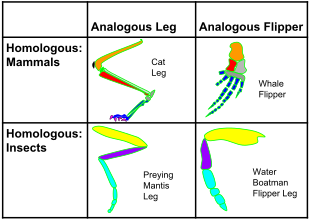Two succulent plant genera, Euphorbia and Astrophytum, are only distantly related, but the species within each have converged on a similar body form.
Convergent evolution is the independent evolution of similar features in species of different lineages. Convergent evolution creates analogous structures that have similar form or function but were not present in the last common ancestor of those groups. The cladistic term for the same phenomenon is homoplasy. The recurrent evolution of flight is a classic example, as flying insects, birds, pterosaurs, and bats have independently evolved the useful capacity of flight. Functionally similar features that have arisen through convergent evolution are analogous, whereas homologous structures or traits have a common origin but can have dissimilar functions. Bird, bat, and pterosaur wings are analogous structures, but their forelimbs are homologous, sharing an ancestral state despite serving different functions.
The opposite of convergence is divergent evolution, where related species evolve different traits. Convergent evolution is similar to parallel evolution, which occurs when two independent species evolve in the same direction and thus independently acquire similar characteristics; for instance, gliding frogs have evolved in parallel from multiple types of tree frog.
Many instances of convergent evolution are known in plants, including the repeated development of C4 photosynthesis, seed dispersal by fleshy fruits adapted to be eaten by animals, and carnivory.
Overview
Homology
and analogy in mammals and insects: on the horizontal axis, the
structures are homologous in morphology, but different in function due
to differences in habitat. On the vertical axis, the structures are
analogous in function due to similar lifestyles but anatomically
different with different phylogeny.
In morphology, analogous traits arise when different species live in
similar ways and/or a similar environment, and so face the same
environmental factors. When occupying similar ecological niches (that is, a distinctive way of life) similar problems can lead to similar solutions. The British anatomist Richard Owen was the first to identify the fundamental difference between analogies and homologies.
In biochemistry, physical and chemical constraints on mechanisms have caused some active site arrangements such as the catalytic triad to evolve independently in separate enzyme superfamilies.
In his 1989 book Wonderful Life, Stephen Jay Gould
argued that if one could "rewind the tape of life [and] the same
conditions were encountered again, evolution could take a very different
course". Simon Conway Morris
disputes this conclusion, arguing that convergence is a dominant force
in evolution, and given that the same environmental and physical
constraints are at work, life will inevitably evolve toward an "optimum"
body plan, and at some point, evolution is bound to stumble upon
intelligence, a trait presently identified with at least primates, corvids, and cetaceans.
Distinctions
Cladistics
In cladistics, a homoplasy is a trait shared by two or more taxa for any reason other than that they share a common ancestry. Taxa which do share ancestry are part of the same clade; cladistics seeks to arrange them according to their degree of relatedness to describe their phylogeny.
Homoplastic traits caused by convergence are therefore, from the point
of view of cladistics, confounding factors which could lead to an
incorrect analysis.
Atavism
In some cases, it is difficult to tell whether a trait has been lost and then re-evolved convergently, or whether a gene has simply been switched off and then re-enabled later. Such a re-emerged trait is called an atavism. From a mathematical standpoint, an unused gene (selectively neutral) has a steadily decreasing probability of retaining potential functionality over time. The time scale of this process varies greatly in different phylogenies; in mammals and birds, there is a reasonable probability of remaining in the genome in a potentially functional state for around 6 million years.Parallel vs. convergent evolution
Evolution at an amino acid
position. In each case, the left-hand species changes from having
alanine (A) at a specific position in a protein in a hypothetical
ancestor, and now has serine (S) there. The right-hand species may
undergo divergent, parallel, or convergent evolution at this amino acid position relative to the first species.
When two species are similar in a particular character, evolution is
defined as parallel if the ancestors were also similar, and convergent
if they were not.
Some scientists have argued that there is a continuum between parallel
and convergent evolution, while others maintain
When the ancestral forms are unspecified or unknown, or the range
of traits considered is not clearly specified, the distinction between
parallel and convergent evolution becomes more subjective. For
instance, the striking example of similar placental and marsupial forms
is described by Richard Dawkins in The Blind Watchmaker
as a case of convergent evolution, because mammals on each continent
had a long evolutionary history prior to the extinction of the dinosaurs
under which to accumulate relevant differences.
At molecular level
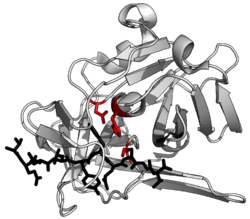
Evolutionary convergence of serine and cysteine protease towards the same catalytic triads organisation of acid-base-nucleophile in different protease superfamilies. Shown are the triads of subtilisin, prolyl oligopeptidase, TEV protease, and papain.
Protease active sites
The enzymology of proteases
provides some of the clearest examples of convergent evolution. These
examples reflect the intrinsic chemical constraints on enzymes, leading
evolution to converge on equivalent solutions independently and
repeatedly.
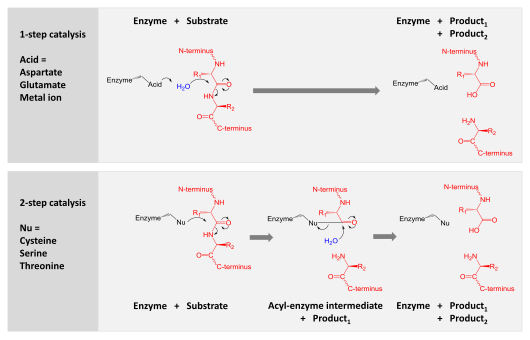
Serine and cysteine proteases use different amino acid functional groups (alcohol or thiol) as a nucleophile. In order to activate that nucleophile, they orient an acidic and a basic residue in a catalytic triad. The chemical and physical constraints on enzyme catalysis have caused identical triad arrangements to evolve independently more than 20 times in different enzyme superfamilies.
Threonine proteases use the amino acid threonine as their catalytic nucleophile. Unlike cysteine and serine, threonine is a secondary alcohol
(i.e. has a methyl group). The methyl group of threonine greatly
restricts the possible orientations of triad and substrate, as the
methyl clashes with either the enzyme backbone or the histidine base.
Consequently, most threonine proteases use an N-terminal threonine in
order to avoid such steric clashes.
Several evolutionarily independent enzyme superfamilies with different protein folds use the N-terminal residue as a nucleophile. This commonality of active site but difference of protein fold indicates that the active site evolved convergently in those families.
Nucleic acids
Convergence occurs at the level of DNA and the amino acid sequences produced by translating structural genes into proteins. Studies have found convergence in amino acid sequences in echolocating bats and the dolphin; among marine mammals; between giant and red pandas; and between the thylacine and canids. Convergence has also been detected in a type of non-coding DNA, cis-regulatory elements, such as in their rates of evolution; this could indicate either positive selection or relaxed purifying selection.
In animal morphology
Dolphins and ichthyosaurs converged on many adaptations for fast swimming.
Bodyplans
Swimming animals including fish such as herrings, marine mammals such as dolphins, and ichthyosaurs (of the Mesozoic) all converged on the same streamlined shape. The fusiform bodyshape (a tube tapered at both ends) adopted by many aquatic animals is an adaptation to enable them to travel at high speed in a high drag environment. Similar body shapes are found in the earless seals and the eared seals: they still have four legs, but these are strongly modified for swimming.
The marsupial fauna of Australia and the placental mammals of the Old
World have several strikingly similar forms, developed in two clades,
isolated from each other. The body and especially the skull shape of the thylacine (Tasmanian wolf) converged with those of Canidae such as the red fox, Vulpes vulpes.
Echolocation
As a sensory adaptation, echolocation has evolved separately in cetaceans (dolphins and whales) and bats, but from the same genetic mutations.
Eyes
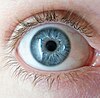
The camera eyes of vertebrates (left) and cephalopods (right) developed independently and are wired differently; for instance, optic nerve fibres reach the vertebrate retina from the front, creating a blind spot.
One of the best-known examples of convergent evolution is the camera eye of cephalopods (such as squid and octopus), vertebrates (including mammals) and cnidaria (such as jellyfish). Their last common ancestor had at most a simple photoreceptive spot, but a range of processes led to the progressive refinement of camera eyes
— with one sharp difference: the cephalopod eye is "wired" in the
opposite direction, with blood and nerve vessels entering from the back
of the retina, rather than the front as in vertebrates. As a result,
cephalopods lack a blind spot.
Flight
Vertebrate wings are partly homologous (from forelimbs), but analogous as organs of flight in (1) pterosaurs, (2) bats, (3) birds, evolved separately.
Birds and bats have homologous limbs because they are both ultimately derived from terrestrial tetrapods,
but their flight mechanisms are only analogous, so their wings are
examples of functional convergence. The two groups have powered flight,
evolved independently. Their wings differ substantially in
construction. The bat wing is a membrane stretched across four extremely
elongated fingers and the legs. The airfoil of the bird wing is made of
feathers, strongly attached to the forearm (the ulna) and the highly fused bones of the wrist and hand (the carpometacarpus),
with only tiny remnants of two fingers remaining, each anchoring a
single feather. So, while the wings of bats and birds are functionally
convergent, they are not anatomically convergent. Birds and bats also share a high concentration of cerebrosides
in the skin of their wings. This improves skin flexibility, a trait
useful for flying animals; other mammals have a far lower concentration. The extinct pterosaurs independently evolved wings from their fore- and hindlimbs, while insects have wings that evolved separately from different organs.
Flying squirrels and sugar gliders
are much alike in their body plans, with gliding wings stretched
between their limbs, but flying squirrels are placental mammals while
sugar gliders are marsupials, widely separated within the mammal
lineage.
Insect mouthparts
Insect mouthparts show many examples of convergent evolution. The mouthparts of different insect groups consist of a set of homologous
organs, specialised for the dietary intake of that insect group.
Convergent evolution of many groups of insects led from original
biting-chewing mouthparts to different, more specialised, derived
function types. These include, for example, the proboscis of flower-visiting insects such as bees and flower beetles, or the biting-sucking mouthparts of blood-sucking insects such as fleas and mosquitos.
Opposable thumbs
Opposable thumbs allowing the grasping of objects are most often associated with primates, like humans, monkeys, apes, and lemurs. Opposable thumbs also evolved in giant pandas,
but these are completely different in structure, having six fingers
including the thumb, which develops from a wrist bone entirely
separately from other fingers.
Primates
 Despite the similar lightening of skin colour after moving out of Africa, different genes were involved in European (left) and East-Asian (right) lineages. Despite the similar lightening of skin colour after moving out of Africa, different genes were involved in European (left) and East-Asian (right) lineages.
| ||
Convergent evolution in humans includes blue eye colour and light skin colour. When humans migrated out of Africa, they moved to more northern latitudes with less intense sunlight. It was beneficial to them to reduce their skin pigmentation. It appears certain that there was some lightening of skin colour before
European and East Asian lineages diverged, as there are some
skin-lightening genetic differences that are common to both groups.
However, after the lineages diverged and became genetically isolated,
the skin of both groups lightened more, and that additional lightening
was due to different genetic changes.
| Humans | Lemurs | ||
|---|---|---|---|
| Despite the similarity of appearance, the genetic basis of blue eyes is different in humans and lemurs. | |||
Lemurs and humans are both primates. Ancestral primates had brown eyes, as most primates do today. The genetic basis of blue eyes in humans has been studied in detail and much is known about it. It is not the case that one gene locus is responsible, say with brown dominant to blue eye colour. However, a single locus is responsible for about 80% of the variation. In lemurs, the differences between blue and brown eyes are not completely known, but the same gene locus is not involved.[42]
In plants
In myrmecochory, seeds such as those of Chelidonium majus have a hard coating and an attached oil body, an elaiosome, for dispersal by ants.
Carbon fixation
While convergent evolution is often illustrated with animal examples, it has often occurred in plant evolution. For instance, C4 photosynthesis, one of the three major carbon-fixing biochemical processes, has arisen independently up to 40 times. About 7,600 plant species of angiosperms use C4 carbon fixation, with many monocots including 46% of grasses such as maize and sugar cane, and dicots including several species in the Chenopodiaceae and the Amaranthaceae.
Fruits
A good example of convergence in plants is the evolution of edible fruits such as apples. These pomes incorporate (five) carpels and their accessory tissues forming the apple's core, surrounded by structures from outside the botanical fruit, the receptacle or hypanthium. Other edible fruits include other plant tissues; for example, the fleshy part of a tomato is the walls of the pericarp.
This implies convergent evolution under selective pressure, in this
case the competition for seed dispersal by animals through consumption
of fleshy fruits.
Seed dispersal by ants (myrmecochory)
has evolved independently more than 100 times, and is present in more
than 11,000 plant species. It is one of the most dramatic examples of
convergent evolution in biology.
Carnivory
Carnivory has evolved multiple times independently in plants in widely separated groups. In three species studied, Cephalotus follicularis, Nepenthes alata and Sarracenia purpurea, there has been convergence at the molecular level. Carnivorous plants secrete enzymes into the digestive fluid they produce. By studying phosphatase, glycoside hydrolase, glucanase, RNAse and chitinase enzymes as well as a pathogenesis-related protein and a thaumatin-related protein, the authors found many convergent amino acid
substitutions. These changes were not at the enzymes' catalytic sites,
but rather on the exposed surfaces of the proteins, where they might
interact with other components of the cell or the digestive fluid. The
authors also found that homologous genes in the non-carnivorous plant Arabidopsis thaliana
tend to have their expression increased when the plant is stressed,
leading the authors to suggest that stress-responsive proteins have
often been co-opted[c] in the repeated evolution of carnivory.
Methods of inference
Angiosperm
phylogeny of orders based on classification by the Angiosperm Phylogeny
Group. The figure shows the number of inferred independent origins of C3-C4 photosynthesis and C4 photosynthesis in parentheses.
Phylogenetic reconstruction and ancestral state reconstruction
proceed by assuming that evolution has occurred without convergence.
Convergent patterns may, however, appear at higher levels in a
phylogenetic reconstruction, and are sometimes explicitly sought by
investigators. The methods applied to infer convergent evolution depend
on whether pattern-based or process-based convergence is expected.
Pattern-based convergence is the broader term, for when two or more
lineages independently evolve patterns of similar traits. Process-based
convergence is when the convergence is due to similar forces of natural selection.
Pattern-based measures
Earlier methods for measuring convergence incorporate ratios of phenotypic and phylogenetic distance by simulating evolution with a Brownian motion model of trait evolution along a phylogeny. More recent methods also quantify the strength of convergence.
One drawback to keep in mind is that these methods can confuse
long-term stasis with convergence due to phenotypic similarities. Stasis
occurs when there is little evolutionary change among taxa.
Distance-based measures assess the degree of similarity between
lineages over time. Frequency-based measures assess the number of
lineages that have evolved in a particular trait space.
Process-based measures
Methods to infer process-based convergence fit models of selection to
a phylogeny and continuous trait data to determine whether the same
selective forces have acted upon lineages. This uses the Ornstein-Uhlenbeck (OU) process to test different scenarios of selection. Other methods rely on an a priori specification of where shifts in selection have occurred.