valley and Sinai) and the "Southern Route"
via the Bab al Mandab strait.
- Posth et al. (2017) suggest that early Homo sapiens, or "another species in Africa closely related to us," might have first migrated out of Africa around 270,000 years ago.
- Finds at Misliya cave, which include a partial jawbone with eight teeth, have been dated to around 185,000 years ago. Layers dating from between 250,000 and 140,000 years ago in the same cave contained tools of the Levallois type which could put the date of the first migration even earlier if the tools can be associated with the modern human jawbone finds.
- An Eastward Dispersal from Northeast Africa to Arabia 150,000–130,000 years ago based on the finds at Jebel Faya dated to 127,000 years ago (discovered in 2011). Possibly related to this wave are the finds from Zhirendong cave, Southern China, dated to more than 100,000 years ago. Other evidence of modern human presence in China has been dated to 80,000 years ago.
- The most significant dispersal took place around 60–70,000 years ago via the so-called Southern Route, either before or after the Toba event, which happened between 69,000 and 77,000 years ago. This dispersal followed the southern coastline of Asia, and reached Australia around 65,000-50,000 years ago. Western Asia was "re-occupied" by a different derivation from this wave around 50,000 years ago, and Europe was populated from Western Asia beginning around 43,000 years ago.
- Wells (2003) describes an additional wave of migration after the southern coastal route, namely a northern migration into Europe at circa 45,000 years ago. However, this possibility is ruled out by Macaulay et al. (2005) and Posth et al. (2016), who argue for a single coastal dispersal, with an early offshoot into Europe.
Northern Route dispersal
Anatomically
Modern Humans known archaeological remains in Europe and Africa,
directly dated, calibrated carbon dates as of 2013.
Beginning 135,000 years ago, tropical Africa experienced megadroughts which drove humans from the land and towards the sea shores, and forced them to cross over to other continents.
Modern humans crossed the Straits of Bab el Mandab in the southern Red Sea, and moved along the green coastlines around Arabia, and thence to the rest of Eurasia. Fossils of early Homo sapiens were found in Qafzeh
cave in Israel and have been dated 80,000 to 100,000 years ago. These
humans seem to have either become extinct or retreated back to Africa
70,000 to 80,000 years ago, possibly replaced by southbound Neanderthals
escaping the colder regions of ice-age Europe. Hua Liu et al. analyzed autosomal microsatellite markers dating to about 56,000 years ago. They interpret the paleontological fossil as an isolated early offshoot that retracted back to Africa.
The discovery of stone tools in the United Arab Emirates in 2011 at the Faya-1 site in Mleiha, Sharjah, indicated the presence of modern humans at least 125,000 years ago, leading to a resurgence of the "long-neglected" North African route.
In Oman,
a site was discovered by Bien Joven in 2011 containing more than 100
surface scatters of stone tools belonging to the late Nubian Complex,
known previously only from archaeological excavations in the Sudan.
Two optically stimulated luminescence age estimates placed the Arabian
Nubian Complex at approximately 106,000 years old. This provides
evidence for a distinct stone age technocomplex in southern Arabia, around the earlier part of the Marine Isotope Stage 5.
According to Kuhlwilm and his co-authors, Neanderthals contributed genetically
to modern humans then living outside of Africa around 100,000 years
ago: humans which had already split off from other modern humans around
200,000 years ago, and this early wave of modern humans outside Africa
also contributed genetically to the Altai Neanderthals.
They found that "the ancestors of Neanderthals from the Altai Mountains
and early modern humans met and interbred, possibly in the Near East,
many thousands of years earlier than previously thought".
According to co-author Ilan Gronau, "This actually complements
archaeological evidence of the presence of early modern humans out of
Africa around and before 100,000 years ago by providing the first
genetic evidence of such populations." Similar genetic admixture events have been noted in other regions as well.
In China, the Liujiang man (Chinese: 柳江人) is among the earliest modern humans found in East Asia.
The date most commonly attributed to the remains is 67,000 years ago.
High rates of variability yielded by various dating techniques carried
out by different researchers place the most widely accepted range of
dates with 67,000 BP as a minimum, but do not rule out dates as old as
159,000 BP. Liu, Martinón-Torres et al. (2015) claim that modern human teeth have been found in China dating to at least 80,000 years ago.
Southern Route dispersal
Coastal route
Red Sea crossing
By some 50-70,000 years ago, a subset of the bearers of mitochondrial haplogroup L3 migrated from East Africa into the Near East.
It has been estimated that from a population of 2,000 to 5,000
individuals in Africa, only a small group, possibly as few as 150 to
1,000 people, crossed the Red Sea. The group that crossed the Red Sea travelled along the coastal route around Arabia and Persia to India, which appears to have been the first major settling point. Wells (2003) argued for the route along the southern coastline of Asia, across about 250 kilometres (155 mi), reaching Australia by around 50,000 years ago.
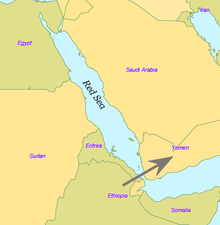
Today at the Bab-el-Mandeb straits, the Red Sea
is about 20 kilometres (12 mi) wide, but 50,000 years ago sea levels
were 70 m (230 ft) lower (owing to glaciation) and the water was much
narrower. Though the straits were never completely closed, they were
narrow enough to have enabled crossing using simple rafts, and there
may have been islands in between. Shell middens 125,000 years old have been found in Eritrea, indicating the diet of early humans included seafood obtained by beachcombing.
The dating of the Southern Dispersal is a matter of dispute.
It may have happened either pre- or post-Toba, a catastrophic volcanic
eruption that took place between 69,000 and 77,000 years ago at the site
of present-day Lake Toba.
Stone tools discovered below the layers of ash disposed in India may
point to a pre-Toba dispersal but the source of the tools is disputed.
An indication for post-Toba is haplo-group L3, that originated before
the dispersal of humans out of Africa and can be dated to 60,000–70,000
years ago, "suggesting that humanity left Africa a few thousand years
after Toba".
New research showing slower than expected genetic mutations in human
DNA was published in 2012, indicating a revised dating for the migration
to between 90,000 and 130,000 years ago.
Western Asia
A fossil of a modern human dated to 54,700 years ago was found in Manot Cave in Israel, named Manot 1, though the dating was questioned by Groucutt et al. (2015).
South-Asia and Australia
It
is thought that Australia was inhabited around 65,000–50,000 years ago.
As of 2017, the earliest evidence of humans in Australia is at least
65,000 years old, while McChesney stated that
...genetic evidence suggests that a small band with the marker M168 migrated out of Africa along the coasts of the Arabian Peninsula and India, through Indonesia, and reached Australia very early, between 60,000 and 50,000 years ago. This very early migration into Australia is also supported by Rasmussen et al. (2011).
Fossils from Lake Mungo, Australia, have been dated to about 42,000 years ago. Other fossils from a site called Madjedbebe have been dated to at least 65,000 years ago.
East Asia
Tianyuan man from China has a probable date range between 38,000 and 42,000 years ago, while Liujiang man
from the same region has a probable date range between 67,000 and
159,000 years ago. According to 2013 DNA tests, Tianyuan man is related
"to many present-day Asians and Native Americans". Tianyuan is similar in morphology to Minatogawa Man, modern humans dated between 17,000 and 19,000 years ago and found on Okinawa Island, Japan.
Europe
According to Macaulay et al. (2005),
an early offshoot from the southern dispersal with haplogroup N
followed the Nile from East Africa, heading northwards and crossing into
Asia through the Sinai. This group then branched, some moving into Europe and others heading east into Asia.
This hypothesis is supported by the relatively late date of the arrival
of modern humans in Europe as well as by archaeological and DNA
evidence. Based on an analysis of 55 human mitochondrial genomes (mtDNAs) of hunter-gatherers, Posth et al. (2016) argue for a "rapid single dispersal of all non-Africans less than 55,000 years ago."
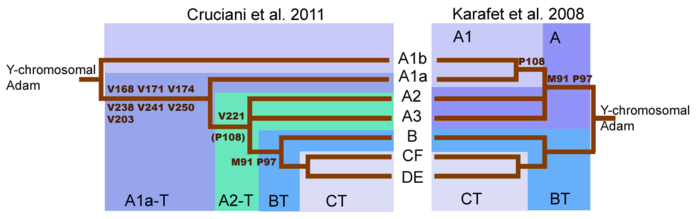
Genetic reconstruction
Mitochondrial haplogroups
Within Africa
Map of early diversification of modern humans according to mitochondrial population genetics.
The first lineage to branch off from Mitochondrial Eve was L0. This haplogroup is found in high proportions among the San of Southern Africa and the Sandawe of East Africa. It is also found among the Mbuti people. These groups branched off early in human history and have remained relatively genetically isolated since then. Haplogroups L1, L2 and L3 are descendants of L1–L6, and are largely confined to Africa. The macro haplogroups M and N,
which are the lineages of the rest of the world outside Africa, descend
from L3. L3 is about 84,000 years old, while haplogroups M and N are
about 63,000 years old. The relationship between such gene trees and demographic history is still debated when applied to dispersals.
Of all the lineages present in Africa, only the female descendants of one lineage, mtDNA haplogroup L3,
are found outside Africa. If there had been several migrations, one
would expect descendants of more than one lineage to be found. L3's
female descendants, the M and N haplogroup lineages, are found in very low frequencies in Africa (although haplogroup M1 populations are very ancient and diversified in North and North-east Africa)
and appear to be more recent arrivals. A possible explanation is that
these mutations occurred in East Africa shortly before the exodus and
became the dominant haplogroups thereafter by means of the founder effect. Alternatively, the mutations may have arisen shortly afterwards.
Southern Route and haplogroups M and N
Results from mtDNA collected from aboriginal Malaysians called Orang Asli
indicate that the hapologroups M and N share characteristics with
original African groups from approximately 85,000 years ago, and share
characteristics with sub-haplogroups found in coastal south-east Asian
regions, such as Australasia, the Indian subcontinent and throughout
continental Asia, which had dispersed and separated from their African
progenitor approximately 65,000 years ago. This southern coastal
dispersal would have occurred before the dispersal through the Levant
approximately 45,000 years ago.
This hypothesis attempts to explain why haplogroup N is predominant in
Europe and why haplogroup M is absent in Europe. Evidence of the coastal
migration is thought to have been destroyed by the rise in sea levels
during the Holocene epoch.
Alternatively, a small European founder population that had expressed
haplogroup M and N at first, could have lost haplogroup M through random
genetic drift resulting from a bottleneck (i.e. a founder effect).
The group that crossed the Red Sea travelled along the coastal route around Arabia and Persia until reaching India. Haplogroup M is found in high frequencies along the southern coastal regions of Pakistan and India and it has the greatest diversity in India, indicating that it is here where the mutation may have occurred. Sixty percent of the Indian population belong to Haplogroup M. The indigenous people of the Andaman Islands
also belong to the M lineage. The Andamanese are thought to be
offshoots of some of the earliest inhabitants in Asia because of their
long isolation from the mainland. They are evidence of the coastal route
of early settlers that extends from India to Thailand and Indonesia all the way to Papua New Guinea. Since M is found in high frequencies in highlanders from New Guinea and the Andamanese and New Guineans have dark skin and Afro-textured hair, some scientists think they are all part of the same wave of migrants who departed across the Red Sea ~60,000 years ago in the Great Coastal Migration. The proportion of haplogroup M increases eastwards from Arabia
to India; in eastern India, M outnumbers N by a ratio of 3:1. Crossing
into Southeast Asia, haplogroup N (mostly in the form of derivatives of
its R subclade) reappears as the predominant lineage. M is predominant in East Asia, but amongst Indigenous Australians, N is the more common lineage. This haphazard distribution of Haplogroup N from Europe to Australia can be explained by founder effects and population bottlenecks.
Autosomal DNA
A 2002 study of African, European and Asian populations, found
greater genetic diversity among Africans than among Eurasians, and that
genetic diversity among Eurasians is largely a subset of that among
Africans, supporting the out of Africa model. A large study by Coop et al. (2009) found evidence for natural selection in autosomal DNA outside of Africa. The study distinguishes non-African sweeps (notably KITLG variants associated with skin color), West-Eurasian sweeps (SLC24A5) and East-Asian sweeps (MC1R,
relevant to skin color). Based on this evidence, the study concluded
that human populations encountered novel selective pressures as they
expanded out of Africa. MC1R and its relation to skin color had already been discussed by Liu, Harding et al. (2000),
p. 135. According to this study, Papua New Guineans continued to be
exposed to selection for dark skin color so that, although these groups
are distinct from Africans in other places, the allele for dark skin
color shared by contemporary Africans, Andamanese and New Guineans is an
archaism. Endicott et al. (2003) suggest convergent evolution.
A 2014 study by Gurdasani et al. indicate that higher genetic diversity
in Africa was caused by relatively recent Eurasian migrations into Africa.
Pathogen DNA
Another promising route towards reconstructing human genetic genealogy is via the JC virus (JCV), a type of human polyomavirus
which is carried by 70–90 percent of humans and which is usually
transmitted vertically, from parents to offspring, suggesting
codivergence with human populations. For this reason, JCV has been used
as a genetic marker for human evolution and migration.
This method does not appear to be reliable for the migration out of
Africa, in contrast to human genetics, JCV strains associated with
African populations are not basal. From this Shackelton et al. (2006)
conclude that either a basal African strain of JCV has become extinct
or that the original infection with JCV post-dates the migration from
Africa.
Admixture of archaic and modern humans
Evidence for archaic human species (descended from Homo heidelbergensis) having interbred with modern humans outside of Africa, was discovered in the 2010s. This concerns primarily Neanderthal admixture in all modern populations except for Sub-Saharan Africans but evidence has also been presented for Denisova hominin admixture in Australasia (i.e. in Melanesians, Aboriginal Australians and some Negritos).
The rate of admixture of Neanderthal admixture to European and
Asian populations as of 2017 has been estimated at between about 2–3%.
Archaic admixture in some Sub-Saharan African populations hunter-gatherer groups (Biaka Pygmies and San),
derived from archaic hominins that broke away from the modern human
lineage around 700,000 years, was discovered in 2011. The rate of
admixture was estimated at around 2%.
Admixture from archaic hominins of still earlier divergence times, estimated at 1.2 to 1.3 million years ago, was found in Pygmies, Hadza and five Sandawe in 2012.
From an analysis of Mucin 7,
a highly divergent haplotype that has an estimated coalescence time
with other variants around 4.5 million years BP and is specific to
African populations is inferred to have been derived from interbreeding
between African modern and archaic humans.
Stone tools
In addition to genetic analysis, Petraglia et al. also examines the small stone tools (microlithic
materials) from the Indian subcontinent and explains the expansion of
population based on the reconstruction of paleoenvironment. He proposed
that the stone tools could be dated to 35 ka in South Asia, and the new
technology might be influenced by environmental change and population
pressure.
History of the theory
Classical paleoanthropology
The frontispiece to Huxley's Evidence as to Man's Place in Nature (1863): the image compares the skeleton of a human to other apes.
The cladistic relationship of humans with the African apes was suggested by Charles Darwin after studying the behaviour of African apes, one of which was displayed at the London Zoo. The anatomist Thomas Huxley had also supported the hypothesis and suggested that African apes have a close evolutionary relationship with humans. These views were opposed by the German biologist Ernst Haeckel, who was a proponent of the Out of Asia theory.
Haeckel argued that humans were more closely related to the primates of
South-east Asia and rejected Darwin's African hypothesis.
In the Descent of Man,
Darwin speculated that humans had descended from apes, which still had
small brains but walked upright, freeing their hands for uses which
favoured intelligence; he thought such apes were African:
In each great region of the world the living mammals are closely related to the extinct species of the same region. It is, therefore, probable that Africa was formerly inhabited by extinct apes closely allied to the gorilla and chimpanzee; and as these two species are now man's nearest allies, it is somewhat more probable that our early progenitors lived on the African continent than elsewhere. But it is useless to speculate on this subject, for an ape nearly as large as a man, namely the Dryopithecus of Lartet, which was closely allied to the anthropomorphous Hylobates, existed in Europe during the Upper Miocene period; and since so remote a period the earth has certainly undergone many great revolutions, and there has been ample time for migration on the largest scale.
— Charles Darwin, Descent of Man
In 1871 there were hardly any human fossils of ancient hominins
available. Almost fifty years later, Darwin's speculation was supported
when anthropologists began finding fossils of ancient small-brained
hominins in several areas of Africa (list of hominina fossils). The hypothesis of recent (as opposed to archaic)
African origin developed in the 20th century. The "Recent African
origin" of modern humans means "single origin" (monogenism) and has been
used in various contexts as an antonym to polygenism. The debate in
anthropology had swung in favour of monogenism by the mid-20th century.
Isolated proponents of polygenism held forth in the mid-20th century,
such as Carleton Coon, who thought as late as 1962 that H. sapiens arose five times from H. erectus in five places.
Multiregional origin hypothesis
The historical alternative to the recent origin model is the multiregional origin of modern humans, initially proposed by Milford Wolpoff in the 1980s. This view proposes that the derivation of anatomically modern human populations from H. erectus at the beginning of the Pleistocene
1.8 million years BP, has taken place within a continuous world
population. The hypothesis necessarily rejects the assumption of an infertility barrier between ancient Eurasian and African populations of Homo. The hypothesis was controversially debated during the late 1980s and the 1990s. The now-current terminology of "recent-origin" and "Out of Africa" became current in the context of this debate in the 1990s.
Originally seen as an antithetical alternative to the recent origin
model, the multiregional hypothesis in its original "strong" form is
obsolete, while its various modified weaker variants have become
variants of a view of "recent origin" combined with archaic admixture.
Stringer (2014) distinguishes the original or "classic" Multiregional
model as having existed from 1984 (its formulation) until 2003, to a
"weak" post-2003 variant that has "shifted close to that of the
Assimilation Model".
Genetics
In the 1980s, Allan Wilson together with Rebecca L. Cann and Mark Stoneking worked on genetic dating of the matrilineal most recent common ancestor of modern human populations (dubbed "Mitochondrial Eve"). To identify informative genetic markers for tracking human evolutionary history, Wilson concentrated on mitochondrial DNA
(mtDNA), passed from mother to child. This DNA material mutates
quickly, making it easy to plot changes over relatively short times.
With his discovery that human mtDNA is genetically much less diverse
than chimpanzee mtDNA, Wilson concluded that modern human populations
had diverged recently from a single population while older human species
such as Neanderthals and Homo erectus had become extinct. With the advent of archaeogenetics in the 1990s, the dating of mitochondrial and Y-chromosomal haplogroups became possible with some confidence. By 1999, estimates ranged around 150,000 years for the mt-MRCA and 60,000 to 70,000 years for the migration out of Africa.
From 2000–2003, there was controversy about the mitochondrial DNA of "Mungo Man 3"
(LM3) and its possible bearing on the multiregional hypothesis. LM3 was
found to have more than the expected number of sequence differences
when compared to modern human DNA (CRS).
Comparison of the mitochondrial DNA with that of ancient and modern
aborigines, led to the conclusion that Mungo Man fell outside the range
of genetic variation seen in Aboriginal Australians and was used to
support the multiregional origin hypothesis. A reanalysis on LM3 and
other ancient specimens from the area published in 2016, showed it to be
akin to modern Aboriginal Australian sequences, inconsistent with the
results of the earlier study.