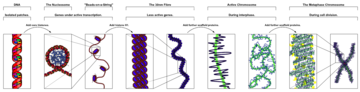
The major structures in DNA compaction: DNA, the nucleosome, the 10 nm "beads-on-a-string" fibre, the 30 nm chromatin fibre and the metaphase chromosome.
Chromatin is a complex of DNA and protein found in eukaryotic cells. Its primary function is packaging
long DNA molecules into more compact, denser structures. This prevents
the strands from becoming tangled and also plays important roles in
reinforcing the DNA during cell division, preventing DNA damage, and regulating gene expression and DNA replication. During mitosis and meiosis, chromatin facilitates proper segregation of the chromosomes in anaphase;
the characteristic shapes of chromosomes visible during this stage are
the result of DNA being coiled into highly condensed chromatin.
The primary protein components of chromatin are histones,
which bind to DNA and function as "anchors" around which the strands
are wound. In general, there are three levels of chromatin organization:
- DNA wraps around histone proteins, forming nucleosomes and the so-called "beads on a string" structure (euchromatin).
- Multiple histones wrap into a 30-nanometer fibre consisting of nucleosome arrays in their most compact form (heterochromatin).
- Higher-level DNA supercoiling of the 30-nm fiber produces the metaphase chromosome (during mitosis and meiosis).
Many organisms, however, do not follow this organization scheme. For example, spermatozoa and avian red blood cells have more tightly packed chromatin than most eukaryotic cells, and trypanosomatid protozoa do not condense their chromatin into visible chromosomes at all. Prokaryotic cells have entirely different structures for organizing their DNA (the prokaryotic chromosome equivalent is called a genophore and is localized within the nucleoid region).
The overall structure of the chromatin network further depends on the stage of the cell cycle. During interphase, the chromatin is structurally loose to allow access to RNA and DNA polymerases that transcribe and replicate the DNA. The local structure of chromatin during interphase depends on the specific genes
present in the DNA. Regions of DNA containing genes which are actively
transcribed ("turned on") are less tightly compacted and closely
associated with RNA polymerases in a structure known as euchromatin,
while regions containing inactive genes ("turned off") are generally
more condensed and associated with structural proteins in heterochromatin. Epigenetic modification of the structural proteins in chromatin via methylation and acetylation
also alters local chromatin structure and therefore gene expression.
The structure of chromatin networks is currently poorly understood and
remains an active area of research in molecular biology.
Dynamic chromatin structure and hierarchy
Chromatin undergoes various structural changes during a cell cycle. Histone
proteins are the basic packer and arranger of chromatin and can be
modified by various post-translational modifications to alter chromatin
packing (Histone modification).
Most of the modifications occur on the histone tail. The consequences
in terms of chromatin accessibility and compaction depend both on the
amino-acid that is modified and the type of modification. For example,
Histone acetylation results in loosening and increased accessibility of
chromatin for replication and transcription. Lysine tri-methylation can
either be correlated with transcriptional activity (tri-methylation of
histone H3 Lysine 4) or transcriptional repression and chromatin
compaction (tri-methylation of histone H3 Lysine 9 or 27). Several
studies suggested that different modifications could occur
simultaneously. For example, it was proposed that a bivalent structure (with tri-methylation of both Lysine 4 and 27 on histone H3) was involved in mammalian early development.
Polycomb-group proteins play a role in regulating genes through modulation of chromatin structure.
DNA structure
The structures of A-, B-, and Z-DNA.
In nature, DNA can form three structures, A-, B-, and Z-DNA.
A- and B-DNA are very similar, forming right-handed helices, whereas
Z-DNA is a left-handed helix with a zig-zag phosphate backbone. Z-DNA is
thought to play a specific role in chromatin structure and
transcription because of the properties of the junction between B- and
Z-DNA.
At the junction of B- and Z-DNA, one pair of bases is flipped out
from normal bonding. These play a dual role of a site of recognition by
many proteins and as a sink for torsional stress from RNA polymerase or nucleosome binding.
Nucleosomes and beads-on-a-string
The basic repeat element of chromatin is the nucleosome, interconnected by sections of linker DNA, a far shorter arrangement than pure DNA in solution.
In addition to the core histones, there is the linker histone,
H1, which contacts the exit/entry of the DNA strand on the nucleosome.
The nucleosome core particle, together with histone H1, is known as a
chromatosome. Nucleosomes, with about 20 to 60 base pairs of linker
DNA, can form, under non-physiological conditions, an approximately
10 nm "beads-on-a-string" fibre. (Fig. 1-2).
The nucleosomes bind DNA non-specifically, as required by their
function in general DNA packaging. There are, however, large DNA
sequence preferences that govern nucleosome positioning. This is due
primarily to the varying physical properties of different DNA sequences:
For instance, adenine and thymine
are more favorably compressed into the inner minor grooves. This means
nucleosomes can bind preferentially at one position approximately every
10 base pairs (the helical repeat of DNA)- where the DNA is rotated to
maximise the number of A and T bases that will lie in the inner minor
groove.
30-nanometer chromatin fiber
Two proposed structures of the 30 nm chromatin filament.
Left: 1 start helix "solenoid" structure.
Right: 2 start loose helix structure.
Note: the histones are omitted in this diagram - only the DNA is shown.
Left: 1 start helix "solenoid" structure.
Right: 2 start loose helix structure.
Note: the histones are omitted in this diagram - only the DNA is shown.
With addition of H1, the beads-on-a-string structure in turn coils
into a 30 nm diameter helical structure known as the 30 nm fibre or
filament. The precise structure of the chromatin fiber in the cell is
not known in detail, and there is still some debate over this.
This level of chromatin structure is thought to be the form of heterochromatin,
which contains mostly transcriptionally silent genes. EM (electron
microscopy) studies have demonstrated that the 30 NM fiber is highly
dynamic such that it unfolds into a 10 nm fiber ("beads-on-a-string")
structure when transversed by an RNA polymerase engaged in
transcription.
Four proposed structures of the 30 nm chromatin filament for DNA repeat length per nucleosomes ranging from 177 to 207 bp.
Linker DNA in yellow and nucleosomal DNA in pink.
Linker DNA in yellow and nucleosomal DNA in pink.
The existing models commonly accept that the nucleosomes lie
perpendicular to the axis of the fibre, with linker histones arranged
internally.
A stable 30 nm fibre relies on the regular positioning of nucleosomes
along DNA. Linker DNA is relatively resistant to bending and rotation.
This makes the length of linker DNA critical to the stability of the
fibre, requiring nucleosomes to be separated by lengths that permit
rotation and folding into the required orientation without excessive
stress to the DNA.
In this view, different lengths of the linker DNA should produce
different folding topologies of the chromatin fiber. Recent theoretical
work, based on electron-microscopy images
of reconstituted fibers supports this view.
Spatial organization of chromatin in the cell nucleus
The
spatial arrangement of the chromatin within the nucleus is not random -
specific regions of the chromatin can be found in certain territories.
Territories are, for example, the lamina-associated domains (LADs), and the topologically associating domains (TADs), which are bound together by protein complexes. Currently, polymer models such as the Strings & Binders Switch (SBS) model and the Dynamic Loop (DL) model are used to describe the folding of chromatin within the nucleus.
Cell-cycle dependent structural organization
- Metaphase: The metaphase structure of chromatin differs vastly to that of interphase. It is optimised for physical strength[citation needed] and manageability, forming the classic chromosome structure seen in karyotypes. The structure of the condensed chromatin is thought to be loops of 30 nm fibre to a central scaffold of proteins. It is, however, not well-characterised.The physical strength of chromatin is vital for this stage of division to prevent shear damage to the DNA as the daughter chromosomes are separated. To maximise strength the composition of the chromatin changes as it approaches the centromere, primarily through alternative histone H1 analogues. During mitosis, although most of the chromatin is tightly compacted, there are small regions that are not as tightly compacted. These regions often correspond to promoter regions of genes that were active in that cell type prior to entry into chromatosis. The lack of compaction of these regions is called bookmarking, which is an epigenetic mechanism believed to be important for transmitting to daughter cells the "memory" of which genes were active prior to entry into mitosis. This bookmarking mechanism is needed to help transmit this memory because transcription ceases during mitosis.
Chromatin and bursts of transcription
Chromatin and its interaction with enzymes has been researched, and a conclusion being made is that it is relevant and an important factor in gene expression. Vincent G. Allfrey, a professor at Rockefeller University, stated that RNA synthesis is related to histone acetylation. The lysine amino acid attached to the end of the histones is positively charged. The acetylation of these tails would make the chromatin ends neutral, allowing for DNA access.When the chromatin decondenses, the DNA is open to entry of molecular machinery. Fluctuations between open and closed chromatin may contribute to the discontinuity of transcription, or transcriptional bursting. Other factors are probably involved, such as the association and dissociation of transcription factor complexes with chromatin. The phenomenon, as opposed to simple probabilistic models of transcription, can account for the high variability in gene expression occurring between cells in isogenic populations.
Alternative chromatin organizations
During metazoan spermiogenesis, the spermatid's chromatin is remodeled into a more spaced-packaged, widened, almost crystal-like structure. This process is associated with the cessation of transcription and involves nuclear protein exchange. The histones are mostly displaced, and replaced by protamines (small, arginine-rich proteins). It is proposed that in yeast, regions devoid of histones become very fragile after transcription; HMO1, an HMG-box protein, helps in stabilizing nucleosomes-free chromatin.Chromatin and DNA repair
The packaging of eukaryotic DNA into chromatin presents a barrier to all DNA-based processes that require recruitment of enzymes to their sites of action. To allow the critical cellular process of DNA repair, the chromatin must be remodeled. In eukaryotes, ATP dependent chromatin remodeling complexes and histone-modifying enzymes are two predominant factors employed to accomplish this remodeling process.Chromatin relaxation occurs rapidly at the site of a DNA damage. This process is initiated by PARP1 protein that starts to appear at DNA damage in less than a second, with half maximum accumulation within 1.6 seconds after the damage occurs. Next the chromatin remodeler Alc1 quickly attaches to the product of PARP1, and completes arrival at the DNA damage within 10 seconds of the damage. About half of the maximum chromatin relaxation, presumably due to action of Alc1, occurs by 10 seconds. This then allows recruitment of the DNA repair enzyme MRE11, to initiate DNA repair, within 13 seconds.
γH2AX, the phosphorylated form of H2AX is also involved in the early steps leading to chromatin decondensation after DNA damage occurrence. The histone variant H2AX constitutes about 10% of the H2A histones in human chromatin. γH2AX (H2AX phosphorylated on serine 139) can be detected as soon as 20 seconds after irradiation of cells (with DNA double-strand break formation), and half maximum accumulation of γH2AX occurs in one minute. The extent of chromatin with phosphorylated γH2AX is about two million base pairs at the site of a DNA double-strand break. γH2AX does not, itself, cause chromatin decondensation, but within 30 seconds of irradiation, RNF8 protein can be detected in association with γH2AX. RNF8 mediates extensive chromatin decondensation, through its subsequent interaction with CHD4, a component of the nucleosome remodeling and deacetylase complex NuRD.
After undergoing relaxation subsequent to DNA damage, followed by DNA repair, chromatin recovers to a compaction state close to its pre-damage level after about 20 min.
Methods to investigate chromatin
- ChIP-seq (Chromatin immunoprecipitation sequencing), aimed against different histone modifications, can be used to identify chromatin states throughout the genome. Different modifications have been linked to various states of chromatin.
- DNase-seq (DNase I hypersensitive sites Sequencing) uses the sensitivity of accessible regions in the genome to the DNase I enzyme to map open or accessible regions in the genome.
- FAIRE-seq (Formaldehyde-Assisted Isolation of Regulatory Elements sequencing) uses the chemical properties of protein-bound DNA in a two-phase separation method to extract nucleosome depleted regions from the genome.
- ATAC-seq (Assay for Transposable Accessible Chromatin sequencing) uses the Tn5 transposase to integrate (synthetic) transposons into accessible regions of the genome consequentially highlighting the localisation of nucleosomes and transcription factors across the genome.
- DNA footprinting is a method aimed at identifying protein-bound DNA. It uses labeling and fragmentation coupled to gel electrophoresis to identify areas of the genome that have been bound by proteins.
- MNase-seq (Micrococcal Nuclease sequencing) uses the micrococcal nuclease enzyme to identify nucleosome positioning throughout the genome.
- Chromosome conformation capture determines the spatial organization of chromatin in the nucleus, by inferring genomic locations that physically interact.
- MACC profiling (Micrococcal nuclease ACCessibility profiling) uses titration series of chromatin digests with micrococcal nuclease to identify chromatin accessibility as well as to map nucleosomes and non-histone DNA-binding proteins in both open and closed regions of the genome.
Chromatin and knots
It has been a puzzle how decondensed interphase chromosomes remain essentially unknotted. The natural expectation is that in the presence of type II DNA topoisomerases that permit passages of double-stranded DNA regions through each other, all chromosomes should reach the state of topological equilibrium. The topological equilibrium in highly crowded interphase chromosomes forming chromosome territories would result in formation of highly knotted chromatin fibres. However, Chromosome Conformation Capture (3C) methods revealed that the decay of contacts with the genomic distance in interphase chromosomes is practically the same as in the crumpled globule state that is formed when long polymers condense without formation of any knots. To remove knots from highly crowded chromatin, one would need an active process that should not only provide the energy to move the system from the state of topological equilibrium but also guide topoisomerase-mediated passages in such a way that knots would be efficiently unknotted instead of making the knots even more complex. It has been shown that the process of chromatin-loop extrusion is ideally suited to actively unknot chromatin fibres in interphase chromosomes.Chromatin: alternative definitions
The term, introduced by Walther Flemming, has multiple meanings:- Simple and concise definition: Chromatin is a macromolecular complex of a DNA macromolecule and protein macromolecules (and RNA). The proteins package and arrange the DNA and control its functions within the cell nucleus.
- A biochemists’ operational definition: Chromatin is the DNA/protein/RNA complex extracted from eukaryotic lysed interphase nuclei. Just which of the multitudinous substances present in a nucleus will constitute a part of the extracted material partly depends on the technique each researcher uses. Furthermore, the composition and properties of chromatin vary from one cell type to the another, during development of a specific cell type, and at different stages in the cell cycle.
- The DNA + histone = chromatin definition: The DNA double helix in the cell nucleus is packaged by special proteins termed histones. The formed protein/DNA complex is called chromatin. The basic structural unit of chromatin is the nucleosome.
Nobel Prizes
The following scientists were recognized for their contributions to chromatin research with Nobel Prizes:| Year | Who | Award |
|---|---|---|
| 1910 | Albrecht Kossel (University of Heidelberg) | Nobel Prize in Physiology or Medicine for his discovery of the five nuclear bases: adenine, cytosine, guanine, thymine, and uracil. |
| 1933 | Thomas Hunt Morgan (California Institute of Technology) | Nobel Prize in Physiology or Medicine for his discoveries of the role played by the gene and chromosome in heredity, based on his studies of the white-eyed mutation in the fruit fly Drosophila. |
| 1962 | Francis Crick, James Watson and Maurice Wilkins (MRC Laboratory of Molecular Biology, Harvard University and London University respectively) | Nobel Prize in Physiology or Medicine for their discoveries of the double helix structure of DNA and its significance for information transfer in living material. |
| 1982 | Aaron Klug (MRC Laboratory of Molecular Biology) | Nobel Prize in Chemistry "for his development of crystallographic electron microscopy and his structural elucidation of biologically important nucleic acid-protein complexes" |
| 1993 | Richard J. Roberts and Phillip A. Sharp | Nobel Prize in Physiology "for their independent discoveries of split genes," in which DNA sections called exons express proteins, and are interrupted by DNA sections called introns, which do not express proteins. |
| 2006 | Roger Kornberg (Stanford University) | Nobel Prize in Chemistry for his discovery of the mechanism by which DNA is transcribed into messenger RNA. |