Micrograph showing condensed chromosomes in blue, kinetochores in pink, and microtubules in green during metaphase of mitosis
In cell biology, the spindle apparatus (or mitotic spindle) refers to the cytoskeletal structure of eukaryotic cells that forms during cell division to separate sister chromatids between daughter cells. It is referred to as the mitotic spindle during mitosis, a process that produces genetically identical daughter cells, or the meiotic spindle during meiosis, a process that produces gametes with half the number of chromosomes of the parent cell.
Besides chromosomes, the spindle apparatus is composed of hundreds of proteins. Microtubules comprise the most abundant components of the machinery.
Spindle structure
This diagram depicts the organization of a typical mitotic spindle found in animal cells. Chromosomes are attached to kinetochore microtubules
via a multiprotein complex called the kinetochore. Polar microtubules
interdigitate at the spindle midzone and push the spindle poles apart
via motor proteins. Astral microtubules anchor the spindle poles to the cell membrane. Microtubule polymerization is nucleated at the microtubule organizing center.
Attachment of microtubules to chromosomes is mediated by kinetochores, which actively monitor spindle formation and prevent premature anaphase
onset. Microtubule polymerization and depolymerization dynamic drive
chromosome congression. Depolymerization of microtubules generates
tension at kinetochores;
bipolar attachment of sister kinetochores to microtubules emanating
from opposite cell poles couples opposing tension forces, aligning
chromosomes at the cell equator and poising them for segregation to
daughter cells. Once every chromosome is bi-oriented, anaphase commences
and cohesin, which couples sister chromatids, is severed, permitting the transit of the sister chromatids to opposite poles.
The cellular spindle apparatus includes the spindle microtubules, associated proteins, which include kinesin and dynein molecular motors, condensed chromosomes, and any centrosomes or asters that may be present at the spindle poles depending on the cell type. The spindle apparatus is vaguely ellipsoid in cross section and tapers at the ends. In the wide middle portion, known as the spindle midzone, antiparallel microtubules are bundled by kinesins. At the pointed ends, known as spindle poles, microtubules are nucleated by the centrosomes in most animal cells. Acentrosomal or anastral
spindles lack centrosomes or asters at the spindle poles, respectively,
and occur for example during female meiosis in most animals. In this instance, a Ran GTP gradient is the main regulator of spindle microtubule organization and assembly. In fungi, spindles form between spindle pole bodies embedded in the nuclear envelope, which does not break down during mitosis.
Microtubule-associated proteins and spindle dynamics
The dynamic lengthening and shortening of spindle microtubules, through a process known as dynamic instability
determines to a large extent the shape of the mitotic spindle and
promotes the proper alignment of chromosomes at the spindle midzone. Microtubule-associated proteins (MAPs) associate with microtubules at the midzone and the spindle poles to regulate their dynamics. γ-tubulin is a specialized tubulin variant that assembles into a ring complex called γ-TuRC which nucleates polymerization of α/β tubulin heterodimers
into microtubules. Recruitment of γ-TuRC to the pericentrosomal region
stabilizes microtubule minus-ends and anchors them near the microtubule-organizing center.
The microtubule-associated protein Augmin acts in conjunction with
γ-TURC to nucleate new microtubules off of existing microtubules.
The growing ends of microtubules are protected against
catastrophe by the action of plus-end microtubule tracking proteins
(+TIPs) to promote their association with kinetochores at the midzone. CLIP170 was shown to localize near microtubule plus-ends in HeLa cells and to accumulate in kinetochores during prometaphase.
Although how CLIP170 recognizes plus-ends remains unclear, it has been
shown that its homologues protect against catastrophe and promote
rescue, suggesting a role for CLIP170 in stabilizing plus-ends and possibly mediating their direct attachment to kinetochores. CLIP-associated proteins like CLASP1
in humans have also been shown to localize to plus-ends and the outer
kinetochore as well as to modulate the dynamics of kinetochore
microtubules (Maiato 2003). CLASP homologues in Drosophila, Xenopus, and yeast
are required for proper spindle assembly; in mammals, CLASP1 and CLASP2
both contribute to proper spindle assembly and microtubule dynamics in
anaphase.
Plus-end polymerization may be further moderated by the EB1 protein,
which directly binds the growing ends of microtubules and coordinates
the binding of other +TIPs.
Opposing the action of these microtubule-stabilizing proteins are
a number of microtubule-depolymerizing factors which permit the dynamic
remodeling of the mitotic spindle to promote chromosome congression and
attainment of bipolarity. The kinesin-13
superfamily of MAPs contains a class of plus-end-directed motor
proteins with associated microtubule depolymerization activity including
the well-studied mammalian MCAK and Xenopus XKCM1. MCAK
localizes to the growing tips of microtubules at kinetochores where it
can trigger catastrophe in direct competition with stabilizing +TIP
activity. These proteins harness the energy of ATP hydrolysis
to induce destabilizing conformational changes in protofilament
structure that cause kinesin release and microtubule depolymerization. Loss of their activity results in numerous mitotic defects. Additional microtubule destabilizing proteins include Op18/stathmin and katanin which have roles in remodeling the mitotic spindle as well as promoting chromosome segregation during anaphase.
The activities of these MAPs are carefully regulated to maintain
proper microtubule dynamics during spindle assembly, with many of these
proteins serving as Aurora and Polo-like kinase substrates.
Organizing the spindle apparatus
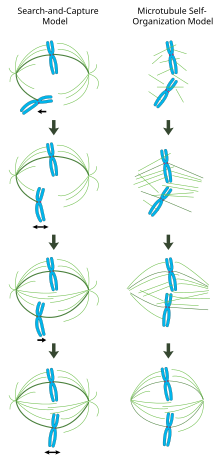
In
the centrosome-mediated “search and capture” model (left), microtubules
nucleated from centrosomes contact chromosomes by chance and become
stabilized at kinetochores to form the spindle. In the
chromatin-mediated “self-organization” model (right), microtubules are
nucleated around the vicinity of mitotic chromatin and organized into a
bipolar array by motor proteins.
In a properly formed mitotic spindle, bi-oriented chromosomes are
aligned along the equator of the cell with spindle microtubules oriented
roughly perpendicular to the chromosomes, their plus-ends embedded in
kinetochores and their minus-ends anchored at the cell poles. The
precise orientation of this complex is required to ensure accurate
chromosome segregation and to specify the cell division plane. However,
it remains unclear how the spindle becomes organized. Two models
predominate the field, which are synergistic and not mutually exclusive.
In the search-and-capture model, the spindle is predominantly
organized by the poleward separation of centrosomal microtubule
organizing centers (MTOCs). Spindle microtubules emanate from
centrosomes and 'seek' out kinetochores; when they bind a kinetochore
they become stabilized and exert tension on the chromosomes. In an
alternative self assembly model, microtubules undergo
acentrosomal nucleation among the condensed chromosomes. Constrained by
cellular dimensions, lateral associations with antiparallel microtubules
via motor proteins, and end-on attachments to kinetochores,
microtubules naturally adopt a spindle-like structure with chromosomes
aligned along the cell equator.
Centrosome-mediated "search-and-capture" model
In
this model, microtubules are nucleated at microtubule organizing
centers and undergo rapid growth and catastrophe to 'search' the
cytoplasm for kinetochores. Once they bind a kinetochore, they are
stabilized and their dynamics are reduced. The newly mono-oriented
chromosome oscillates in space near the pole to which it is attached
until a microtubule from the opposite pole binds the sister kinetochore.
This second attachment further stabilizes kinetochore attachment to the
mitotic spindle. Gradually, the bi-oriented chromosome is pulled
towards the center of the cell until microtubule tension is balanced on
both sides of the centromere;
the congressed chromosome then oscillates at the metaphase plate until
anaphase onset releases cohesion of the sister chromatids.
In this model, microtubule organizing centers are localized to
the cell poles, their separation driven by microtubule polymerization
and 'sliding' of antiparallel spindle microtubules with respect to one
another at the spindle midzone mediated by bipolar, plus-end-directed
kinesins.[19][20]
Such sliding forces may account not only for spindle pole separation
early in mitosis, but also spindle elongation during late anaphase.
Chromatin-mediated self-organization of the mitotic spindle
In
contrast to the search-and-capture mechanism in which centrosomes
largely dictate the organization of the mitotic spindle, this model
proposes that microtubules are nucleated acentrosomally near chromosomes
and spontaneously assemble into anti-parallel bundles and adopt a
spindle-like structure.
Classic experiments by Heald and Karsenti show that functional mitotic
spindles and nuclei form around DNA-coated beads incubated in Xenopus egg extracts and that bipolar arrays of microtubules are formed in the absence of centrosomes and kinetochores.
Indeed, it has also been shown that laser ablation of centrosomes in
vertebrate cells inhibits neither spindle assembly nor chromosome
segregation.
Under this scheme, the shape and size of the mitotic spindle are a
function of the biophysical properties of the cross-linking motor
proteins.
Chromatin-mediated microtubule nucleation by the Ran GTP gradient
The guanine nucleotide exchange factor for the small GTPase Ran (Regulator of chromosome condensation 1 or RCC1) is attached to nucleosomes via core histones H2A and H2B.
Thus, a gradient of GTP-bound Ran is generated around the vicinity of
mitotic chromatin. Glass beads coated with RCC1 induce microtubule
nucleation and bipolar spindle formation in Xenopus egg extracts, revealing that the Ran GTP gradient alone is sufficient for spindle assembly.
The gradient triggers release of spindle assembly factors (SAFs) from
inhibitory interactions via the transport proteins importin β/α. The
unbound SAFs then promote microtubule nucleation and stabilization
around mitotic chromatin, and spindle bipolarity is organized by
microtubule motor proteins.
Regulation of spindle assembly
Spindle
assembly is largely regulated by phosphorylation events catalyzed by
mitotic kinases.
Cyclin dependent kinase complexes (CDKs) are activated by mitotic
cyclins, whose translation increases during mitosis. CDK1 (also called
CDC2) is considered the main mitotic kinase in mammalian cells and is
activated by Cyclin B1.
Aurora kinases are required for proper spindle assembly and separation. Aurora A associates with centrosomes and is believed to regulate mitotic entry. Aurora B is a member of the chromosomal passenger complex and mediates chromosome-microtubule attachment and sister chromatid cohesion.
Polo-like kinase, also known as PLK, especially PLK1 has important roles in the spindle maintenance by regulating microtubule dynamics.
Mitotic chromosome structure
By the end of DNA replication, sister chromatids
are bound together in an amorphous mass of tangled DNA and protein that
would be virtually impossible to partition into each daughter cell. To
avoid this problem, mitotic entry triggers a dramatic reorganization of
the duplicated genome. Sister chromatids are disentangled and resolved
from one another. Chromosomes also shorten in length, up to 10,000 fold
in animal cells,
in a process called condensation. Condensation begins in prophase and
chromosomes are maximally compacted into rod-shaped structures by the
time they are aligned in the middle of the spindle at metaphase. This
gives mitotic chromosomes the classic “X” shape seen in karyotypes, with each condensed sister chromatid linked along their lengths by cohesin proteins and joined, often near the center, at the centromere.
While these dynamic rearrangements are vitally important to
ensure accurate and high-fidelity segregation of the genome, our
understanding of mitotic chromosome structure remains largely
incomplete. A few specific molecular players have been identified,
however: Topoisomerase II uses ATP hydrolysis to catalyze decatenation
of DNA entanglements, promoting sister chromatid resolution. Condensins are 5-subunit complexes that also use ATP-hydrolysis to promote chromosome condensation. Experiments in Xenopus egg extracts have also implicated linker Histone H1 as an important regulator of mitotic chromosome compaction.
Mitotic spindle assembly checkpoint
The completion of spindle formation is a crucial transition point in the cell cycle called the spindle assembly checkpoint.
If chromosomes are not properly attached to the mitotic spindle by the
time of this checkpoint, the onset of anaphase will be delayed. Failure of this spindle assembly checkpoint can result in aneuploidy and may be involved in aging and the formation of cancer.
Spindle apparatus orientation
Cartoon
of the dividing epithelium cell surrounded by epithelium tissue.
Spindle apparatus rotates inside the cell. The rotation is a result of
astral microtubules pulling towards tri-cellular-junctions (TCJ),
signaling centers localized at the regions where three cells meet.
Cell division orientation
is of major importance for tissue architecture, cell fates and
morphogenesis. Cells tend to divide along their long axis according to
the so-called Hertwig rule.
The axis of cell division is determined by the orientation of the
spindle apparatus. Cells divide along the line connecting two
centrosomes of the spindle apparatus. After formation, the spindle
apparatus undergoes rotation inside the cell. The astral microtubules
originating from centrosomes reach the cell membrane where they are
pulled towards specific cortical clues. In vitro, the distribution of cortical clues is set up by the adhesive pattern. In vivo polarity cues are determined by localization of Tricellular junctions localized at cell vertices.
The spatial distribution of cortical clues leads to the force field
that determine final spindle apparatus orientation and the subsequent
orientation of cell division.