MALDI TOF mass spectrometer
In mass spectrometry, matrix-assisted laser desorption/ionization (MALDI) is an ionization technique that uses a laser energy absorbing matrix to create ions from large molecules with minimal fragmentation. It has been applied to the analysis of biomolecules (biopolymers such as DNA, proteins, peptides and sugars) and large organic molecules (such as polymers, dendrimers and other macromolecules), which tend to be fragile and fragment when ionized by more conventional ionization methods. It is similar in character to electrospray ionization
(ESI) in that both techniques are relatively soft (low fragmentation)
ways of obtaining ions of large molecules in the gas phase, though MALDI
typically produces far fewer multi-charged ions.
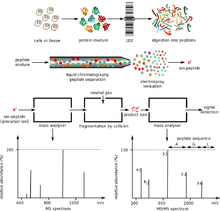
MALDI methodology is a three-step process. First, the sample is
mixed with a suitable matrix material and applied to a metal plate.
Second, a pulsed laser irradiates the sample, triggering ablation and desorption of the sample and matrix material. Finally, the analyte molecules are ionized by being protonated or deprotonated in the hot plume of ablated gases, and then they can be accelerated into whichever mass spectrometer is used to analyse them.
History
The term matrix-assisted laser desorption ionization (MALDI) was coined in 1985 by Franz Hillenkamp, Michael Karas and their colleagues. These researchers found that the amino acid alanine could be ionized more easily if it was mixed with the amino acid tryptophan
and irradiated with a pulsed 266 nm laser. The tryptophan was absorbing
the laser energy and helping to ionize the non-absorbing alanine.
Peptides up to the 2843 Da peptide melittin could be ionized when mixed with this kind of “matrix”. The breakthrough for large molecule laser desorption ionization came in 1987 when Koichi Tanaka
of Shimadzu Corporation and his co-workers used what they called the
“ultra fine metal plus liquid matrix method” that combined 30 nm cobalt particles in glycerol with a 337 nm nitrogen laser for ionization.
Using this laser and matrix combination, Tanaka was able to ionize
biomolecules as large as the 34,472 Da protein carboxypeptidase-A.
Tanaka received one-quarter of the 2002 Nobel Prize in Chemistry for demonstrating that, with the proper combination of laser wavelength and matrix, a protein can be ionized.
Karas and Hillenkamp were subsequently able to ionize the 67 kDa
protein albumin using a nicotinic acid matrix and a 266 nm laser. Further improvements were realized through the use of a 355 nm laser and the cinnamic acid derivatives ferulic acid, caffeic acid and sinapinic acid as the matrix. The availability of small and relatively inexpensive nitrogen lasers
operating at 337 nm wavelength and the first commercial instruments
introduced in the early 1990s brought MALDI to an increasing number of
researchers. Today, mostly organic matrices are used for MALDI mass spectrometry.
Matrix
| Compound | Other Names | Solvent | Wavelength (nm) | Applications |
|---|---|---|---|---|
| 2,5-dihydroxy benzoic acid | DHB, Gentisic acid | acetonitrile, water, methanol, acetone, chloroform | 337, 355, 266 | peptides, nucleotides, oligonucleotides, oligosaccharides |
| 3,5-dimethoxy-4-hydroxycinnamic acid | sinapic acid; sinapinic acid; SA | acetonitrile, water, acetone, chloroform | 337, 355, 266 | peptides, proteins, lipids |
| 4-hydroxy-3-methoxycinnamic acid | ferulic acid | acetonitrile, water, propanol | 337, 355, 266 | proteins |
| α-Cyano-4-hydroxycinnamic acid[12] | CHCA | acetonitrile, water, ethanol, acetone | 337, 355 | peptides, lipids, nucleotides |
| Picolinic acid | PA | Ethanol | 266 | oligonucleotides |
| 3-hydroxy picolinic acid | HPA | Ethanol | 337, 355 | oligonucleotides |
The matrix consists of crystallized molecules, of which the three most commonly used are 3,5-dimethoxy-4-hydroxycinnamic acid (sinapinic acid), α-cyano-4-hydroxycinnamic acid (α-CHCA, alpha-cyano or alpha-matrix) and 2,5-dihydroxybenzoic acid (DHB). A solution of one of these molecules is made, often in a mixture of highly purified water and an organic solvent such as acetonitrile (ACN) or ethanol. A counter ion source such as Trifluoroacetic acid (TFA) is usually added to generate the [M+H] ions. A good example of a matrix-solution would be 20 mg/mL sinapinic acid in ACN:water:TFA (50:50:0.1).
Notation for cinnamic acid substitutions.
The identification of suitable matrix compounds is determined to some
extent by trial and error, but they are based on some specific
molecular design considerations. They are of a fairly low molecular
weight (to allow easy vaporization), but are large enough (with a low
enough vapor pressure) not to evaporate during sample preparation or
while standing in the mass spectrometer. They are often acidic,
therefore act as a proton source to encourage ionization of the analyte.
Basic matrices have also been reported. They have a strong optical absorption in either the UV or IR range,
so that they rapidly and efficiently absorb the laser irradiation. This
efficiency is commonly associated with chemical structures
incorporating several conjugated double bonds,
as seen in the structure of cinnamic acid. They are functionalized with
polar groups, allowing their use in aqueous solutions. They typically
contain a chromophore.
The matrix solution is mixed with the analyte (e.g. protein-sample). A mixture of water and organic solvent allows both hydrophobic and water-soluble (hydrophilic)
molecules to dissolve into the solution. This solution is spotted onto a
MALDI plate (usually a metal plate designed for this purpose). The
solvents vaporize, leaving only the recrystallized matrix, but now with
analyte molecules embedded into MALDI crystals. The matrix and the
analyte are said to be co-crystallized. Co-crystallization is a key
issue in selecting a proper matrix to obtain a good quality mass
spectrum of the analyte of interest.
In analysis of biological systems, Inorganic salts, which are
also part of protein extracts, interfere with the ionization process.
The salts can be removed by solid phase extraction or by washing the
dried-droplet MALDI spots with cold water.
Both methods can also remove other substances from the sample. The
matrix-protein mixture is not homogenous because the polarity difference
leads to a separation of the two substances during co-crystallization.
The spot diameter of the target is much larger than that of the laser,
which makes it necessary to make many laser shots at different places of
the target, to get the statistical average of the substance
concentration within the target spot.
Naphthalene and naphthalene-like compounds can also be used as a matrix to ionize a sample
The matrix can be used to tune the instrument to ionize the sample in
different ways. As mentioned above, acid-base like reactions are often
utilized to ionize the sample, however, molecules with conjugated pi
systems, such as naphthalene like compounds, can also serve as an
electron acceptor and thus a matrix for MALDI/TOF. This is particularly useful in studying molecules that also possess conjugated pi systems. The most widely used application for these matrices is studying porphyrin like compounds such as chlorophyll.
These matrices have been shown to have better ionization patterns that
do not result in odd fragmentation patterns or complete loss of side
chains.
It has also been suggested that conjugated porphryin like molecules can
serve as a matrix and cleave themselves eliminating the need for a
separate matrix compound.
Instrumentation
Diagram
of a MALDI TOF instrument. Sample matrix ionized by radiant energy is
ejected from surface. Sample travels into mass analyzer and is
substantially detected.
There are several variations of the MALDI technology and comparable
instruments are today produced for very different purposes. From more
academic and analytical, to more industrial and high throughput. The MS
field has expanded into requiring ultrahigh resolution mass spectrometry
such as the FT-ICR instruments as well as more high-throughput instruments. As many MALDI MS instruments can be bought with an interchangeable ionization source (Electrospray ionization, MALDI, Atmospheric pressure ionization,
etc.) the technologies often overlap and many times any soft ionization
method could potentially be used. For more variations of soft
ionization methods go to Soft laser desorption or Ion source.
Laser
MALDI techniques typically employ the use of UV lasers such as nitrogen lasers (337 nm) and frequency-tripled and quadrupled Nd:YAG lasers (355 nm and 266 nm respectively).
Infrared laser wavelengths used for infrared MALDI include the 2.94 µm Er:YAG laser, mid-IR optical parametric oscillator, and 10.6 µm carbon dioxide laser. Although not as common, infrared lasers are used due to their softer mode of ionization.
IR-MALDI also has the advantage of greater material removal (useful for
biological samples), less low-mass interferences, and compatibility
with other matrix-free laser desorption mass spectrometry methods.
Time of flight
Sample target for a MALDI mass spectrometer
The type of a mass spectrometer most widely used with MALDI is the TOF
(time-of-flight mass spectrometer), mainly due to its large mass range.
The TOF measurement procedure is also ideally suited to the MALDI
ionization process since the pulsed laser takes individual 'shots'
rather than working in continuous operation. MALDI-TOF instrument or reflectron
is equipped with an "ion mirror" that reflects ions using an electric
field, thereby doubling the ion flight path and increasing the
resolution. Today, commercial reflectron
TOF instruments reach a resolving power m/Δm of well above 20,000 FWHM
(full-width half-maximum, Δm defined as the peak width at 50% of peak
height).
MALDI has been coupled with IMS-TOF MS to identify phosphorylated and non-phosphorylated peptides.
MALDI-FT-ICR MS has been demonstrated to be a useful technique where high resolution MALDI-MS measurements are desired.
Atmospheric pressure
Atmospheric
pressure (AP) matrix-assisted laser desorption/ionization (MALDI) is an
ionization technique (ion source) that in contrast to vacuum MALDI
operates at normal atmospheric environment.
The main difference between vacuum MALDI and AP-MALDI is the pressure
in which the ions are created. In vacuum MALDI, ions are typically
produced at 10 mTorr or less while in AP-MALDI ions are formed in atmospheric pressure.
In the past the main disadvantage of AP MALDI technique compared to the
conventional vacuum MALDI has been its limited sensitivity; however,
ions can be transferred into the mass spectrometer with high efficiency
and attomole detection limits have been reported.
AP-MALDI is used in mass spectrometry (MS) in a variety of applications
ranging from proteomics to drug discovery. Popular topics that are
addressed by AP-MALDI mass spectrometry include: proteomics; mass
analysis of DNA, RNA, PNA, lipids, oligosaccharides, phosphopeptides,
bacteria, small molecules and synthetic polymers, similar applications
as available also for vacuum MALDI instruments. The AP-MALDI ion source
is easily coupled to an ion trap mass spectrometer or any other MS system equipped with ESI (electrospray ionization) or nanoESI source.
Ionization mechanism
The laser
is fired at the matrix crystals in the dried-droplet spot. The matrix
absorbs the laser energy and it is thought that primarily the matrix is
desorbed and ionized (by addition of a proton)
by this event. The hot plume produced during ablation contains many
species: neutral and ionized matrix molecules, protonated and
deprotonated matrix molecules, matrix clusters and nanodroplets.
Ablated species may participate in the ionization of analyte, though
the mechanism of MALDI is still debated. The matrix is then thought to
transfer protons to the analyte molecules (e.g., protein molecules),
thus charging the analyte.
An ion observed after this process will consist of the initial neutral
molecule [M] with ions added or removed. This is called a quasimolecular
ion, for example [M+H]+ in the case of an added proton, [M+Na]+ in the case of an added sodium ion, or [M-H]− in the case of a removed proton. MALDI is capable of creating singly charged ions or multiply charged ions ([M+nH]n+)
depending on the nature of the matrix, the laser intensity, and/or the
voltage used. Note that these are all even-electron species. Ion signals
of radical cations (photoionized molecules) can be observed, e.g., in
the case of matrix molecules and other organic molecules.
The gas phase proton transfer model, implemented as the coupled physical and chemical dynamics (CPCD) model, of UV laser MALDI postulates primary and secondary processes leading to ionization.
Primary processes involve initial charge separation through absorption
of photons by the matrix and pooling of the energy to form matrix ion
pairs. Primary ion formation occurs through absorption of a UV photon to
create excited state molecules by
- S0 + hν → S1
- S1 + S1 → S0 + Sn
- S1 + Sn → M+ + M−
where S0 is the ground electronic state, S1 the first electronic excited state, and Sn is a higher electronic excited state. The product ions can be proton transfer or electron transfer ion pairs, indicated by M+ and M− above. Secondary processes involve ion-molecule reactions to form analyze ions.
In
the lucky survivor model, positive ions can be formed from highly
charged clusters produced during break-up of the matrix- and
analyte-containing solid.
The lucky survivor model (cluster ionization mechanism) postulates that analyte molecules are incorporated in the matrix maintaining the charge state from solution. Ion formation occurs through charge separation upon fragmentation of laser ablated clusters. Ions that are not neutralized by recombination with photoelectrons or counter ions are the so-called lucky survivors.
The thermal model postulates that the high temperature
facilitates the proton transfer between matrix and analyte in melted
matrix liquid.
Ion-to-neutral ratio is an important parameter to justify the
theoretical model, and the mistaken citation of ion-to-neutral ratio
could result in an erroneous determination of the ionization mechanism.
The model quantitatively predicts the increase in total ion intensity
as a function of the concentration and proton affinity of the analytes,
and the ion-to-neutral ratio as a function of the laser fluences. This model also suggests that metal ion adducts (e.g., [M+Na]+ or [M+K]+) are mainly generated from the thermally induced dissolution of salt.
The matrix-assisted ionization
(MAI) method uses matrix preparation similar to MALDI but does not
require laser ablation to produce analyte ions of volatile or
nonvolatile compounds.
Simply exposing the matrix with analyte to the vacuum of the mass
spectrometer creates ions with nearly identical charge states to
electrospray ionization. It is suggested that there are likely mechanistic commonality between this process and MALDI.
Applications
Biochemistry
In proteomics, MALDI is used for the rapid identification of proteins isolated by using gel electrophoresis: SDS-PAGE, size exclusion chromatography, affinity chromatography, strong/weak ion exchange, isotope coded protein labelling (ICPL), and two-dimensional gel electrophoresis. Peptide mass fingerprinting
is the most popular analytical application of MALDI-TOF mass
spectrometers. MALDI TOF/TOF mass spectrometers are used to reveal amino
acid sequence of peptides using post-source decay or high energy collision-induced dissociation (further use see mass spectrometry).
Loss of sialic acid has been identified in papers when DHB has
been used as a matrix for MALDI MS analysis of glycosylated peptides.
Using sinapinic acid, 4-HCCA and DHB as matrices, S. Martin studied loss
of sialic acid in glycosylated peptides by metastable decay in
MALDI/TOF in linear mode and reflector mode. A group at Shimadzu Corporation derivatized the sialic acid by an amidation reaction as a way to improve detection sensitivity
and also demonstrated that ionic liquid matrix reduces a loss of sialic
acid during MALDI/TOF MS analysis of sialylated oligosaccharides. THAP, DHAP, and a mixture of 2-aza-2-thiothymine and phenylhydrazine
have been identified as matrices that could be used to minimize loss of
sialic acid during MALDI MS analysis of glycosylated peptides.
It has been reported that a reduction in loss of some
post-translational modifications can be accomplished if IR MALDI is used
instead of UV MALDI.
In molecular biology, a mixture of 5-methoxysalicylic acid and spermine can be used as a matrix for oligonucleotides analysis in MALDI mass spectrometry, for instance after oligonucleotide synthesis.
Organic chemistry
Some synthetic macromolecules, such as catenanes and rotaxanes, dendrimers and hyperbranched polymers,
and other assemblies, have molecular weights extending into the
thousands or tens of thousands, where most ionization techniques have
difficulty producing molecular ions. MALDI is a simple and fast
analytical method that can allow chemists to rapidly analyze the results
of such syntheses and verify their results.
Polymers
In polymer chemistry MALDI can be used to determine the molar mass distribution. Polymers with polydispersity
greater than 1.2 are difficult to characterize with MALDI due to the
signal intensity discrimination against higher mass oligomers.
A good matrix for polymers is dithranol or AgTFA.
The sample must first be mixed with dithranol and the AgTFA added
afterwards; otherwise the sample would precipitate out of solution.
Microbiology
MALDI/TOF
spectra are used for the identification of micro-organisms such as
bacteria or fungi. A portion of a colony of the microbe in question is
placed onto the sample target and overlaid with matrix. The mass spectra
generated are analyzed by dedicated software and compared with stored
profiles. Species diagnosis by this procedure is much faster, more
accurate and cheaper than other procedures based on immunological or
biochemical tests. MALDI/TOF is becoming a standard method for species
identification in medical microbiological laboratories.
One main advantage over other microbiological identification
methods is its ability to rapidly and reliably identify, at low cost, a
wide variety of microorganisms directly from the selective medium used
to isolate them. The absence of the need to purify the suspect or
"presumptive" colony allows for a much faster turn-around times.
Another advantage is the potential to predict antibiotic susceptibility of bacteria. A single mass spectral peak can predict methicillin resistance of Staphylococcus aureus MALDI can also detect carbapenemase of carbapenem-resistant enterobacteriaceae, including Acinetobacter baumannii and Klebsiella pneumoniae.
However, most proteins that mediate antibiotic resistance are larger
than MALDI-TOF´s 2000-20,000 Da range for protein peak interpretation
and only occasionally, as in the 2011 KPC outbreak at the NIH, a
correlation between a peak and resistance conferring protein can be
made.
Parasitology
MALDI/TOF spectra have been used for the detection and identification of various parasites such as trypanosomatids, Leishmania and Plasmodium.
In addition to these unicellular parasites, MALDI/TOF can be used for the identification of cercariae, the free-swimming stage of trematodes.
Medicine
MALDI/TOF
spectra are often utilized in tandem with other analysis and
spectroscopy techniques in the diagnosis of diseases. MALDI/TOF is a
diagnostic tool with much potential because it allows for the rapid
identification of proteins and changes to proteins without the cost or
computing power of sequencing nor the skill or time needed to solve a crystal structure in X-ray crystallography.
One example of this is necrotizing enterocolitis
(NEC), which is a devastating disease that affects the bowels of
premature infants. The symptoms of NEC are very similar to those of sepsis,
and many infants die awaiting diagnosis and treatment. MALDI/TOF was
used to quickly analyze fecal samples and find differences between the
mutant and the functional protein responsible for NEC. There is hope
that a similar technique could be used as a quick, diagnostic tool that
would not require sequencing.
Another example of the diagnostic power of MALDI/TOF is in the area of cancer. Pancreatic cancer remains one of the most deadly and difficult to diagnose cancers. Impaired cellular signaling due to mutations in membrane proteins has been long suspected to contribute to pancreatic cancer. MALDI/TOF has been used to identify a membrane protein associated with pancreatic cancer and at one point may even serve as an early detection technique.
MALDI/TOF can also potentially be used to dictate treatment as
well as diagnosis. MALDI/TOF serves as a method for determining the drug resistance of bacteria, especially to β-lactams
(Penicillin family). The MALDI/TOF detects the presence of
carbapenemases, which indicates drug resistance to standard antibiotics.
It is predicted that this could serve as a method for identifying a
bacterium as drug resistant in as little as three hours. This technique
could help physicians decide whether to prescribe more aggressive
antibiotics initially.