From Wikipedia, the free encyclopedia
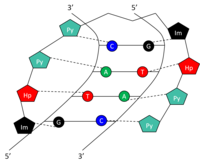
Gene therapy using an
adenovirus
vector. In some cases, the adenovirus will insert the new gene into a
cell. If the treatment is successful, the new gene will make a
functional
protein to treat a disease.
In the
medicine field,
gene therapy (also called
human gene transfer) is the therapeutic delivery of
nucleic acid into a patient's cells as a
drug to treat disease. The first attempt at modifying human
DNA was performed in 1980 by
Martin Cline, but the first successful nuclear gene transfer in humans, approved by the
National Institutes of Health, was performed in May 1989.
The first therapeutic use of gene transfer as well as the first direct
insertion of human DNA into the nuclear genome was performed by
French Anderson in a trial starting in September 1990.
Between 1989 and February 2016, over 2,300 clinical trials had been conducted, more than half of them in
phase I.
Not all medical procedures that introduce alterations to a patient's genetic makeup can be considered gene therapy.
Bone marrow transplantation and
organ transplants in general have been found to introduce foreign DNA into patients.
[5] Gene therapy is defined by the precision of the procedure and the intention of direct therapeutic effects.
Background
Gene therapy was conceptualized in 1972, by authors who urged caution before commencing human gene therapy studies.
The first attempt, an unsuccessful one, at gene therapy (as well
as the first case of medical transfer of foreign genes into humans not
counting
organ transplantation) was performed by
Martin Cline on 10 July 1980.
[6][7]
Cline claimed that one of the genes in his patients was active six
months later, though he never published this data or had it verified
[8] and even if he is correct, it's unlikely it produced any significant beneficial effects treating
beta-thalassemia.
After extensive research on animals throughout the 1980s and a
1989 bacterial gene tagging trial on humans, the first gene therapy
widely accepted as a success was demonstrated in a trial that started on
14 September 1990, when Ashi DeSilva was treated for
ADA-
SCID.
[9]
The first somatic treatment that produced a permanent genetic change was performed in 1993.
[citation needed]
Gene therapy is a way to fix a genetic problem at its source. The polymers are either
translated into
proteins, interfere with target
gene expression, or possibly correct
genetic mutations.
The most common form uses
DNA that encodes a functional, therapeutic
gene to replace a
mutated gene. The polymer molecule is packaged within a "
vector", which carries the molecule inside cells.
Early clinical failures led to dismissals of gene therapy.
Clinical successes since 2006 regained researchers' attention, although
as of 2014, it was still largely an experimental technique.
[10] These include treatment of
retinal diseases Leber's congenital amaurosis[11][12][13][14] and
choroideremia,
[15] X-linked SCID,
[16] ADA-SCID,
[17][18] adrenoleukodystrophy,
[19] chronic lymphocytic leukemia (CLL),
[20] acute lymphocytic leukemia (ALL),
[21] multiple myeloma,
[22] haemophilia,
[18] and
Parkinson's disease.
[23] Between 2013 and April 2014, US companies invested over $600 million in the field.
[24]
The first commercial gene therapy,
Gendicine, was approved in China in 2003 for the treatment of certain cancers.
[25] In 2011
Neovasculgen was registered in Russia as the first-in-class gene-therapy drug for treatment of
peripheral artery disease, including
critical limb ischemia.
[26]
In 2012
Glybera, a treatment for a rare
inherited disorder, became the first treatment to be approved for clinical use in either Europe or the United States after its endorsement by the
European Commission.
[10][27]
Approaches
Following early advances in
genetic engineering
of bacteria, cells, and small animals, scientists started considering
how to apply it to medicine. Two main approaches were considered –
replacing or disrupting defective genes.
[28] Scientists focused on diseases caused by single-gene defects, such as
cystic fibrosis,
haemophilia,
muscular dystrophy,
thalassemia, and
sickle cell anemia.
Glybera treats one such disease, caused by a defect in
lipoprotein lipase.
[27]
DNA must be administered, reach the damaged cells, enter the cell and either express or disrupt a protein.
[29] Multiple delivery techniques have been explored. The initial approach incorporated DNA into an engineered
virus to deliver the DNA into a
chromosome.
[30][31] Naked DNA approaches have also been explored, especially in the context of
vaccine development.
[32]
Generally, efforts focused on administering a gene that causes a
needed protein to be expressed. More recently, increased understanding
of
nuclease function has led to more direct DNA editing, using techniques such as
zinc finger nucleases and
CRISPR.
The vector incorporates genes into chromosomes. The expressed nucleases
then knock out and replace genes in the chromosome. As of 2014 these
approaches involve removing cells from patients, editing a chromosome
and returning the transformed cells to patients.
[33]
Gene editing is a potential approach to alter the human genome to treat genetic diseases,
[34] viral diseases,
[35] and cancer.
[36] As of 2016 these approaches were still years from being medicine.
[37][38]
A duplex of crRNA and
tracrRNA
acts as guide RNA to
introduce a specifically located gene modification
based on
the RNA 5’ upstream of the crRNA. Cas9 binds the tracrRNA
and
needs a DNA binding sequence (5’NGG3’), which is
called protospacer
adjacent motif (PAM). After binding, Cas9
introduces a DNA double strand
break, which is then followed
by gene modification via homologous
recombination (HDR) or
non-homologous end joining (NHEJ).
Cell types
Gene therapy may be classified into two types:
Somatic
In
somatic cell gene therapy (SCGT), the therapeutic genes are transferred into any cell other than a
gamete,
germ cell,
gametocyte, or undifferentiated
stem cell. Any such modifications affect the individual patient only, and are not inherited by
offspring. Somatic gene therapy represents mainstream basic and clinical research, in which therapeutic DNA (either integrated in the
genome or as an external
episome or
plasmid) is used to treat disease.
Over 600
clinical trials utilizing SCGT are underway in the US. Most focus on severe genetic disorders, including
immunodeficiencies,
haemophilia,
thalassaemia, and
cystic fibrosis.
Such single gene disorders are good candidates for somatic cell
therapy. The complete correction of a genetic disorder or the
replacement of multiple genes is not yet possible. Only a few of the
trials are in the advanced stages.
[39]
Germline
In
germline gene therapy (GGT),
germ cells (
sperm or
egg cells)
are modified by the introduction of functional genes into their
genomes. Modifying a germ cell causes all the organism's cells to
contain the modified gene. The change is therefore
heritable and passed on to later generations. Australia, Canada, Germany, Israel, Switzerland, and the Netherlands
[40]
prohibit GGT for application in human beings, for technical and ethical
reasons, including insufficient knowledge about possible risks to
future generations
[40] and higher risks versus SCGT.
[41]
The US has no federal controls specifically addressing human genetic
modification (beyond FDA regulations for therapies in general).
[40][42][43][44]
Vectors
The delivery of DNA into cells can be accomplished by multiple
methods. The two major classes are
recombinant viruses (sometimes called biological nanoparticles or viral vectors) and
naked DNA or DNA complexes (non-viral methods).
Viruses
In order to
replicate,
viruses
introduce their genetic material into the host cell, tricking the
host's cellular machinery into using it as blueprints for viral
proteins.
Retroviruses
go a stage further by having their genetic material copied into the
genome of the host cell. Scientists exploit this by substituting a
virus's genetic material with therapeutic DNA. (The term 'DNA' may be an
oversimplification, as some viruses contain RNA, and gene therapy could
take this form as well.) A number of viruses have been used for human
gene therapy, including
retroviruses,
adenoviruses,
herpes simplex,
vaccinia, and
adeno-associated virus.
[4]
Like the genetic material (DNA or RNA) in viruses, therapeutic DNA can
be designed to simply serve as a temporary blueprint that is degraded
naturally or (at least theoretically) to enter the host's genome,
becoming a permanent part of the host's DNA in infected cells.
Non-viral
Non-viral methods present certain advantages over viral methods, such as large scale production and low host
immunogenicity. However, non-viral methods initially produced lower levels of
transfection and
gene expression, and thus lower therapeutic efficacy. Later technology remedied this deficiency.
Methods for non-viral gene therapy include the injection of naked DNA,
electroporation, the
gene gun,
sonoporation,
magnetofection, the use of
oligonucleotides, lipoplexes, dendrimers, and inorganic nanoparticles.
Hurdles
Some of the unsolved problems include:
- Short-lived nature – Before gene therapy can become a permanent
cure for a condition, the therapeutic DNA introduced into target cells
must remain functional and the cells containing the therapeutic DNA must
be stable. Problems with integrating therapeutic DNA into the genome
and the rapidly dividing nature of many cells prevent it from achieving
long-term benefits. Patients require multiple treatments.
- Immune response – Any time a foreign object is introduced into human
tissues, the immune system is stimulated to attack the invader.
Stimulating the immune system in a way that reduces gene therapy
effectiveness is possible. The immune system's enhanced response to viruses that it has seen before reduces the effectiveness to repeated treatments.
- Problems with viral vectors – Viral vectors carry the risks of
toxicity, inflammatory responses, and gene control and targeting issues.
- Multigene disorders – Some commonly occurring disorders, such as heart disease, high blood pressure, Alzheimer's disease, arthritis, and diabetes, are affected by variations in multiple genes, which complicate gene therapy.
- Some therapies may breach the Weismann barrier
(between soma and germ-line) protecting the testes, potentially
modifying the germline, falling afoul of regulations in countries that
prohibit the latter practice.[45]
- Insertional mutagenesis – If the DNA is integrated in a sensitive spot in the genome, for example in a tumor suppressor gene, the therapy could induce a tumor. This has occurred in clinical trials for X-linked severe combined immunodeficiency (X-SCID) patients, in which hematopoietic stem cells were transduced with a corrective transgene using a retrovirus, and this led to the development of T cell leukemia in 3 of 20 patients.[46][47]
One possible solution is to add a functional tumor suppressor gene to
the DNA to be integrated. This may be problematic since the longer the
DNA is, the harder it is to integrate into cell genomes. CRISPR technology allows researchers to make much more precise genome changes at exact locations.[48]
- Cost – Alipogene tiparvovec or Glybera, for example, at a cost of $1.6 million per patient, was reported in 2013 to be the world's most expensive drug.[49][50]
Deaths
Three patients' deaths have been reported in gene therapy trials, putting the field under close scrutiny. The first was that of
Jesse Gelsinger in 1999. Jesse Gelsinger died because of immune rejection response.
[51] One X-SCID patient died of leukemia in 2003.
[9] In 2007, a
rheumatoid arthritis patient died from an infection; the subsequent investigation concluded that the death was not related to gene therapy.
[52]
History
1970s and earlier
In 1972 Friedmann and Roblin authored a paper in
Science titled "Gene therapy for human genetic disease?"
[53] Rogers (1970) was cited for proposing that
exogenous good DNA be used to replace the defective DNA in those who suffer from genetic defects.
[54]
1980s
In 1984 a retrovirus vector system was designed that could efficiently insert foreign genes into mammalian chromosomes.
[55]
1990s
The first approved gene therapy clinical research in the US took place on 14 September 1990, at the
National Institutes of Health (NIH), under the direction of
William French Anderson.
[56] Four-year-old Ashanti DeSilva received treatment for a genetic defect that left her with
ADA-
SCID,
a severe immune system deficiency. The defective gene of the patient's
blood cells was replaced by the functional variant. Ashanti’s immune
system was partially restored by the therapy. Production of the missing
enzyme was temporarily stimulated, but the new cells with functional
genes were not generated. She led a normal life only with the regular
injections performed every two months. The effects were successful, but
temporary.
[57]
Cancer gene therapy was introduced in 1992/93 (Trojan et al. 1993).
[58]
The treatment of glioblastoma multiforme, the malignant brain tumor
whose outcome is always fatal, was done using a vector expressing
antisense IGF-I RNA (clinical trial approved by NIH protocolno.1602
November 24, 1993,
[59]
and by the FDA in 1994). This therapy also represents the beginning of
cancer immunogene therapy, a treatment which proves to be effective due
to the anti-tumor mechanism of IGF-I antisense, which is related to
strong immune and apoptotic phenomena.
In 1992
Claudio Bordignon, working at the
Vita-Salute San Raffaele University, performed the first gene therapy procedure using
hematopoietic stem cells as vectors to deliver genes intended to correct
hereditary diseases.
[60] In 2002 this work led to the publication of the first successful gene therapy treatment for
adenosine deaminase deficiency (ADA-SCID). The success of a multi-center trial for treating children with SCID (
severe combined immune deficiency
or "bubble boy" disease) from 2000 and 2002, was questioned when two of
the ten children treated at the trial's Paris center developed a
leukemia-like condition. Clinical trials were halted temporarily in
2002, but resumed after regulatory review of the protocol in the US, the
United Kingdom, France, Italy, and Germany.
[61]
In 1993 Andrew Gobea was born with SCID following prenatal
genetic screening. Blood was removed from his mother's
placenta and
umbilical cord immediately after birth, to acquire stem cells. The
allele that codes for
adenosine deaminase
(ADA) was obtained and inserted into a retrovirus. Retroviruses and
stem cells were mixed, after which the viruses inserted the gene into
the stem cell chromosomes. Stem cells containing the working ADA gene
were injected into Andrew's blood. Injections of the ADA enzyme were
also given weekly. For four years
T cells (white blood cells), produced by stem cells, made ADA enzymes using the ADA gene. After four years more treatment was needed.
[62]
Jesse Gelsinger's death in 1999 impeded gene therapy research in the US.
[63][64] As a result, the FDA suspended several clinical trials pending the reevaluation of ethical and procedural practices.
[65]
2000s
The modified cancer gene therapy strategy of antisense IGF-I RNA (NIH n˚ 1602)
[59]
using antisense / triple helix anti-IGF-I approach was registered in
2002 by Wiley gene therapy clinical trial - n˚ 635 and 636. The approach
has shown promising results in the treatment of six different malignant
tumors: glioblastoma, cancers of liver, colon, prostate, uterus, and
ovary (Collaborative NATO Science Programme on Gene Therapy USA, France,
Poland n˚ LST 980517 conducted by J. Trojan) (Trojan et al., 2012).
This anti-gene antisense/triple helix therapy has proven to be
efficient, due to the mechanism stopping simultaneously IGF-I expression
on translation and transcription levels, strengthening anti-tumor
immune and apoptotic phenomena.
2002
Sickle-cell disease can be treated in mice.
[66] The mice – which have essentially the same defect that causes human cases – used a viral vector to induce production of
fetal hemoglobin (HbF), which normally ceases to be produced shortly after birth. In humans, the use of
hydroxyurea
to stimulate the production of HbF temporarily alleviates sickle cell
symptoms. The researchers demonstrated this treatment to be a more
permanent means to increase therapeutic HbF production.
[67]
A new gene therapy approach repaired errors in
messenger RNA derived from defective genes. This technique has the potential to treat
thalassaemia,
cystic fibrosis and some cancers.
[68]
Researchers created
liposomes 25 nanometers across that can carry therapeutic DNA through pores in the
nuclear membrane.
[69]
2003
In 2003 a research team inserted genes into the brain for the first time. They used
liposomes coated in a
polymer called
polyethylene glycol, which unlike viral vectors, are small enough to cross the
blood–brain barrier.
[70]
Short pieces of
double-stranded RNA (short, interfering RNAs or
siRNAs)
are used by cells to degrade RNA of a particular sequence. If a siRNA
is designed to match the RNA copied from a faulty gene, then the
abnormal protein product of that gene will not be produced.
[71]
Gendicine is a cancer gene therapy that delivers the
tumor suppressor gene
p53 using an engineered
adenovirus. In 2003, it was approved in China for the treatment of
head and neck squamous cell carcinoma.
[25]
2006
In March researchers announced the successful use of gene therapy to treat two adult patients for X-linked
chronic granulomatous disease, a disease which affects
myeloid cells and damages the
immune system. The study is the first to show that gene therapy can treat the
myeloid system.
[72]
In May a team reported a way to prevent the immune system from rejecting a newly delivered gene.
[73] Similar to
organ transplantation, gene therapy has been plagued by this problem. The
immune system
normally recognizes the new gene as foreign and rejects the cells
carrying it. The research utilized a newly uncovered network of genes
regulated by molecules known as
microRNAs. This natural function selectively obscured their therapeutic gene in
immune system cells and protected it from discovery. Mice infected with
the gene containing an immune-cell microRNA target sequence did not
reject the gene.
In August scientists successfully treated metastatic
melanoma in two patients using
killer T cells genetically retargeted to attack the cancer cells.
[74]
In November researchers reported on the use of VRX496, a gene-based
immunotherapy for the treatment of
HIV that uses a
lentiviral vector to deliver an
antisense gene against the
HIV envelope. In a
phase I clinical trial, five subjects with chronic HIV infection who had failed to respond to at least two
antiretroviral regimens were treated. A single intravenous infusion of
autologous CD4
T cells genetically modified with VRX496 was well tolerated. All
patients had stable or decreased viral load; four of the five patients
had stable or increased CD4 T cell counts. All five patients had stable
or increased immune response to HIV
antigens and other
pathogens. This was the first evaluation of a lentiviral vector administered in a US human clinical trial.
[75][76]
2007
In May researchers announced the first gene therapy trial for inherited
retinal disease. The first operation was carried out on a 23-year-old British male, Robert Johnson, in early 2007.
[77]
2008
Leber's congenital amaurosis is an inherited blinding disease caused by mutations in the
RPE65 gene. The results of a small clinical trial in children were published in April.
[11] Delivery of recombinant
adeno-associated virus
(AAV) carrying RPE65 yielded positive results. In May two more groups
reported positive results in independent clinical trials using gene
therapy to treat the condition. In all three clinical trials, patients
recovered functional vision without apparent side-effects.
[11][12][13][14]
2009
In September researchers were able to give
trichromatic vision to
squirrel monkeys.
[78] In November 2009, researchers halted a fatal
genetic disorder called
adrenoleukodystrophy in two children using a
lentivirus vector to deliver a functioning version of
ABCD1, the gene that is mutated in the disorder.
[79]
2010s
2010
An April paper reported that gene therapy addressed
achromatopsia (color blindness) in dogs by targeting
cone
photoreceptors. Cone function and day vision were restored for at least
33 months in two young specimens. The therapy was less efficient for
older dogs.
[80]
In September it was announced that an 18-year-old male patient in France with
beta-thalassemia major had been successfully treated.
[81] Beta-thalassemia major is an inherited
blood disease in which
beta haemoglobin is missing and patients are dependent on regular lifelong
blood transfusions.
[82] The technique used a lentiviral vector to transduce the human ß-globin gene into purified blood and
marrow cells obtained from the patient in June 2007.
[83]
The patient's haemoglobin levels were stable at 9 to 10 g/dL. About a
third of the hemoglobin contained the form introduced by the viral
vector and blood transfusions were not needed.
[83][84] Further clinical trials were planned.
[85] Bone marrow transplants are the only cure for thalassemia, but 75% of patients do not find a matching donor.
[84]
Cancer immunogene therapy using modified antigene,
antisense/triple helix approach was introduced in South America in
2010/11 in La Sabana University, Bogota (Ethical Committee 14 December
2010, no P-004-10). Considering the ethical aspect of gene diagnostic
and gene therapy targeting IGF-I, the IGF-I expressing tumors i.e. lung
and epidermis cancers were treated (Trojan et al. 2016).
2011
In 2007 and 2008, a man (
Timothy Ray Brown) was cured of HIV by repeated
hematopoietic stem cell transplantation (see also
allogeneic stem cell transplantation,
allogeneic bone marrow transplantation,
allotransplantation) with double-delta-32 mutation which disables the
CCR5 receptor. This cure was accepted by the medical community in 2011.
[88] It required complete
ablation of existing
bone marrow, which is very debilitating.
In August two of three subjects of a pilot study were confirmed to have been cured from
chronic lymphocytic leukemia (CLL). The therapy used genetically modified
T cells to attack cells that expressed the
CD19 protein to fight the disease.
[20]
In 2013, the researchers announced that 26 of 59 patients had achieved
complete remission and the original patient had remained tumor-free.
[89]
Human HGF plasmid DNA therapy of
cardiomyocytes is being examined as a potential treatment for
coronary artery disease as well as treatment for the damage that occurs to the heart after
myocardial infarction.
[90][91]
In 2011
Neovasculgen was registered in Russia as the first-in-class gene-therapy drug for treatment of
peripheral artery disease, including
critical limb ischemia; it delivers the gene encoding for
VEGF.
[92][26] Neovasculogen is a
plasmid encoding the
CMV promoter and the 165 amino acid form of
VEGF.
[93][94]
2012
The FDA approved Phase 1 clinical trials on
thalassemia major patients in the US for 10 participants in July.
[95] The study was expected to continue until 2015.
[85]
In July 2012, the
European Medicines Agency recommended approval of a gene therapy treatment for the first time in either Europe or the United States. The treatment used
Alipogene tiparvovec (Glybera) to compensate for
lipoprotein lipase deficiency, which can cause severe
pancreatitis.
[96] The recommendation was endorsed by the
European Commission in November 2012
[10][27][97][98] and commercial rollout began in late 2014.
[99] Alipogene tiparvovec was expected to cost around $1.6 million per treatment in 2012,
[100] revised to $1 million in 2015,
[101] making it the most expensive medicine in the world at the time.
[102] As of 2016, only one person had been treated with drug.
[103]
In December 2012, it was reported that 10 of 13 patients with
multiple myeloma were in remission "or very close to it" three months after being injected with a treatment involving genetically engineered
T cells to target proteins
NY-ESO-1 and
LAGE-1, which exist only on cancerous myeloma cells.
[22]
2013
In March researchers reported that three of five adult subjects who had
acute lymphocytic leukemia (ALL) had been in remission for five months to two years after being treated with genetically modified
T cells which attacked cells with
CD19 genes on their surface, i.e. all
B-cells,
cancerous or not. The researchers believed that the patients' immune
systems would make normal T-cells and B-cells after a couple of months.
They were also given bone marrow. One patient relapsed and died and one
died of a blood clot unrelated to the disease.
[21]
Following encouraging Phase 1 trials, in April, researchers
announced they were starting Phase 2 clinical trials (called CUPID2 and
SERCA-LVAD) on 250 patients
[104] at several hospitals to combat
heart disease. The therapy was designed to increase the levels of
SERCA2, a protein in heart muscles, improving muscle function.
[105] The
FDA granted this a
Breakthrough Therapy Designation to accelerate the trial and approval process.
[106] In 2016 it was reported that no improvement was found from the CUPID 2 trial.
[107]
In July researchers reported promising results for six children
with two severe hereditary diseases had been treated with a partially
deactivated lentivirus to replace a faulty gene and after 7–32 months. Three of the children had
metachromatic leukodystrophy, which causes children to lose cognitive and motor skills.
[108] The other children had
Wiskott-Aldrich syndrome, which leaves them to open to infection, autoimmune diseases, and cancer.
[109] Follow up trials with gene therapy on another six children with Wiskott-Aldrich syndrome were also reported as promising.
[110][111]
In October researchers reported that two children born with
adenosine deaminase severe combined immunodeficiency disease (
ADA-SCID)
had been treated with genetically engineered stem cells 18 months
previously and that their immune systems were showing signs of full
recovery. Another three children were making progress.
[18] In 2014 a further 18 children with ADA-SCID were cured by gene therapy.
[112] ADA-SCID children have no functioning immune system and are sometimes known as "bubble children."
[18]
Also in October researchers reported that they had treated six
hemophilia sufferers in early 2011 using an adeno-associated virus.
Over two years later all six were producing
clotting factor.
[18][113]
2014
In January researchers reported that six
choroideremia patients had been treated with adeno-associated virus with a copy of
REP1. Over a six-month to two-year period all had improved their sight.
[114][115] By 2016, 32 patients had been treated with positive results and researchers were hopeful the treatment would be long-lasting.
[15] Choroideremia is an inherited genetic eye disease with no approved treatment, leading to loss of sight.
In March researchers reported that 12 HIV patients had been
treated since 2009 in a trial with a genetically engineered virus with a
rare mutation (
CCR5 deficiency) known to protect against HIV with promising results.
[116][117]
Clinical trials of gene therapy for
sickle cell disease were started in 2014.
[118][119] There is a need for high quality
randomised controlled trials assessing the risks and benefits involved with gene therapy for people with sickle cell disease.
[120]
2015
In February
LentiGlobin BB305, a gene therapy treatment undergoing clinical trials for treatment of
beta thalassemia
gained FDA "breakthrough" status after several patients were able to
forgo the frequent blood transfusions usually required to treat the
disease.
[121]
In March researchers delivered a
recombinant gene encoding a
broadly neutralizing antibody into monkeys infected with simian
HIV; the monkeys' cells produced the
antibody,
which cleared them of HIV. The technique is named immunoprophylaxis by
gene transfer (IGT). Animal tests for antibodies to ebola, malaria,
influenza, and hepatitis were underway.
[122][123]
In March, scientists, including an inventor of
CRISPR,
Jennifer Doudna,
urged a worldwide moratorium on germline gene therapy, writing
"scientists should avoid even attempting, in lax jurisdictions, germline
genome modification for clinical application in humans" until the full
implications "are discussed among scientific and governmental
organizations".
[124][125][126][127]
In October, researchers announced that they had treated a baby
girl, Layla Richards, with an experimental treatment using donor T-cells
genetically engineered using
TALEN to attack cancer cells. One year after the treatment she was still free of her cancer (a highly aggressive form of
acute lymphoblastic leukaemia [ALL]).
[128]
Children with highly aggressive ALL normally have a very poor prognosis
and Layla's disease had been regarded as terminal before the treatment.
[129]
In December, scientists of major world academies called for a moratorium on inheritable
human genome edits, including those related to
CRISPR-Cas9 technologies
[130] but that basic research including embryo gene editing should continue.
[131]
2016
In April the
Committee for Medicinal Products for Human Use of the
European Medicines Agency endorsed a gene therapy treatment called
Strimvelis[132][133] and the European Commission approved it in June.
[134] This treats children born with
adenosine deaminase deficiency and who have no functioning immune system. This was the second gene therapy treatment to be approved in Europe.
[135]
In October, Chinese scientists reported they had started a trial
to genetically modify T-cells from 10 adult patients with lung cancer
and reinject the modified T-cells back into their bodies to attack the
cancer cells. The T-cells had the
PD-1 protein (which stops or slows the immune response) removed using CRISPR-Cas9.
[136][137]
A 2016
Cochrane systematic review looking at data from four trials on
topical cystic fibrosis transmembrane conductance regulator
(CFTR) gene therapy does not support its clinical use as a mist inhaled
into the lungs to treat cystic fibrosis patients with lung infections.
One of the four trials did find weak evidence that liposome-based CFTR
gene transfer therapy may lead to a small respiratory improvement for
people with CF. This weak evidence is not enough to make a clinical
recommendation for routine CFTR gene therapy.
[138]
2017
In February
Kite Pharma announced results from a clinical trial of
CAR-T cells in around a hundred people with advanced
Non-Hodgkin lymphoma.
[139]
In March, French scientists reported on clinical research of gene therapy to treat
sickle-cell disease.
[140]
In August, the FDA approved
tisagenlecleucel for acute lymphoblastic leukemia.
[141] Tisagenlecleucel is an
adoptive cell transfer therapy for
B-cell acute lymphoblastic leukemia;
T cells from a person with cancer are removed,
genetically engineered to make a specific
T-cell receptor
(a chimeric T cell receptor, or "CAR-T") that reacts to the cancer, and
are administered back to the person. The T cells are engineered to
target a protein called
CD19
that is common on B cells. This is the first form of gene therapy to be
approved in the United States. In October, a similar therapy called
axicabtagene ciloleucel was approved for
non-Hodgkin lymphoma.
[142]
In December the results of using an adeno-associated virus with blood clotting
factor VIII to treat nine
haemophilia A
patients were published. Six of the seven patients on the high dose
regime increased the level of the blood clotting VIII to normal levels.
The low and medium dose regimes had no effect on the patient's blood
clotting levels.
[143][144]
In December, the FDA approved
Luxturna, the first
in vivo gene therapy, for the treatment of blindness due to
Leber's congenital amaurosis.
[145] The price of this treatment was 850,000 US dollars for both eyes.
[146][147]
Speculative uses
Speculated uses for gene therapy include:
Fertility
Gene
Therapy techniques have the potential to provide alternative treatments
for those with infertility. Recently, successful experimentation on
mice has proven that fertility can be restored by using the gene therapy
method, CRISPR.
[148]
Spermatogenical stem cells from another organism were transplanted into
the testes of an infertile male mouse. The stem cells re-established
spermatogenesis and fertility.
[149]
Gene doping
Athletes might adopt gene therapy technologies to improve their performance.
[150] Gene doping is not known to occur, but multiple gene therapies may have such effects. Kayser et al. argue that gene doping could
level the playing field
if all athletes receive equal access. Critics claim that any
therapeutic intervention for non-therapeutic/enhancement purposes
compromises the ethical foundations of medicine and sports.
[151]
Human genetic engineering
Genetic engineering could be used to cure diseases, but also to change physical appearance,
metabolism, and even
improve physical capabilities and mental faculties such as
memory and
intelligence. Ethical claims about germline engineering include beliefs that every
fetus
has a right to remain genetically unmodified, that parents hold the
right to genetically modify their offspring, and that every child has
the right to be born free of preventable diseases.
[152][153][154]
For parents, genetic engineering could be seen as another child
enhancement technique to add to diet, exercise, education, training,
cosmetics, and plastic surgery.
[155][156] Another theorist claims that moral concerns limit but do not prohibit germline engineering.
[157]
Possible regulatory schemes include a complete ban, provision to everyone, or professional self-regulation. The
American Medical Association’s
Council on Ethical and Judicial Affairs stated that "genetic
interventions to enhance traits should be considered permissible only in
severely restricted situations: (1) clear and meaningful benefits to
the fetus or child; (2) no trade-off with other characteristics or
traits; and (3) equal access to the genetic technology, irrespective of
income or other socioeconomic characteristics."
[158]
As early in the history of
biotechnology as 1990, there have been scientists opposed to attempts to modify the human
germline using these new tools,
[159] and such concerns have continued as technology progressed.
[160][161] With the advent of new techniques like
CRISPR, in March 2015 a group of scientists urged a worldwide moratorium on clinical use of gene editing technologies to edit the
human genome in a way that can be inherited.
[124][125][126][127] In April 2015, researchers sparked controversy when they
reported results of
basic research to edit the
DNA of non-viable
human embryos using CRISPR.
[148][162] A committee of the American
National Academy of Sciences and
National Academy of Medicine gave qualified support to human genome editing in 2017
[163][164] once answers have been found to safety and efficiency problems "but only for serious conditions under stringent oversight."
[165]
Regulations
Regulations
covering genetic modification are part of general guidelines about
human-involved biomedical research. There are no international treaties
which are legally binding in this area, but there are recommendations
for national laws from various bodies.
The
Helsinki Declaration (Ethical Principles for Medical Research Involving Human Subjects) was amended by the
World Medical Association's
General Assembly in 2008. This document provides principles physicians
and researchers must consider when involving humans as research
subjects. The Statement on Gene Therapy Research initiated by the
Human Genome Organization
(HUGO) in 2001 provides a legal baseline for all countries. HUGO’s
document emphasizes human freedom and adherence to human rights, and
offers recommendations for somatic gene therapy, including the
importance of recognizing public concerns about such research.
[166]
United States
No
federal legislation lays out protocols or restrictions about human
genetic engineering. This subject is governed by overlapping regulations
from local and federal agencies, including the
Department of Health and Human Services,
the FDA and NIH's Recombinant DNA Advisory Committee. Researchers
seeking federal funds for an investigational new drug application,
(commonly the case for somatic human genetic engineering,) must obey
international and federal guidelines for the protection of human
subjects.
[167]
NIH serves as the main gene therapy regulator for federally
funded research. Privately funded research is advised to follow these
regulations. NIH provides funding for research that develops or enhances
genetic engineering techniques and to evaluate the ethics and quality
in current research. The NIH maintains a mandatory registry of human
genetic engineering research protocols that includes all federally
funded projects.
An NIH advisory committee published a set of guidelines on gene manipulation.
[168]
The guidelines discuss lab safety as well as human test subjects and
various experimental types that involve genetic changes. Several
sections specifically pertain to human genetic engineering, including
Section III-C-1. This section describes required review processes and
other aspects when seeking approval to begin clinical research involving
genetic transfer into a human patient.
[169]
The protocol for a gene therapy clinical trial must be approved by the
NIH's Recombinant DNA Advisory Committee prior to any clinical trial
beginning; this is different from any other kind of clinical trial.
[168]
As with other kinds of drugs, the FDA regulates the quality and
safety of gene therapy products and supervises how these products are
used clinically. Therapeutic alteration of the human genome falls under
the same regulatory requirements as any other medical treatment.
Research involving human subjects, such as
clinical trials, must be reviewed and approved by the FDA and an
Institutional Review Board.
[170][171]
Popular culture
Gene therapy is the basis for the plotline of the film
I Am Legend[172] and the TV show
Will Gene Therapy Change the Human Race?.
[173] In 1994, gene therapy was a plot element in
The Erlenmeyer Flask, The X-Files' first season finale. It is also used in
Stargate as a means of allowing humans to use Ancient technology.