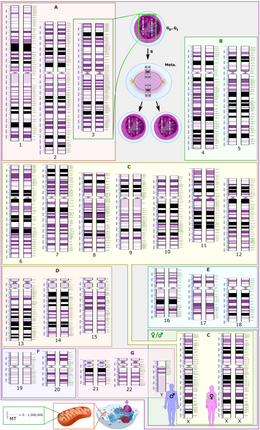
Schematic representation of the human diploid karyotype, showing the organization of the genome into chromosomes, as well as annotated bands and sub-bands as seen on G banding. This drawing shows both the female (XX) and male (XY) versions of the 23rd chromosome pair. Chromosomal changes during the cell cycle are displayed at top center. The mitochondrial genome is shown to scale at bottom left. | |
| NCBI genome ID | 51 |
|---|---|
| Ploidy | diploid |
| Genome size | 3,117,275,501 base pairs (bp) |
| Number of chromosomes | 23 pairs |
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly repetitive sequences. Introns make up a large percentage of non-coding DNA. Some of this non-coding DNA is non-functional junk DNA, such as pseudogenes, but there is no firm consensus on the total amount of junk DNA.
Haploid human genomes, which are contained in germ cells (the egg and sperm gamete cells created in the meiosis phase of sexual reproduction before fertilization) consist of 3,054,815,472 DNA base pairs (if X chromosome is used), while female diploid genomes (found in somatic cells) have twice the DNA content.
While there are significant differences among the genomes of human individuals (on the order of 0.1% due to single-nucleotide variants and 0.6% when considering indels), these are considerably smaller than the differences between humans and their closest living relatives, the bonobos and chimpanzees (~1.1% fixed single-nucleotide variants and 4% when including indels). Size in basepairs can vary too; the telomere length decreases after every round of DNA replication.
Although the sequence of the human genome has been completely determined by DNA sequencing in 2022 (including methylation), it is not yet fully understood. Most, but not all, genes have been identified by a combination of high throughput experimental and bioinformatics approaches, yet much work still needs to be done to further elucidate the biological functions of their protein and RNA products (in particular, annotation of the complete CHM13v2.0 sequence is still ongoing). And yet, overlapping genes are quite common, in some cases allowing two protein coding genes from each strand to reuse base pairs twice (for example, genes DCDC2 and KAAG1). Recent results suggest that most of the vast quantities of noncoding DNA within the genome have associated biochemical activities, including regulation of gene expression, organization of chromosome architecture, and signals controlling epigenetic inheritance. There are also a significant number of retroviruses in human DNA, at least 3 of which have been proven to possess an important function (i.e., HIV-like HERV-K, HERV-W, and HERV-FRD play a role in placenta formation by inducing cell-cell fusion).
In 2003, scientists reported the sequencing of 85% of the entire human genome, but as of 2020 at least 8% was still missing. In 2021, scientists reported sequencing the complete female genome (i.e., without the Y chromosome). This sequence identified 19,969 protein-coding sequences, accounting for approximately 1.5% of the genome, and 63,494 genes in total, most of them being non-coding RNA genes. The genome consists of regulatory DNA sequences, LINEs, SINEs, introns, and sequences for which as yet no function has been determined. The human Y chromosome, consisting of 62,460,029 base pairs from a different cell line and found in all males, was sequenced completely in January 2022.
In 2023, a draft human pangenome reference was published. It is based on 47 genomes from persons of varied ethnicity. Plans are underway for an improved reference capturing still more biodiversity from a still wider sample.
Sequencing
The first human genome sequences were published in nearly complete draft form in February 2001 by the Human Genome Project and Celera Corporation. Completion of the Human Genome Project's sequencing effort was announced in 2004 with the publication of a draft genome sequence, leaving just 341 gaps in the sequence, representing highly repetitive and other DNA that could not be sequenced with the technology available at the time. The human genome was the first of all vertebrates to be sequenced to such near-completion, and as of 2018, the diploid genomes of over a million individual humans had been determined using next-generation sequencing.
These data are used worldwide in biomedical science, anthropology, forensics and other branches of science. Such genomic studies have led to advances in the diagnosis and treatment of diseases, and to new insights in many fields of biology, including human evolution.
By 2018, the total number of genes had been raised to at least 46,831, plus another 2300 micro-RNA genes. A 2018 population survey found another 300 million bases of human genome that was not in the reference sequence. Prior to the acquisition of the full genome sequence, estimates of the number of human genes ranged from 50,000 to 140,000 (with occasional vagueness about whether these estimates included non-protein coding genes). As genome sequence quality and the methods for identifying protein-coding genes improved, the count of recognized protein-coding genes dropped to 19,000–20,000.
In June 2016, scientists formally announced HGP-Write, a plan to synthesize the human genome.
In 2022 the Telomere-to-Telomere (T2T) consortium reported the complete sequence of a human female genome, filling all the gaps in the X chromosome (2020) and the 22 autosomes (May 2021). The previously unsequenced parts contain immune response genes that help to adapt to and survive infections, as well as genes that are important for predicting drug response. The completed human genome sequence will also provide better understanding of human formation as an individual organism and how humans vary both between each other and other species.
Achieving completeness
Although the 'completion' of the human genome project was announced in 2001, there remained hundreds of gaps, with about 5–10% of the total sequence remaining undetermined. The missing genetic information was mostly in repetitive heterochromatic regions and near the centromeres and telomeres, but also some gene-encoding euchromatic regions. There remained 160 euchromatic gaps in 2015 when the sequences spanning another 50 formerly unsequenced regions were determined. Only in 2020 was the first truly complete telomere-to-telomere sequence of a human chromosome determined, namely of the X chromosome. The first complete telomere-to-telomere sequence of a human autosomal chromosome, chromosome 8, followed a year later. The complete human genome (without Y chromosome) was published in 2021, while with Y chromosome in January 2022.
In 2023, a draft human pangenome reference was published. It is based on 47 genomes from persons of varied ethnicity. Plans are underway for an improved reference capturing still more biodiversity from a still wider sample.
Molecular organization and gene content
The total length of the human reference genome does not represent the sequence of any specific individual. The genome is organized into 22 paired chromosomes, termed autosomes, plus the 23rd pair of sex chromosomes (XX) in the female and (XY) in the male. The haploid genome is 3 054 815 472 base pairs, when the X chromosome is included, and 2 963 015 935 base pairs when the Y chromosome is substituted for the X chromosome. These chromosomes are all large linear DNA molecules contained within the cell nucleus. The genome also includes the mitochondrial DNA, a comparatively small circular molecule present in multiple copies in each mitochondrion.
| Chromo- some |
Length | Base pairs |
Varia- tions |
Protein- coding genes |
Pseudo- genes |
Total long ncRNA |
Total small ncRNA |
miRNA | rRNA | snRNA | snoRNA | Misc ncRNA |
Links | Centromere position (Mbp) |
Cumu- lative (%) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 8.5 cm | 248,387,328 | 12,151,146 | 2058 | 1220 | 1200 | 496 | 134 | 66 | 221 | 145 | 192 | EBI | 125 | 7.9 |
| 2 | 8.3 cm | 242,696,752 | 12,945,965 | 1309 | 1023 | 1037 | 375 | 115 | 40 | 161 | 117 | 176 | EBI | 93.3 | 16.2 |
| 3 | 6.7 cm | 201,105,948 | 10,638,715 | 1078 | 763 | 711 | 298 | 99 | 29 | 138 | 87 | 134 | EBI | 91 | 23 |
| 4 | 6.5 cm | 193,574,945 | 10,165,685 | 752 | 727 | 657 | 228 | 92 | 24 | 120 | 56 | 104 | EBI | 50.4 | 29.6 |
| 5 | 6.2 cm | 182,045,439 | 9,519,995 | 876 | 721 | 844 | 235 | 83 | 25 | 106 | 61 | 119 | EBI | 48.4 | 35.8 |
| 6 | 5.8 cm | 172,126,628 | 9,130,476 | 1048 | 801 | 639 | 234 | 81 | 26 | 111 | 73 | 105 | EBI | 61 | 41.6 |
| 7 | 5.4 cm | 160,567,428 | 8,613,298 | 989 | 885 | 605 | 208 | 90 | 24 | 90 | 76 | 143 | EBI | 59.9 | 47.1 |
| 8 | 5.0 cm | 146,259,331 | 8,221,520 | 677 | 613 | 735 | 214 | 80 | 28 | 86 | 52 | 82 | EBI | 45.6 | 52 |
| 9 | 4.8 cm | 150,617,247 | 6,590,811 | 786 | 661 | 491 | 190 | 69 | 19 | 66 | 51 | 96 | EBI | 49 | 56.3 |
| 10 | 4.6 cm | 134,758,134 | 7,223,944 | 733 | 568 | 579 | 204 | 64 | 32 | 87 | 56 | 89 | EBI | 40.2 | 60.9 |
| 11 | 4.6 cm | 135,127,769 | 7,535,370 | 1298 | 821 | 710 | 233 | 63 | 24 | 74 | 76 | 97 | EBI | 53.7 | 65.4 |
| 12 | 4.5 cm | 133,324,548 | 7,228,129 | 1034 | 617 | 848 | 227 | 72 | 27 | 106 | 62 | 115 | EBI | 35.8 | 70 |
| 13 | 3.9 cm | 113,566,686 | 5,082,574 | 327 | 372 | 397 | 104 | 42 | 16 | 45 | 34 | 75 | EBI | 17.9 | 73.4 |
| 14 | 3.6 cm | 101,161,492 | 4,865,950 | 830 | 523 | 533 | 239 | 92 | 10 | 65 | 97 | 79 | EBI | 17.6 | 76.4 |
| 15 | 3.5 cm | 99,753,195 | 4,515,076 | 613 | 510 | 639 | 250 | 78 | 13 | 63 | 136 | 93 | EBI | 19 | 79.3 |
| 16 | 3.1 cm | 96,330,374 | 5,101,702 | 873 | 465 | 799 | 187 | 52 | 32 | 53 | 58 | 51 | EBI | 36.6 | 82 |
| 17 | 2.8 cm | 84,276,897 | 4,614,972 | 1197 | 531 | 834 | 235 | 61 | 15 | 80 | 71 | 99 | EBI | 24 | 84.8 |
| 18 | 2.7 cm | 80,542,538 | 4,035,966 | 270 | 247 | 453 | 109 | 32 | 13 | 51 | 36 | 41 | EBI | 17.2 | 87.4 |
| 19 | 2.0 cm | 61,707,364 | 3,858,269 | 1472 | 512 | 628 | 179 | 110 | 13 | 29 | 31 | 61 | EBI | 26.5 | 89.3 |
| 20 | 2.1 cm | 66,210,255 | 3,439,621 | 544 | 249 | 384 | 131 | 57 | 15 | 46 | 37 | 68 | EBI | 27.5 | 91.4 |
| 21 | 1.6 cm | 45,090,682 | 2,049,697 | 234 | 185 | 305 | 71 | 16 | 5 | 21 | 19 | 24 | EBI | 13.2 | 92.6 |
| 22 | 1.7 cm | 51,324,926 | 2,135,311 | 488 | 324 | 357 | 78 | 31 | 5 | 23 | 23 | 62 | EBI | 14.7 | 93.8 |
| X | 5.3 cm | 154,259,566 | 5,753,881 | 842 | 874 | 271 | 258 | 128 | 22 | 85 | 64 | 100 | EBI | 60.6 | 99.1 |
| Y | 2.0 cm | 62,460,029 | 211,643 | 71 | 388 | 71 | 30 | 15 | 7 | 17 | 3 | 8 | EBI | 10.4 | 100 |
| mtDNA | 5.4 μm | 16,569 | 929 | 13 | 0 | 0 | 24 | 0 | 2 | 0 | 0 | 0 | EBI | N/A | 100 |
|
| |||||||||||||||
| hapl 1-23 + X | 104 cm | 3,054,815,472 | 20328 | 14212 | 14656 | 4983 | 1741 | 523 | 1927 | 1518 | 2205 |
| |||
| hapl 1-23 + Y | 101 cm | 2,963,015,935 | 19557 | 13726 | 14456 | 4755 | 1628 | 508 | 1859 | 1457 | 2113 |
| |||
dipl + mt♀ |
208.23 cm | 6,109,647,513 | 40669 | 28424 | 29312 | 9990 | 3482 | 1048 | 3854 | 3036 | 4410 |
| |||
dipl + mt♂ |
205.00 cm | 6,017,847,976 | 39898 | 27938 | 29112 | 9762 | 3369 | 1033 | 3786 | 2975 | 4318 |
| |||
Variations are unique DNA sequence differences that have been identified in the individual human genome sequences analyzed by Ensembl as of December 2016. The number of identified variations is expected to increase as further personal genomes are sequenced and analyzed. In addition to the gene content shown in this table, a large number of non-expressed functional sequences have been identified throughout the human genome (see below). Links open windows to the reference chromosome sequences in the EBI genome browser.
Small non-coding RNAs are RNAs of as many as 200 bases that do not have protein-coding potential. These include: microRNAs, or miRNAs (post-transcriptional regulators of gene expression), small nuclear RNAs, or snRNAs (the RNA components of spliceosomes), and small nucleolar RNAs, or snoRNA (involved in guiding chemical modifications to other RNA molecules). Long non-coding RNAs are RNA molecules longer than 200 bases that do not have protein-coding potential. These include: ribosomal RNAs, or rRNAs (the RNA components of ribosomes), and a variety of other long RNAs that are involved in regulation of gene expression, epigenetic modifications of DNA nucleotides and histone proteins, and regulation of the activity of protein-coding genes. Small discrepancies between total-small-ncRNA numbers and the numbers of specific types of small ncNRAs result from the former values being sourced from Ensembl release 87 and the latter from Ensembl release 68.
The number of genes in the human genome is not entirely clear because the function of numerous transcripts remains unclear. This is especially true for non-coding RNA. The number of protein-coding genes is better known but there are still on the order of 1,400 questionable genes which may or may not encode functional proteins, usually encoded by short open reading frames.|
|
Gencode | Ensembl | Refseq | CHESS |
|---|---|---|---|---|
| protein-coding genes | 19,901 | 20,376 | 20,345 | 21,306 |
| lncRNA genes | 15,779 | 14,720 | 17,712 | 18,484 |
| antisense RNA | 5501 |
|
28 | 2694 |
| miscellaneous RNA | 2213 | 2222 | 13,899 | 4347 |
| Pseudogenes | 14,723 | 1740 | 15,952 |
|
| total transcripts | 203,835 | 203,903 | 154,484 | 328,827 |
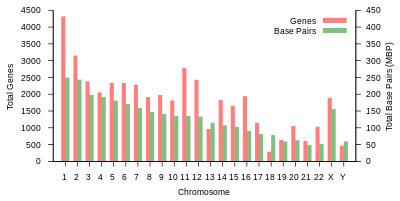
Information content
The haploid human genome (23 chromosomes) is about 3 billion base pairs long and in 2018 was said to contain at least 46,831 genes. In 2022 the number increased again to 63,494 genes. The increase from the previously accepted number of around 20,000 is due to the difficulty of defining what a gene is. It is widely agreed that there are about 20,000 protein-coding genes, with some papers stating exact figures of 21,306. The higher figures include non-protein coding RNA-producing genes that perform other cell functions.
Since every base pair can be coded by 2 bits, this is about 750 megabytes of data. An individual somatic (diploid) cell contains twice this amount, that is, about 6 billion base pairs. Males have fewer than females because the Y chromosome is about 62 million base pairs whereas the X is about 154 million. Since individual genomes vary in sequence by less than 1% from each other, the variations of a given human's genome from a common reference can be losslessly compressed to roughly 4 megabytes.
The entropy rate of the genome differs significantly between coding and non-coding sequences. It is close to the maximum of 2 bits per base pair for the coding sequences (about 45 million base pairs), but less for the non-coding parts. It ranges between 1.5 and 1.9 bits per base pair for the individual chromosome, except for the Y chromosome, which has an entropy rate below 0.9 bits per base pair.
Coding vs. noncoding DNA
The content of the human genome is commonly divided into coding and noncoding DNA sequences. Coding DNA is defined as those sequences that can be transcribed into mRNA and translated into proteins during the human life cycle; these sequences occupy only a small fraction of the genome (<2%). Noncoding DNA is made up of all of those sequences (approx. 98% of the genome) that are not used to encode proteins.
Some noncoding DNA contains genes for RNA molecules with important biological functions (noncoding RNA, for example ribosomal RNA and transfer RNA). The exploration of the function and evolutionary origin of noncoding DNA is an important goal of contemporary genome research, including the ENCODE (Encyclopedia of DNA Elements) project, which aims to survey the entire human genome, using a variety of experimental tools whose results are indicative of molecular activity. It is however disputed whether molecular activity (transcription of DNA into RNA) alone implies that the RNA produced has a meaningful biological function, since experiments have shown that random nonfunctional DNA will also reproducibly recruit transcription factors resulting in transcription into nonfunctional RNA.
There is no consensus on what constitutes a "functional" element in the genome since geneticists, evolutionary biologists, and molecular biologists employ different definitions and methods. Due to the ambiguity in the terminology, different schools of thought have emerged. In evolutionary definitions, "functional" DNA, whether it is coding or non-coding, contributes to the fitness of the organism, and therefore is maintained by negative evolutionary pressure whereas "non-functional" DNA has no benefit to the organism and therefore is under neutral selective pressure. This type of DNA has been described as junk DNA In genetic definitions, "functional" DNA is related to how DNA segments manifest by phenotype and "nonfunctional" is related to loss-of-function effects on the organism. In biochemical definitions, "functional" DNA relates to DNA sequences that specify molecular products (e.g. noncoding RNAs) and biochemical activities with mechanistic roles in gene or genome regulation (i.e. DNA sequences that impact cellular level activity such as cell type, condition, and molecular processes). There is no consensus in the literature on the amount of functional DNA since, depending on how "function" is understood, ranges have been estimated from up to 90% of the human genome is likely nonfunctional DNA (junk DNA) to up to 80% of the genome is likely functional. It is also possible that junk DNA may acquire a function in the future and therefore may play a role in evolution, but this is likely to occur only very rarely. Finally DNA that is deliterious to the organism and is under negative selective pressure is called garbage DNA.
Because non-coding DNA greatly outnumbers coding DNA, the concept of the sequenced genome has become a more focused analytical concept than the classical concept of the DNA-coding gene.
Coding sequences (protein-coding genes)

Protein-coding sequences represent the most widely studied and best understood component of the human genome. These sequences ultimately lead to the production of all human proteins, although several biological processes (e.g. DNA rearrangements and alternative pre-mRNA splicing) can lead to the production of many more unique proteins than the number of protein-coding genes. The complete modular protein-coding capacity of the genome is contained within the exome, and consists of DNA sequences encoded by exons that can be translated into proteins. Because of its biological importance, and the fact that it constitutes less than 2% of the genome, sequencing of the exome was the first major milepost of the Human Genome Project.
Number of protein-coding genes. About 20,000 human proteins have been annotated in databases such as Uniprot. Historically, estimates for the number of protein genes have varied widely, ranging up to 2,000,000 in the late 1960s, but several researchers pointed out in the early 1970s that the estimated mutational load from deleterious mutations placed an upper limit of approximately 40,000 for the total number of functional loci (this includes protein-coding and functional non-coding genes). The number of human protein-coding genes is not significantly larger than that of many less complex organisms, such as the roundworm and the fruit fly. This difference may result from the extensive use of alternative pre-mRNA splicing in humans, which provides the ability to build a very large number of modular proteins through the selective incorporation of exons.
Protein-coding capacity per chromosome. Protein-coding genes are distributed unevenly across the chromosomes, ranging from a few dozen to more than 2000, with an especially high gene density within chromosomes 1, 11, and 19. Each chromosome contains various gene-rich and gene-poor regions, which may be correlated with chromosome bands and GC-content. The significance of these nonrandom patterns of gene density is not well understood.
Size of protein-coding genes. The size of protein-coding genes within the human genome shows enormous variability. For example, the gene for histone H1a (HIST1HIA) is relatively small and simple, lacking introns and encoding an 781 nucleotide-long mRNA that produces a 215 amino acid protein from its 648 nucleotide open reading frame. Dystrophin (DMD) was the largest protein-coding gene in the 2001 human reference genome, spanning a total of 2.2 million nucleotides, while more recent systematic meta-analysis of updated human genome data identified an even larger protein-coding gene, RBFOX1 (RNA binding protein, fox-1 homolog 1), spanning a total of 2.47 million nucleotides. Titin (TTN) has the longest coding sequence (114,414 nucleotides), the largest number of exons (363), and the longest single exon (17,106 nucleotides). As estimated based on a curated set of protein-coding genes over the whole genome, the median size is 26,288 nucleotides (mean = 66,577), the median exon size, 133 nucleotides (mean = 309), the median number of exons, 8 (mean = 11), and the median encoded protein is 425 amino acids (mean = 553) in length.
| Protein | Chrom | Gene | Length | Exons | Exon length | Intron length | Alt splicing |
|---|---|---|---|---|---|---|---|
| Breast cancer type 2 susceptibility protein | 13 | BRCA2 | 83,736 | 27 | 11,386 | 72,350 | yes |
| Cystic fibrosis transmembrane conductance regulator | 7 | CFTR | 202,881 | 27 | 4,440 | 198,441 | yes |
| Cytochrome b | MT | MTCYB | 1,140 | 1 | 1,140 | 0 | no |
| Dystrophin | X | DMD | 2,220,381 | 79 | 10,500 | 2,209,881 | yes |
| Glyceraldehyde-3-phosphate dehydrogenase | 12 | GAPDH | 4,444 | 9 | 1,425 | 3,019 | yes |
| Hemoglobin beta subunit | 11 | HBB | 1,605 | 3 | 626 | 979 | no |
| Histone H1A | 6 | HIST1H1A | 781 | 1 | 781 | 0 | no |
| Titin | 2 | TTN | 281,434 | 364 | 104,301 | 177,133 | yes |
Noncoding DNA (ncDNA)
Noncoding DNA is defined as all of the DNA sequences within a genome that are not found within protein-coding exons, and so are never represented within the amino acid sequence of expressed proteins. By this definition, more than 98% of the human genomes is composed of ncDNA.
Numerous classes of noncoding DNA have been identified, including genes for noncoding RNA (e.g. tRNA and rRNA), pseudogenes, introns, untranslated regions of mRNA, regulatory DNA sequences, repetitive DNA sequences, and sequences related to mobile genetic elements.
Numerous sequences that are included within genes are also defined as noncoding DNA. These include genes for noncoding RNA (e.g. tRNA, rRNA), and untranslated components of protein-coding genes (e.g. introns, and 5' and 3' untranslated regions of mRNA).
Protein-coding sequences (specifically, coding exons) constitute less than 1.5% of the human genome. In addition, about 26% of the human genome is introns. Aside from genes (exons and introns) and known regulatory sequences (8–20%), the human genome contains regions of noncoding DNA. The exact amount of noncoding DNA that plays a role in cell physiology has been hotly debated. An analysis by the ENCODE project indicates that 80% of the entire human genome is either transcribed, binds to regulatory proteins, or is associated with some other biochemical activity.
It however remains controversial whether all of this biochemical activity contributes to cell physiology, or whether a substantial portion of this is the result of transcriptional and biochemical noise, which must be actively filtered out by the organism. Excluding protein-coding sequences, introns, and regulatory regions, much of the non-coding DNA is composed of: Many DNA sequences that do not play a role in gene expression have important biological functions. Comparative genomics studies indicate that about 5% of the genome contains sequences of noncoding DNA that are highly conserved, sometimes on time-scales representing hundreds of millions of years, implying that these noncoding regions are under strong evolutionary pressure and purifying selection.
Many of these sequences regulate the structure of chromosomes by limiting the regions of heterochromatin formation and regulating structural features of the chromosomes, such as the telomeres and centromeres. Other noncoding regions serve as origins of DNA replication. Finally several regions are transcribed into functional noncoding RNA that regulate the expression of protein-coding genes (for example), mRNA translation and stability (see miRNA), chromatin structure (including histone modifications, for example), DNA methylation (for example), DNA recombination (for example), and cross-regulate other noncoding RNAs (for example). It is also likely that many transcribed noncoding regions do not serve any role and that this transcription is the product of non-specific RNA Polymerase activity.
Pseudogenes
Pseudogenes are inactive copies of protein-coding genes, often generated by gene duplication, that have become nonfunctional through the accumulation of inactivating mutations. The number of pseudogenes in the human genome is on the order of 13,000, and in some chromosomes is nearly the same as the number of functional protein-coding genes. Gene duplication is a major mechanism through which new genetic material is generated during molecular evolution.
For example, the olfactory receptor gene family is one of the best-documented examples of pseudogenes in the human genome. More than 60 percent of the genes in this family are non-functional pseudogenes in humans. By comparison, only 20 percent of genes in the mouse olfactory receptor gene family are pseudogenes. Research suggests that this is a species-specific characteristic, as the most closely related primates all have proportionally fewer pseudogenes. This genetic discovery helps to explain the less acute sense of smell in humans relative to other mammals.
Genes for noncoding RNA (ncRNA)
Noncoding RNA molecules play many essential roles in cells, especially in the many reactions of protein synthesis and RNA processing. Noncoding RNA include tRNA, ribosomal RNA, microRNA, snRNA and other non-coding RNA genes including about 60,000 long non-coding RNAs (lncRNAs). Although the number of reported lncRNA genes continues to rise and the exact number in the human genome is yet to be defined, many of them are argued to be non-functional.
Many ncRNAs are critical elements in gene regulation and expression. Noncoding RNA also contributes to epigenetics, transcription, RNA splicing, and the translational machinery. The role of RNA in genetic regulation and disease offers a new potential level of unexplored genomic complexity.
Introns and untranslated regions of mRNA
In addition to the ncRNA molecules that are encoded by discrete genes, the initial transcripts of protein coding genes usually contain extensive noncoding sequences, in the form of introns, 5'-untranslated regions (5'-UTR), and 3'-untranslated regions (3'-UTR). Within most protein-coding genes of the human genome, the length of intron sequences is 10- to 100-times the length of exon sequences.
Regulatory DNA sequences
The human genome has many different regulatory sequences which are crucial to controlling gene expression. Conservative estimates indicate that these sequences make up 8% of the genome, however extrapolations from the ENCODE project give that 20-40% of the genome is gene regulatory sequence. Some types of non-coding DNA are genetic "switches" that do not encode proteins, but do regulate when and where genes are expressed (called enhancers).
Regulatory sequences have been known since the late 1960s. The first identification of regulatory sequences in the human genome relied on recombinant DNA technology. Later with the advent of genomic sequencing, the identification of these sequences could be inferred by evolutionary conservation. The evolutionary branch between the primates and mouse, for example, occurred 70–90 million years ago. So computer comparisons of gene sequences that identify conserved non-coding sequences will be an indication of their importance in duties such as gene regulation.
Other genomes have been sequenced with the same intention of aiding conservation-guided methods, for exampled the pufferfish genome. However, regulatory sequences disappear and re-evolve during evolution at a high rate.
As of 2012, the efforts have shifted toward finding interactions between DNA and regulatory proteins by the technique ChIP-Seq, or gaps where the DNA is not packaged by histones (DNase hypersensitive sites), both of which tell where there are active regulatory sequences in the investigated cell type.
Repetitive DNA sequences
Repetitive DNA sequences comprise approximately 50% of the human genome.
About 8% of the human genome consists of tandem DNA arrays or tandem repeats, low complexity repeat sequences that have multiple adjacent copies (e.g. "CAGCAGCAG..."). The tandem sequences may be of variable lengths, from two nucleotides to tens of nucleotides. These sequences are highly variable, even among closely related individuals, and so are used for genealogical DNA testing and forensic DNA analysis.
Repeated sequences of fewer than ten nucleotides (e.g. the dinucleotide repeat (AC)n) are termed microsatellite sequences. Among the microsatellite sequences, trinucleotide repeats are of particular importance, as sometimes occur within coding regions of genes for proteins and may lead to genetic disorders. For example, Huntington's disease results from an expansion of the trinucleotide repeat (CAG)n within the Huntingtin gene on human chromosome 4. Telomeres (the ends of linear chromosomes) end with a microsatellite hexanucleotide repeat of the sequence (TTAGGG)n.
Tandem repeats of longer sequences (arrays of repeated sequences 10–60 nucleotides long) are termed minisatellites.
Mobile genetic elements (transposons) and their relics
Transposable genetic elements, DNA sequences that can replicate and insert copies of themselves at other locations within a host genome, are an abundant component in the human genome. The most abundant transposon lineage, Alu, has about 50,000 active copies, and can be inserted into intragenic and intergenic regions. One other lineage, LINE-1, has about 100 active copies per genome (the number varies between people). Together with non-functional relics of old transposons, they account for over half of total human DNA. Sometimes called "jumping genes", transposons have played a major role in sculpting the human genome. Some of these sequences represent endogenous retroviruses, DNA copies of viral sequences that have become permanently integrated into the genome and are now passed on to succeeding generations.
Mobile elements within the human genome can be classified into LTR retrotransposons (8.3% of total genome), SINEs (13.1% of total genome) including Alu elements, LINEs (20.4% of total genome), SVAs (SINE-VNTR-Alu) and Class II DNA transposons (2.9% of total genome).
Genomic variation in humans
Human reference genome
With the exception of identical twins, all humans show significant variation in genomic DNA sequences. The human reference genome (HRG) is used as a standard sequence reference.
There are several important points concerning the human reference genome:
- The HRG is a haploid sequence. Each chromosome is represented once.
- The HRG is a composite sequence, and does not correspond to any actual human individual.
- The HRG is periodically updated to correct errors, ambiguities, and unknown "gaps".
- The HRG in no way represents an "ideal" or "perfect" human individual. It is simply a standardized representation or model that is used for comparative purposes.
The Genome Reference Consortium is responsible for updating the HRG. Version 38 was released in December 2013.
Measuring human genetic variation
Most studies of human genetic variation have focused on single-nucleotide polymorphisms (SNPs), which are substitutions in individual bases along a chromosome. Most analyses estimate that SNPs occur 1 in 1000 base pairs, on average, in the euchromatic human genome, although they do not occur at a uniform density. Thus follows the popular statement that "we are all, regardless of race, genetically 99.9% the same", although this would be somewhat qualified by most geneticists. For example, a much larger fraction of the genome is now thought to be involved in copy number variation. A large-scale collaborative effort to catalog SNP variations in the human genome is being undertaken by the International HapMap Project.
The genomic loci and length of certain types of small repetitive sequences are highly variable from person to person, which is the basis of DNA fingerprinting and DNA paternity testing technologies. The heterochromatic portions of the human genome, which total several hundred million base pairs, are also thought to be quite variable within the human population (they are so repetitive and so long that they cannot be accurately sequenced with current technology). These regions contain few genes, and it is unclear whether any significant phenotypic effect results from typical variation in repeats or heterochromatin.
Most gross genomic mutations in gamete germ cells probably result in inviable embryos; however, a number of human diseases are related to large-scale genomic abnormalities. Down syndrome, Turner Syndrome, and a number of other diseases result from nondisjunction of entire chromosomes. Cancer cells frequently have aneuploidy of chromosomes and chromosome arms, although a cause and effect relationship between aneuploidy and cancer has not been established.
Mapping human genomic variation
Whereas a genome sequence lists the order of every DNA base in a genome, a genome map identifies the landmarks. A genome map is less detailed than a genome sequence and aids in navigating around the genome.
An example of a variation map is the HapMap being developed by the International HapMap Project. The HapMap is a haplotype map of the human genome, "which will describe the common patterns of human DNA sequence variation." It catalogs the patterns of small-scale variations in the genome that involve single DNA letters, or bases.
Researchers published the first sequence-based map of large-scale structural variation across the human genome in the journal Nature in May 2008. Large-scale structural variations are differences in the genome among people that range from a few thousand to a few million DNA bases; some are gains or losses of stretches of genome sequence and others appear as re-arrangements of stretches of sequence. These variations include differences in the number of copies individuals have of a particular gene, deletions, translocations and inversions.
Structural variation
Structural variation refers to genetic variants that affect larger segments of the human genome, as opposed to point mutations. Often, structural variants (SVs) are defined as variants of 50 base pairs (bp) or greater, such as deletions, duplications, insertions, inversions and other rearrangements. About 90% of structural variants are noncoding deletions but most individuals have more than a thousand such deletions; the size of deletions ranges from dozens of base pairs to tens of thousands of bp. On average, individuals carry ~3 rare structural variants that alter coding regions, e.g. delete exons. About 2% of individuals carry ultra-rare megabase-scale structural variants, especially rearrangements. That is, millions of base pairs may be inverted within a chromosome; ultra-rare means that they are only found in individuals or their family members and thus have arisen very recently.
SNP frequency across the human genome
Single-nucleotide polymorphisms (SNPs) do not occur homogeneously across the human genome. In fact, there is enormous diversity in SNP frequency between genes, reflecting different selective pressures on each gene as well as different mutation and recombination rates across the genome. However, studies on SNPs are biased towards coding regions, the data generated from them are unlikely to reflect the overall distribution of SNPs throughout the genome. Therefore, the SNP Consortium protocol was designed to identify SNPs with no bias towards coding regions and the Consortium's 100,000 SNPs generally reflect sequence diversity across the human chromosomes. The SNP Consortium aims to expand the number of SNPs identified across the genome to 300 000 by the end of the first quarter of 2001.
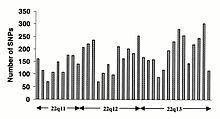
Changes in non-coding sequence and synonymous changes in coding sequence are generally more common than non-synonymous changes, reflecting greater selective pressure reducing diversity at positions dictating amino acid identity. Transitional changes are more common than transversions, with CpG dinucleotides showing the highest mutation rate, presumably due to deamination.
Personal genomes
A personal genome sequence is a (nearly) complete sequence of the chemical base pairs that make up the DNA of a single person. Because medical treatments have different effects on different people due to genetic variations such as single-nucleotide polymorphisms (SNPs), the analysis of personal genomes may lead to personalized medical treatment based on individual genotypes.
The first personal genome sequence to be determined was that of Craig Venter in 2007. Personal genomes had not been sequenced in the public Human Genome Project to protect the identity of volunteers who provided DNA samples. That sequence was derived from the DNA of several volunteers from a diverse population. However, early in the Venter-led Celera Genomics genome sequencing effort the decision was made to switch from sequencing a composite sample to using DNA from a single individual, later revealed to have been Venter himself. Thus the Celera human genome sequence released in 2000 was largely that of one man. Subsequent replacement of the early composite-derived data and determination of the diploid sequence, representing both sets of chromosomes, rather than a haploid sequence originally reported, allowed the release of the first personal genome. In April 2008, that of James Watson was also completed. In 2009, Stephen Quake published his own genome sequence derived from a sequencer of his own design, the Heliscope. A Stanford team led by Euan Ashley published a framework for the medical interpretation of human genomes implemented on Quake's genome and made whole genome-informed medical decisions for the first time. That team further extended the approach to the West family, the first family sequenced as part of Illumina's Personal Genome Sequencing program. Since then hundreds of personal genome sequences have been released, including those of Desmond Tutu, and of a Paleo-Eskimo. In 2012, the whole genome sequences of two family trios among 1092 genomes was made public. In November 2013, a Spanish family made four personal exome datasets (about 1% of the genome) publicly available under a Creative Commons public domain license. The Personal Genome Project (started in 2005) is among the few to make both genome sequences and corresponding medical phenotypes publicly available.
The sequencing of individual genomes further unveiled levels of genetic complexity that had not been appreciated before. Personal genomics helped reveal the significant level of diversity in the human genome attributed not only to SNPs but structural variations as well. However, the application of such knowledge to the treatment of disease and in the medical field is only in its very beginnings. Exome sequencing has become increasingly popular as a tool to aid in diagnosis of genetic disease because the exome contributes only 1% of the genomic sequence but accounts for roughly 85% of mutations that contribute significantly to disease.
Human knockouts
In humans, gene knockouts naturally occur as heterozygous or homozygous loss-of-function gene knockouts. These knockouts are often difficult to distinguish, especially within heterogeneous genetic backgrounds. They are also difficult to find as they occur in low frequencies.
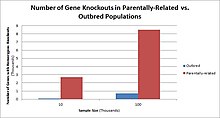
Populations with high rates of consanguinity, such as countries with high rates of first-cousin marriages, display the highest frequencies of homozygous gene knockouts. Such populations include Pakistan, Iceland, and Amish populations. These populations with a high level of parental-relatedness have been subjects of human knock out research which has helped to determine the function of specific genes in humans. By distinguishing specific knockouts, researchers are able to use phenotypic analyses of these individuals to help characterize the gene that has been knocked out.
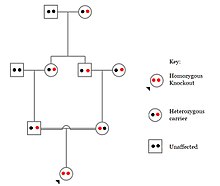
Knockouts in specific genes can cause genetic diseases, potentially have beneficial effects, or even result in no phenotypic effect at all. However, determining a knockout's phenotypic effect and in humans can be challenging. Challenges to characterizing and clinically interpreting knockouts include difficulty calling of DNA variants, determining disruption of protein function (annotation), and considering the amount of influence mosaicism has on the phenotype.
One major study that investigated human knockouts is the Pakistan Risk of Myocardial Infarction study. It was found that individuals possessing a heterozygous loss-of-function gene knockout for the APOC3 gene had lower triglycerides in the blood after consuming a high fat meal as compared to individuals without the mutation. However, individuals possessing homozygous loss-of-function gene knockouts of the APOC3 gene displayed the lowest level of triglycerides in the blood after the fat load test, as they produce no functional APOC3 protein.
Human genetic disorders
Most aspects of human biology involve both genetic (inherited) and non-genetic (environmental) factors. Some inherited variation influences aspects of our biology that are not medical in nature (height, eye color, ability to taste or smell certain compounds, etc.). Moreover, some genetic disorders only cause disease in combination with the appropriate environmental factors (such as diet). With these caveats, genetic disorders may be described as clinically defined diseases caused by genomic DNA sequence variation. In the most straightforward cases, the disorder can be associated with variation in a single gene. For example, cystic fibrosis is caused by mutations in the CFTR gene and is the most common recessive disorder in caucasian populations with over 1,300 different mutations known.
Disease-causing mutations in specific genes are usually severe in terms of gene function and are fortunately rare, thus genetic disorders are similarly individually rare. However, since there are many genes that can vary to cause genetic disorders, in aggregate they constitute a significant component of known medical conditions, especially in pediatric medicine. Molecularly characterized genetic disorders are those for which the underlying causal gene has been identified. Currently there are approximately 2,200 such disorders annotated in the OMIM database.
Studies of genetic disorders are often performed by means of family-based studies. In some instances, population based approaches are employed, particularly in the case of so-called founder populations such as those in Finland, French-Canada, Utah, Sardinia, etc. Diagnosis and treatment of genetic disorders are usually performed by a geneticist-physician trained in clinical/medical genetics. The results of the Human Genome Project are likely to provide increased availability of genetic testing for gene-related disorders, and eventually improved treatment. Parents can be screened for hereditary conditions and counselled on the consequences, the probability of inheritance, and how to avoid or ameliorate it in their offspring.
There are many different kinds of DNA sequence variation, ranging from complete extra or missing chromosomes down to single nucleotide changes. It is generally presumed that much naturally occurring genetic variation in human populations is phenotypically neutral, i.e., has little or no detectable effect on the physiology of the individual (although there may be fractional differences in fitness defined over evolutionary time frames). Genetic disorders can be caused by any or all known types of sequence variation. To molecularly characterize a new genetic disorder, it is necessary to establish a causal link between a particular genomic sequence variant and the clinical disease under investigation. Such studies constitute the realm of human molecular genetics.
With the advent of the Human Genome and International HapMap Project, it has become feasible to explore subtle genetic influences on many common disease conditions such as diabetes, asthma, migraine, schizophrenia, etc. Although some causal links have been made between genomic sequence variants in particular genes and some of these diseases, often with much publicity in the general media, these are usually not considered to be genetic disorders per se as their causes are complex, involving many different genetic and environmental factors. Thus there may be disagreement in particular cases whether a specific medical condition should be termed a genetic disorder.
Additional genetic disorders of mention are Kallman syndrome and Pfeiffer syndrome (gene FGFR1), Fuchs corneal dystrophy (gene TCF4), Hirschsprung's disease (genes RET and FECH), Bardet-Biedl syndrome 1 (genes CCDC28B and BBS1), Bardet-Biedl syndrome 10 (gene BBS10), and facioscapulohumeral muscular dystrophy type 2 (genes D4Z4 and SMCHD1).
Genome sequencing is now able to narrow the genome down to specific locations to more accurately find mutations that will result in a genetic disorder. Copy number variants (CNVs) and single nucleotide variants (SNVs) are also able to be detected at the same time as genome sequencing with newer sequencing procedures available, called Next Generation Sequencing (NGS). This only analyzes a small portion of the genome, around 1–2%. The results of this sequencing can be used for clinical diagnosis of a genetic condition, including Usher syndrome, retinal disease, hearing impairments, diabetes, epilepsy, Leigh disease, hereditary cancers, neuromuscular diseases, primary immunodeficiencies, severe combined immunodeficiency (SCID), and diseases of the mitochondria. NGS can also be used to identify carriers of diseases before conception. The diseases that can be detected in this sequencing include Tay-Sachs disease, Bloom syndrome, Gaucher disease, Canavan disease, familial dysautonomia, cystic fibrosis, spinal muscular atrophy, and fragile-X syndrome. The Next Genome Sequencing can be narrowed down to specifically look for diseases more prevalent in certain ethnic populations.
| Disorder | Prevalence | Chromosome or gene involved |
|---|---|---|
| Chromosomal conditions | ||
| Down syndrome | 1:600 | Chromosome 21 |
| Klinefelter syndrome | 1:500–1000 males | Additional X chromosome |
| Turner syndrome | 1:2000 females | Loss of X chromosome |
| Sickle cell anemia | 1 in 50 births in parts of Africa; rarer elsewhere | β-globin (on chromosome 11) |
| Bloom syndrome | 1:48000 Ashkenazi Jews | BLM |
| Cancers | ||
| Breast/Ovarian cancer (susceptibility) | ~5% of cases of these cancer types | BRCA1, BRCA2 |
| FAP (hereditary nonpolyposis coli) | 1:3500 | APC |
| Lynch syndrome | 5–10% of all cases of bowel cancer | MLH1, MSH2, MSH6, PMS2 |
| Fanconi anemia | 1:130000 births | FANCC |
| Neurological conditions | ||
| Huntington disease | 1:20000 | Huntingtin |
| Alzheimer disease - early onset | 1:2500 | PS1, PS2, APP |
| Tay-Sachs | 1:3600 births in Ashkenazi Jews | HEXA gene (on chromosome 15) |
| Canavan disease | 2.5% Eastern European Jewish ancestry | ASPA gene (on chromosome 17) |
| Familial dysautonomia | 600 known cases worldwide since discovery | IKBKAP gene (on chromosome 9) |
| Fragile X syndrome | 1.4:10000 in males, 0.9:10000 in females | FMR1 gene (on X chromosome) |
| Mucolipidosis type IV | 1:90 to 1:100 in Ashkenazi Jews | MCOLN1 |
| Other conditions | ||
| Cystic fibrosis | 1:2500 | CFTR |
| Duchenne muscular dystrophy | 1:3500 boys | Dystrophin |
| Becker muscular dystrophy | 1.5–6:100000 males | DMD |
| Beta thalassemia | 1:100000 | HBB |
| Congenital adrenal hyperplasia | 1:280 in Native Americans and Yupik Eskimos
1:15000 in American Caucasians |
CYP21A2 |
| Glycogen storage disease type I | 1:100000 births in America | G6PC |
| Maple syrup urine disease | 1:180000 in the U.S.
1:176 in Mennonite/Amish communities 1:250000 in Austria |
BCKDHA, BCKDHB, DBT, DLD |
| Niemann–Pick disease, SMPD1-associated | 1,200 cases worldwide | SMPD1 |
| Usher syndrome | 1:23000 in the U.S.
1:28000 in Norway 1:12500 in Germany |
CDH23, CLRN1, DFNB31, GPR98, MYO7A, PCDH15, USH1C, USH1G, USH2A |
Evolution
−10 — – −9 — – −8 — – −7 — – −6 — – −5 — – −4 — – −3 — – −2 — – −1 — – 0 — |
| |||||||||||||||||||||||||||
Comparative genomics studies of mammalian genomes suggest that approximately 5% of the human genome has been conserved by evolution since the divergence of extant lineages approximately 200 million years ago, containing the vast majority of genes. The published chimpanzee genome differs from that of the human genome by 1.23% in direct sequence comparisons. Around 20% of this figure is accounted for by variation within each species, leaving only ~1.06% consistent sequence divergence between humans and chimps at shared genes. This nucleotide by nucleotide difference is dwarfed, however, by the portion of each genome that is not shared, including around 6% of functional genes that are unique to either humans or chimps.
In other words, the considerable observable differences between humans and chimps may be due as much or more to genome level variation in the number, function and expression of genes rather than DNA sequence changes in shared genes. Indeed, even within humans, there has been found to be a previously unappreciated amount of copy number variation (CNV) which can make up as much as 5–15% of the human genome. In other words, between humans, there could be +/- 500,000,000 base pairs of DNA, some being active genes, others inactivated, or active at different levels. The full significance of this finding remains to be seen. On average, a typical human protein-coding gene differs from its chimpanzee ortholog by only two amino acid substitutions; nearly one third of human genes have exactly the same protein translation as their chimpanzee orthologs. A major difference between the two genomes is human chromosome 2, which is equivalent to a fusion product of chimpanzee chromosomes 12 and 13. (later renamed to chromosomes 2A and 2B, respectively).
Humans have undergone an extraordinary loss of olfactory receptor genes during our recent evolution, which explains our relatively crude sense of smell compared to most other mammals. Evolutionary evidence suggests that the emergence of color vision in humans and several other primate species has diminished the need for the sense of smell.
In September 2016, scientists reported that, based on human DNA genetic studies, all non-Africans in the world today can be traced to a single population that exited Africa between 50,000 and 80,000 years ago.
Mitochondrial DNA
The human mitochondrial DNA is of tremendous interest to geneticists, since it undoubtedly plays a role in mitochondrial disease. It also sheds light on human evolution; for example, analysis of variation in the human mitochondrial genome has led to the postulation of a recent common ancestor for all humans on the maternal line of descent (see Mitochondrial Eve).
Due to the lack of a system for checking for copying errors, mitochondrial DNA (mtDNA) has a more rapid rate of variation than nuclear DNA. This 20-fold higher mutation rate allows mtDNA to be used for more accurate tracing of maternal ancestry. Studies of mtDNA in populations have allowed ancient migration paths to be traced, such as the migration of Native Americans from Siberia or Polynesians from southeastern Asia. It has also been used to show that there is no trace of Neanderthal DNA in the European gene mixture inherited through purely maternal lineage. Due to the restrictive all or none manner of mtDNA inheritance, this result (no trace of Neanderthal mtDNA) would be likely unless there were a large percentage of Neanderthal ancestry, or there was strong positive selection for that mtDNA. For example, going back 5 generations, only 1 of a person's 32 ancestors contributed to that person's mtDNA, so if one of these 32 was pure Neanderthal an expected ~3% of that person's autosomal DNA would be of Neanderthal origin, yet they would have a ~97% chance of having no trace of Neanderthal mtDNA.
Epigenome
Epigenetics describes a variety of features of the human genome that transcend its primary DNA sequence, such as chromatin packaging, histone modifications and DNA methylation, and which are important in regulating gene expression, genome replication and other cellular processes. Epigenetic markers strengthen and weaken transcription of certain genes but do not affect the actual sequence of DNA nucleotides. DNA methylation is a major form of epigenetic control over gene expression and one of the most highly studied topics in epigenetics. During development, the human DNA methylation profile experiences dramatic changes. In early germ line cells, the genome has very low methylation levels. These low levels generally describe active genes. As development progresses, parental imprinting tags lead to increased methylation activity.
Epigenetic patterns can be identified between tissues within an individual as well as between individuals themselves. Identical genes that have differences only in their epigenetic state are called epialleles. Epialleles can be placed into three categories: those directly determined by an individual's genotype, those influenced by genotype, and those entirely independent of genotype. The epigenome is also influenced significantly by environmental factors. Diet, toxins, and hormones impact the epigenetic state. Studies in dietary manipulation have demonstrated that methyl-deficient diets are associated with hypomethylation of the epigenome. Such studies establish epigenetics as an important interface between the environment and the genome.