From Wikipedia, the free encyclopedia
Genomics is an interdisciplinary field of science focusing on the structure, function, evolution, mapping, and editing of
genomes. A genome is an organism's complete set of
DNA, including all of its genes. In contrast to
genetics,
which refers to the study of individual genes and their roles in
inheritance, genomics aims at the collective characterization and
quantification of genes, which direct the production of
proteins
with the assistance of enzymes and messenger molecules. In turn,
proteins make up body structures such as organs and tissues as well as
control chemical reactions and carry signals between cells. Genomics
also involves the sequencing and analysis of genomes through uses of
high throughput
DNA sequencing and
bioinformatics to assemble and analyze the function and structure of entire genomes.
[1][2][3] Advances in genomics have triggered a revolution in discovery-based research and
systems biology to facilitate understanding of even the most complex biological systems such as the brain.
[4]
The field also includes studies of intragenomic (within the genome) phenomena such as
epistasis (effect of one gene on another),
pleiotropy (one gene affecting more than one trait),
heterosis (hybrid vigour), and other interactions between
loci and
alleles within the genome.
[5]
History
Etymology
From the Greek ΓΕΝ
[6] gen,
"gene" (gamma, epsilon, nu, epsilon) meaning "become, create, creation,
birth", and subsequent variants: genealogy, genesis, genetics, genic,
genomere, genotype, genus etc. While the word
genome (from the
German Genom, attributed to
Hans Winkler) was in use in
English as early as 1926,
[7] the term
genomics was coined by Tom Roderick, a
geneticist at the
Jackson Laboratory (
Bar Harbor, Maine), over beer at a meeting held in
Maryland on the mapping of the human genome in 1986.
[8]
Early sequencing efforts
Following
Rosalind Franklin's confirmation of the helical structure of DNA,
James D. Watson and
Francis Crick's publication of the structure of DNA in 1953 and
Fred Sanger's publication of the
Amino acid sequence of insulin in 1955, nucleic acid sequencing became a major target of early
molecular biologists.
[9] In 1964,
Robert W. Holley and colleagues published the first nucleic acid sequence ever determined, the
ribonucleotide sequence of
alanine transfer RNA.
[10][11] Extending this work,
Marshall Nirenberg and
Philip Leder revealed the triplet nature of the
genetic code and were able to determine the sequences of 54 out of 64
codons in their experiments.
[12] In 1972,
Walter Fiers and his team at the Laboratory of Molecular Biology of the
University of Ghent (
Ghent,
Belgium) were the first to determine the sequence of a gene: the gene for
Bacteriophage MS2 coat protein.
[13]
Fiers' group expanded on their MS2 coat protein work, determining the
complete nucleotide-sequence of bacteriophage MS2-RNA (whose genome
encodes just four genes in 3569
base pairs [bp]) and
Simian virus 40 in 1976 and 1978, respectively.
[14][15]
DNA-sequencing technology developed
In addition to his seminal work on the amino acid sequence of insulin,
Frederick Sanger
and his colleagues played a key role in the development of DNA
sequencing techniques that enabled the establishment of comprehensive
genome sequencing projects.
[5]
In 1975, he and Alan Coulson published a sequencing procedure using DNA
polymerase with radiolabelled nucleotides that he called the
Plus and Minus technique.
[16][17]
This involved two closely related methods that generated short
oligonucleotides with defined 3' termini. These could be fractionated by
electrophoresis on a
polyacrylamide
gel (called polyacrylamide gel electrophoresis) and visualised using
autoradiography. The procedure could sequence up to 80 nucleotides in
one go and was a big improvement, but was still very laborious.
Nevertheless, in 1977 his group was able to sequence most of the 5,386
nucleotides of the single-stranded
bacteriophage φX174, completing the first fully sequenced DNA-based genome.
[18] The refinement of the
Plus and Minus method resulted in the chain-termination, or
Sanger method (see
below),
which formed the basis of the techniques of DNA sequencing, genome
mapping, data storage, and bioinformatic analysis most widely used in
the following quarter-century of research.
[19][20] In the same year
Walter Gilbert and
Allan Maxam of
Harvard University independently developed the
Maxam-Gilbert method (also known as the
chemical method) of DNA sequencing, involving the preferential cleavage of DNA at known bases, a less efficient method.
[21][22] For their groundbreaking work in the sequencing of nucleic acids, Gilbert and Sanger shared half the 1980
Nobel Prize in chemistry with
Paul Berg (
recombinant DNA).
Complete genomes
The advent of these technologies resulted in a rapid intensification in the scope and speed of completion of
genome sequencing projects. The first complete genome sequence of an
eukaryotic organelle, the human
mitochondrion (16,568 bp, about 16.6 kb [kilobase]), was reported in 1981,
[23] and the first
chloroplast genomes followed in 1986.
[24][25] In 1992, the first eukaryotic
chromosome, chromosome III of brewer's yeast
Saccharomyces cerevisiae (315 kb) was sequenced.
[26] The first free-living organism to be sequenced was that of
Haemophilus influenzae (1.8 Mb [megabase]) in 1995.
[27] The following year a consortium of researchers from laboratories across
North America,
Europe, and
Japan announced the completion of the first complete genome sequence of a eukaryote,
S. cerevisiae (12.1 Mb), and since then genomes have continued being sequenced at an exponentially growing pace.
[28] As of October 2011, the complete sequences are available for: 2,719
viruses, 1,115
archaea and
bacteria, and 36
eukaryotes, of which about half are
fungi.
[29][30]
The number of genome projects has increased as technological improvements continue to lower the cost of sequencing. (A) Exponential growth of genome sequence databases since 1995. (B) The cost in US Dollars (USD) to sequence one million bases. (C) The cost in USD to sequence a 3,000 Mb (human-sized) genome on a log-transformed scale.
Most of the microorganisms whose genomes have been completely sequenced are problematic
pathogens, such as
Haemophilus influenzae, which has resulted in a pronounced bias in their phylogenetic distribution compared to the breadth of microbial diversity.
[31][32]
Of the other sequenced species, most were chosen because they were
well-studied model organisms or promised to become good models. Yeast (
Saccharomyces cerevisiae) has long been an important
model organism for the
eukaryotic cell, while the fruit fly
Drosophila melanogaster has been a very important tool (notably in early pre-molecular
genetics). The worm
Caenorhabditis elegans is an often used simple model for
multicellular organisms. The zebrafish
Brachydanio rerio is used for many developmental studies on the molecular level, and the plant
Arabidopsis thaliana is a model organism for flowering plants. The
Japanese pufferfish (
Takifugu rubripes) and the
spotted green pufferfish (
Tetraodon nigroviridis) are interesting because of their small and compact genomes, which contain very little
noncoding DNA compared to most species.
[33][34] The mammals dog (
Canis familiaris),
[35] brown rat (
Rattus norvegicus), mouse (
Mus musculus), and chimpanzee (
Pan troglodytes) are all important model animals in medical research.
[22]
A rough draft of the
human genome was completed by the
Human Genome Project in early 2001, creating much fanfare.
[36]
This project, completed in 2003, sequenced the entire genome for one
specific person, and by 2007 this sequence was declared "finished" (less
than one error in 20,000 bases and all chromosomes assembled).
[36] In the years since then, the genomes of many other individuals have been sequenced, partly under the auspices of the
1000 Genomes Project, which announced the sequencing of 1,092 genomes in October 2012.
[37]
Completion of this project was made possible by the development of
dramatically more efficient sequencing technologies and required the
commitment of significant
bioinformatics resources from a large international collaboration.
[38] The continued analysis of human genomic data has profound political and social repercussions for human societies.
[39]
The "omics" revolution
The English-language
neologism omics informally refers to a field of study in
biology ending in
-omics, such as genomics,
proteomics or
metabolomics. The related suffix
-ome is used to address the objects of study of such fields, such as the
genome,
proteome or
metabolome respectively. The suffix
-ome as used in molecular biology refers to a
totality of some sort; similarly
omics
has come to refer generally to the study of large, comprehensive
biological data sets. While the growth in the use of the term has led
some scientists (
Jonathan Eisen, among others
[40]) to claim that it has been oversold,
[41]
it reflects the change in orientation towards the quantitative analysis
of complete or near-complete assortment of all the constituents of a
system.
[42] In the study of
symbioses,
for example, researchers which were once limited to the study of a
single gene product can now simultaneously compare the total complement
of several types of biological molecules.
[43][44]
Genome analysis
After an organism has been selected, genome projects involve three
components: the sequencing of DNA, the assembly of that sequence to
create a representation of the original chromosome, and the annotation
and analysis of that representation.
[5]
Overview of a genome project. First, the genome must be selected, which
involves several factors including cost and relevance. Second, the
sequence is generated and assembled at a given sequencing center (such
as
BGI or
DOE JGI). Third, the genome sequence is annotated at several levels: DNA, protein, gene pathways, or comparatively.
Sequencing
Historically, sequencing was done in
sequencing centers, centralized facilities (ranging from large independent institutions such as
Joint Genome Institute
which sequence dozens of terabases a year, to local molecular biology
core facilities) which contain research laboratories with the costly
instrumentation and technical support necessary. As sequencing
technology continues to improve, however, a new generation of effective
fast turnaround benchtop sequencers has come within reach of the average
academic laboratory.
[45][46] On the whole, genome sequencing approaches fall into two broad categories,
shotgun and
high-throughput (or
next-generation) sequencing.
[5]
Shotgun sequencing
An ABI PRISM 3100 Genetic Analyzer. Such capillary sequencers automated early large-scale genome sequencing efforts.
Shotgun sequencing is a sequencing method designed for analysis of
DNA sequences longer than 1000 base pairs, up to and including entire
chromosomes.
[47] It is named by analogy with the rapidly expanding, quasi-random firing pattern of a
shotgun.
Since gel electrophoresis sequencing can only be used for fairly short
sequences (100 to 1000 base pairs), longer DNA sequences must be broken
into random small segments which are then sequenced to obtain
reads.
Multiple overlapping reads for the target DNA are obtained by
performing several rounds of this fragmentation and sequencing. Computer
programs then use the overlapping ends of different reads to assemble
them into a continuous sequence.
[47][48] Shotgun sequencing is a random sampling process, requiring over-sampling to ensure a given
nucleotide is represented in the reconstructed sequence; the average number of reads by which a genome is over-sampled is referred to as
coverage.
[49]
For much of its history, the technology underlying shotgun sequencing was the classical chain-termination method or '
Sanger method', which is based on the selective incorporation of chain-terminating
dideoxynucleotides by
DNA polymerase during
in vitro DNA replication.
[18][50] Recently, shotgun sequencing has been supplanted by
high-throughput sequencing methods, especially for large-scale, automated
genome
analyses. However, the Sanger method remains in wide use, primarily for
smaller-scale projects and for obtaining especially long contiguous DNA
sequence reads (>500 nucleotides).
[51] Chain-termination methods require a single-stranded DNA template, a DNA
primer, a
DNA polymerase,
normal deoxynucleosidetriphosphates (dNTPs), and modified nucleotides
(dideoxyNTPs) that terminate DNA strand elongation. These
chain-terminating nucleotides lack a 3'-
OH group required for the formation of a
phosphodiester bond
between two nucleotides, causing DNA polymerase to cease extension of
DNA when a ddNTP is incorporated. The ddNTPs may be radioactively or
fluorescently labelled for detection in
DNA sequencers.
[5] Typically, these machines can sequence up to 96 DNA samples in a single batch (run) in up to 48 runs a day.
[52]
High-throughput sequencing
The high demand for low-cost sequencing has driven the development of high-throughput sequencing technologies that
parallelize the sequencing process, producing thousands or millions of sequences at once.
[53][54]
High-throughput sequencing is intended to lower the cost of DNA
sequencing beyond what is possible with standard dye-terminator methods.
In ultra-high-throughput sequencing, as many as 500,000
sequencing-by-synthesis operations may be run in parallel.
[55][56]
Illumina Genome Analyzer II System. Illumina technologies have set the
standard for high-throughput massively parallel sequencing.
[45]
The
Illumina dye sequencing
method is based on reversible dye-terminators and was developed in 1996
at the Geneva Biomedical Research Institute, by Pascal Mayer and
Laurent Farinelli.
[57] In this method, DNA molecules and primers are first attached on a slide and amplified with
polymerase
so that local clonal colonies, initially coined "DNA colonies", are
formed. To determine the sequence, four types of reversible terminator
bases (RT-bases) are added and non-incorporated nucleotides are washed
away. Unlike pyrosequencing, the DNA chains are extended one nucleotide
at a time and image acquisition can be performed at a delayed moment,
allowing for very large arrays of DNA colonies to be captured by
sequential images taken from a single camera. Decoupling the enzymatic
reaction and the image capture allows for optimal throughput and
theoretically unlimited sequencing capacity; with an optimal
configuration, the ultimate throughput of the instrument depends only on
the
A/D conversion rate of the camera. The camera takes images of the
fluorescently labeled nucleotides, then the dye along with the terminal 3' blocker is chemically removed from the DNA, allowing the next cycle.
[58]
An alternative approach,
ion semiconductor sequencing,
is based on standard DNA replication chemistry. This technology
measures the release of a hydrogen ion each time a base is incorporated.
A microwell containing template DNA is flooded with a single
nucleotide,
if the nucleotide is complementary to the template strand it will be
incorporated and a hydrogen ion will be released. This release triggers
an
ISFET ion sensor. If a
homopolymer
is present in the template sequence multiple nucleotides will be
incorporated in a single flood cycle, and the detected electrical signal
will be proportionally higher.
[59]
Assembly
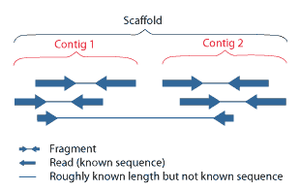
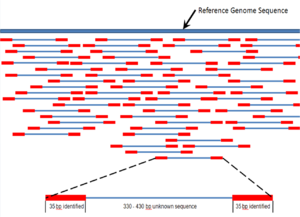
Overlapping reads form contigs; contigs and gaps of known length form scaffolds.
Paired end reads of next generation sequencing data mapped to a reference genome.
Multiple, fragmented sequence reads must be assembled together on the basis of their overlapping areas.
Sequence assembly refers to
aligning and merging fragments of a much longer
DNA sequence in order to reconstruct the original sequence.
[5] This is needed as current
DNA sequencing
technology cannot read whole genomes as a continuous sequence, but
rather reads small pieces of between 20 and 1000 bases, depending on the
technology used. 3rd generation sequencing technologies such as PacBio
or Oxford Nanopore routinly generate sequenceing reads >10 kb in
length; however, they have a high error rate at approximately 15%.
[60][61] Typically the short fragments, called reads, result from
shotgun sequencing genomic DNA, or
gene transcripts (
ESTs).
[5]
Assembly approaches
Assembly can be broadly categorized into two approaches:
de novo
assembly, for genomes which are not similar to any sequenced in the
past, and comparative assembly, which uses the existing sequence of a
closely related organism as a reference during assembly.
[49] Relative to comparative assembly,
de novo assembly is computationally difficult (
NP-hard), making it less favorable for short-read NGS technologies. Within the
de novo
assembly paradigm there are two primary strategies for assembly,
Eulerian path strategies, and overlap-layout-consensus (OLC) strategies.
OLC strategies ultimately try to create a Hamiltonian path through an
overlap graph which is an NP-hard problem. Eulerian path strategies are
computationally more tractable because they try to find a Eulerian path
through a deBruijn graph.
[49]
Finishing
Finished genomes are defined as having a single contiguous sequence with no ambiguities representing each
replicon.
[62]
Annotation
The DNA sequence assembly alone is of little value without additional analysis.
[5] Genome annotation is the process of attaching biological information to
sequences, and consists of three main steps:
[63]
- identifying portions of the genome that do not code for proteins
- identifying elements on the genome, a process called gene prediction, and
- attaching biological information to these elements.
Automatic annotation tools try to perform these steps
in silico, as opposed to manual annotation (a.k.a. curation) which involves human expertise and potential experimental verification.
[64] Ideally, these approaches co-exist and complement each other in the same annotation
pipeline (also see
below).
Traditionally, the basic level of annotation is using
BLAST for finding similarities, and then annotating genomes based on homologues.
[5]
More recently, additional information is added to the annotation
platform. The additional information allows manual annotators to
deconvolute discrepancies between genes that are given the same
annotation. Some databases use genome context information, similarity
scores, experimental data, and integrations of other resources to
provide genome annotations through their Subsystems approach. Other
databases (e.g.
Ensembl) rely on both curated data sources as well as a range of software tools in their automated genome annotation pipeline.
[65] Structural annotation consists of the identification of genomic elements, primarily
ORFs and their localisation, or gene structure.
Functional annotation consists of attaching biological information to genomic elements.
Sequencing pipelines and databases
The need for reproducibility and efficient management of the large amount of data associated with genome projects mean that
computational pipelines have important applications in genomics.
[66]
Research areas
Functional genomics
Functional genomics is a field of
molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as
genome sequencing projects) to describe
gene (and
protein) functions and interactions. Functional genomics focuses on the dynamic aspects such as gene
transcription,
translation, and
protein–protein interactions, as opposed to the static aspects of the genomic information such as
DNA sequence
or structures. Functional genomics attempts to answer questions about
the function of DNA at the levels of genes, RNA transcripts, and protein
products. A key characteristic of functional genomics studies is their
genome-wide approach to these questions, generally involving
high-throughput methods rather than a more traditional “gene-by-gene”
approach.
A major branch of genomics is still concerned with
sequencing the genomes of various organisms, but the knowledge of full genomes has created the possibility for the field of
functional genomics, mainly concerned with patterns of
gene expression during various conditions. The most important tools here are
microarrays and
bioinformatics.
Structural genomics
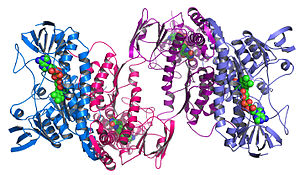
An example of a protein structure determined by the Midwest Center for Structural Genomics.
Structural genomics seeks to describe the
3-dimensional structure of every protein encoded by a given
genome.
[67][68] This genome-based approach allows for a high-throughput method of structure determination by a combination of
experimental and modeling approaches. The principal difference between structural genomics and
traditional structural prediction
is that structural genomics attempts to determine the structure of
every protein encoded by the genome, rather than focusing on one
particular protein. With full-genome sequences available, structure
prediction can be done more quickly through a combination of
experimental and modeling approaches, especially because the
availability of large numbers of sequenced genomes and previously solved
protein structures allow scientists to model protein structure on the
structures of previously solved homologs. Structural genomics involves
taking a large number of approaches to structure determination,
including experimental methods using genomic sequences or modeling-based
approaches based on sequence or
structural homology
to a protein of known structure or based on chemical and physical
principles for a protein with no homology to any known structure. As
opposed to traditional
structural biology, the determination of a
protein structure
through a structural genomics effort often (but not always) comes
before anything is known regarding the protein function. This raises new
challenges in
structural bioinformatics, i.e. determining protein function from its
3D structure.
[69]
Epigenomics
Epigenomics is the study of the complete set of
epigenetic modifications on the genetic material of a cell, known as the
epigenome.
[70]
Epigenetic modifications are reversible modifications on a cell’s DNA
or histones that affect gene expression without altering the DNA
sequence (Russell 2010 p. 475). Two of the most characterized epigenetic
modifications are
DNA methylation and
histone modification.
Epigenetic modifications play an important role in gene expression and
regulation, and are involved in numerous cellular processes such as in
differentiation/development and
tumorigenesis.
[70]
The study of epigenetics on a global level has been made possible only
recently through the adaptation of genomic high-throughput assays.
[71]
Metagenomics
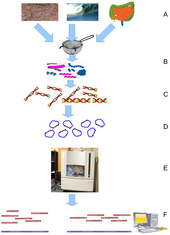
Environmental Shotgun Sequencing (ESS) is a key technique in
metagenomics. (A) Sampling from habitat; (B) filtering particles,
typically by size; (C) Lysis and DNA extraction; (D) cloning and library
construction; (E) sequencing the clones; (F) sequence assembly into
contigs and scaffolds.
Metagenomics is the study of
metagenomes,
genetic material recovered directly from
environmental
samples. The broad field may also be referred to as environmental
genomics, ecogenomics or community genomics. While traditional
microbiology and microbial
genome sequencing rely upon cultivated
clonal cultures, early environmental gene sequencing cloned specific genes (often the
16S rRNA gene) to produce a
profile of diversity in a natural sample. Such work revealed that the vast majority of
microbial biodiversity had been missed by
cultivation-based methods.
[72] Recent studies use "shotgun"
Sanger sequencing or massively parallel
pyrosequencing to get largely unbiased samples of all genes from all the members of the sampled communities.
[73]
Because of its power to reveal the previously hidden diversity of
microscopic life, metagenomics offers a powerful lens for viewing the
microbial world that has the potential to revolutionize understanding of
the entire living world.
[74][75]
Model systems
Viruses and bacteriophages
Bacteriophages have played and continue to play a key role in bacterial
genetics and
molecular biology. Historically, they were used to define
gene structure and gene regulation. Also the first
genome to be sequenced was a
bacteriophage.
However, bacteriophage research did not lead the genomics revolution,
which is clearly dominated by bacterial genomics. Only very recently has
the study of bacteriophage genomes become prominent, thereby enabling
researchers to understand the mechanisms underlying
phage
evolution. Bacteriophage genome sequences can be obtained through
direct sequencing of isolated bacteriophages, but can also be derived as
part of microbial genomes. Analysis of bacterial genomes has shown that
a substantial amount of microbial DNA consists of
prophage sequences and prophage-like elements.
[76] A detailed database mining of these sequences offers insights into the role of prophages in shaping the bacterial genome.
[77][78]
Cyanobacteria
At present there are 24
cyanobacteria for which a total genome sequence is available. 15 of these cyanobacteria come from the marine environment. These are six
Prochlorococcus strains, seven marine
Synechococcus strains,
Trichodesmium erythraeum IMS101 and
Crocosphaera watsonii WH8501.
Several studies have demonstrated how these sequences could be used
very successfully to infer important ecological and physiological
characteristics of marine cyanobacteria. However, there are many more
genome projects currently in progress, amongst those there are further
Prochlorococcus and marine
Synechococcus isolates,
Acaryochloris and
Prochloron, the N
2-fixing filamentous cyanobacteria
Nodularia spumigena,
Lyngbya aestuarii and
Lyngbya majuscula, as well as
bacteriophages
infecting marine cyanobaceria. Thus, the growing body of genome
information can also be tapped in a more general way to address global
problems by applying a comparative approach. Some new and exciting
examples of progress in this field are the identification of genes for
regulatory RNAs, insights into the evolutionary origin of
photosynthesis, or estimation of the contribution of horizontal gene transfer to the genomes that have been analyzed.
[79]
Applications of genomics
Genomics has provided applications in many fields, including
medicine,
biotechnology,
anthropology and other
social sciences.
[39]
Genomic medicine
Next-generation
genomic technologies allow clinicians and biomedical researchers to
drastically increase the amount of genomic data collected on large study
populations.
[80]
When combined with new informatics approaches that integrate many kinds
of data with genomic data in disease research, this allows researchers
to better understand the genetic bases of drug response and disease.
[81][82]
Synthetic biology and bioengineering
The growth of genomic knowledge has enabled increasingly sophisticated applications of
synthetic biology.
[83] In 2010 researchers at the
J. Craig Venter Institute announced the creation of a partially synthetic species of
bacterium,
Mycoplasma laboratorium, derived from the
genome of
Mycoplasma genitalium.
[84]
Conservation genomics
Conservationists
can use the information gathered by genomic sequencing in order to
better evaluate genetic factors key to species conservation, such as the
genetic diversity of a population or whether an individual is heterozygous for a recessive inherited genetic disorder.
[85] By using genomic data to evaluate the effects of
evolutionary processes
and to detect patterns in variation throughout a given population,
conservationists can formulate plans to aid a given species without as
many variables left unknown as those unaddressed by standard
genetic approaches.
[86]