
AMMs have diversified rapidly over the past few decades. A major point is to exploit existing motion in proteins, such as rotation about single bonds or cis-trans isomerization. Different AMMs are produced by introducing various functionalities, such as the introduction of bistability to create switches. A broad range of AMMs has been designed, featuring different properties and applications; some of these include molecular motors, switches, and logic gates. A wide range of applications have been demonstrated for AMMs, including those integrated into polymeric, liquid crystal, and crystalline systems for varied functions (such as materials research, homogenous catalysis and surface chemistry).
Terminology
Several definitions describe a "molecular machine" as a class of molecules typically described as an assembly of a discrete number of molecular components intended to produce mechanical movements in response to specific stimuli. The expression is often more generally applied to molecules that simply mimic functions that occur at the macroscopic level. A few prime requirements for a molecule to be considered a "molecular machine" are: the presence of moving parts, the ability to consume energy, and the ability to perform a task. Molecular machines differ from other stimuli-responsive compounds that can produce motion (such as cis-trans isomers) in their relatively larger amplitude of movement (potentially due to chemical reactions) and the presence of a clear external stimulus to regulate the movements (as compared to random thermal motion). Piezoelectric, magnetostrictive, and other materials that produce a movement due to external stimuli on a macro-scale are generally not included, since despite the molecular origin of the motion the effects are not useable on the molecular scale.
This definition generally applies to synthetic molecular machines, which have historically gained inspiration from the naturally occurring biological molecular machines (also referred to as "nanomachines"). Biological machines are considered to be nanoscale devices (such as molecular proteins) in a living system that convert various forms of energy to mechanical work in order to drive crucial biological processes such as intracellular transport, muscle contractions, ATP generation and cell division.
History
What would be the utility of such machines? Who knows? I cannot see exactly what would happen, but I can hardly doubt that when we have some control of the arrangement of things on a molecular scale we will get an enormously greater range of possible properties that substances can have, and of the different things we can do.
Biological molecular machines have been known and studied for decades given their vital role in sustaining life, and have served as inspiration for synthetically designed systems with similar useful functionality. The advent of conformational analysis, or the study of conformers to analyze complex chemical structures, in the 1950s gave rise to the idea of understanding and controlling relative motion within molecular components for further applications. This led to the design of "proto-molecular machines" featuring conformational changes such as cog-wheeling of the aromatic rings in triptycenes. By 1980, scientists could achieve desired conformations using external stimuli and utilize this for different applications. A major example is the design of a photoresponsive crown ether containing an azobenzene unit, which could switch between cis and trans isomers on exposure to light and hence tune the cation-binding properties of the ether. In his seminal 1959 lecture There's Plenty of Room at the Bottom, Richard Feynman alluded to the idea and applications of molecular devices designed artificially by manipulating matter at the atomic level. This was further substantiated by Eric Drexler during the 1970s, who developed ideas based on molecular nanotechnology such as nanoscale "assemblers", though their feasibility was disputed.

Though these events served as inspiration for the field, the actual breakthrough in practical approaches to synthesize artificial molecular machines (AMMs) took place in 1991 with the invention of a "molecular shuttle" by Sir Fraser Stoddart. Building upon the assembly of mechanically linked molecules such as catenanes and rotaxanes as developed by Jean-Pierre Sauvage in the early 1980s, this shuttle features a rotaxane with a ring that can move across an "axle" between two ends or possible binding sites (hydroquinone units). This design realized the well-defined motion of a molecular unit across the length of the molecule for the first time. In 1994, an improved design allowed control over the motion of the ring by pH variation or electrochemical methods, making it the first example of an AMM. Here the two binding sites are a benzidine and a biphenol unit; the cationic ring typically prefers staying over the benzidine ring, but moves over to the biphenol group when the benzidine gets protonated at low pH or if it gets electrochemically oxidized. In 1998, a study could capture the rotary motion of a decacyclene molecule on a copper-base metallic surface using a scanning tunneling microscope. Over the following decade, a broad variety of AMMs responding to various stimuli were invented for different applications. In 2016, the Nobel Prize in Chemistry was awarded to Sauvage, Stoddart, and Bernard L. Feringa for the design and synthesis of molecular machines.
Artificial molecular machines
Over the past few decades, AMMs have diversified rapidly and their design principles, properties, and characterization methods have been outlined more clearly. A major starting point for the design of AMMs is to exploit the existing modes of motion in molecules. For instance, single bonds can be visualized as axes of rotation, as can be metallocene complexes. Bending or V-like shapes can be achieved by incorporating double bonds, that can undergo cis-trans isomerization in response to certain stimuli (typically irradiation with a suitable wavelength), as seen in numerous designs consisting of stilbene and azobenzene units. Similarly, ring-opening and -closing reactions such as those seen for spiropyran and diarylethene can also produce curved shapes. Another common mode of movement is the circumrotation of rings relative to one another as observed in mechanically interlocked molecules (primarily catenanes). While this type of rotation can not be accessed beyond the molecule itself (because the rings are confined within one another), rotaxanes can overcome this as the rings can undergo translational movements along a dumbbell-like axis. Another line of AMMs consists of biomolecules such as DNA and proteins as part of their design, making use of phenomena like protein folding and unfolding.
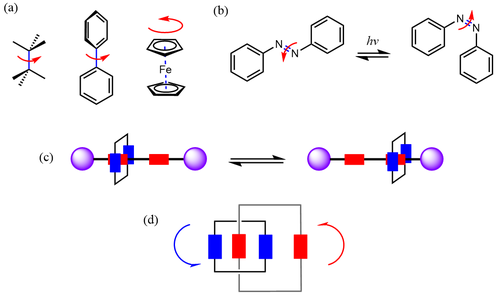
AMM designs have diversified significantly since the early days of the field. A major route is the introduction of bistability to produce molecular switches, featuring two distinct configurations for the molecule to convert between. This has been perceived as a step forward from the original molecular shuttle which consisted of two identical sites for the ring to move between without any preference, in a manner analogous to the ring flip in an unsubstituted cyclohexane. If these two sites are different from each other in terms of features like electron density, this can give rise to weak or strong recognition sites as in biological systems — such AMMs have found applications in catalysis and drug delivery. This switching behavior has been further optimized to acquire useful work that gets lost when a typical switch returns to its original state. Inspired by the use of kinetic control to produce work in natural processes, molecular motors are designed to have a continuous energy influx to keep them away from equilibrium to deliver work.
Various energy sources are employed to drive molecular machines today, but this was not the case during the early years of AMM development. Though the movements in AMMs were regulated relative to the random thermal motion generally seen in molecules, they could not be controlled or manipulated as desired. This led to the addition of stimuli-responsive moieties in AMM design, so that externally applied non-thermal sources of energy could drive molecular motion and hence allow control over the properties. Chemical energy (or "chemical fuels") was an attractive option at the beginning, given the broad array of reversible chemical reactions (heavily based on acid-base chemistry) to switch molecules between different states. However, this comes with the issue of practically regulating the delivery of the chemical fuel and the removal of waste generated to maintain the efficiency of the machine as in biological systems. Though some AMMs have found ways to circumvent this, more recently waste-free reactions such based on electron transfers or isomerization have gained attention (such as redox-responsive viologens). Eventually, several different forms of energy (electric, magnetic, optical and so on) have become the primary energy sources used to power AMMs, even producing autonomous systems such as light-driven motors.
Types
Various AMMs are tabulated below along with indicative images:
| Type | Details | Image |
|---|---|---|
| Molecular balance | A molecule that can interconvert between two or more conformational or configurational states in response to the dynamic of multiple intra- and intermolecular driving forces, such as hydrogen bonding, solvophobic or hydrophobic effects, π interactions, and steric and dispersion interactions. The distinct conformers of a molecular balance can show different interactions with the same molecule, such that analyzing the ratio of the conformers and the energies for these interactions can enable quantification of different properties (such as CH-π or arene-arene interactions, see image). | 
|
| Molecular hinge | A molecular hinge is a molecule that can typically rotate in a crank-like motion around a rigid axis, such as a double bond or aromatic ring, to switch between reversible configurations. Such configurations must have distinguishable geometries; for instance, azobenzene groups in a linear molecule may undergo cis-trans isomerization when irradiated with ultraviolet light, triggering a reversible transition to a bent or V-shaped conformation (see image). Molecular hinges have been adapted for applications such as nucleobase recognition, peptide modifications, and visualizing molecular motion. | 
|
| Molecular logic gate | A molecule that performs a logical operation on one or more logic inputs and produces a single logic output. Modelled on logic gates, these molecules have slowly replaced the conventional silicon-based machinery. Several applications have come forth, such as water quality examination, food safety examination, metal ion detection, and pharmaceutical studies. The first example of a molecular logic gate was reported in 1993, featuring a receptor (see image) where the emission intensity could be treated as a tunable output if the concentrations of protons and sodium ions were to be considered as inputs. | 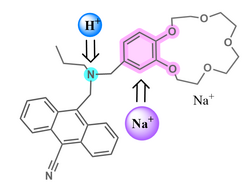
|
| Molecular motor | A molecule that is capable of directional rotary motion around a single or double bond and produce useful work as a result (as depicted in the image). Carbon nanotube nanomotors have also been produced. Single bond rotary motors are generally activated by chemical reactions whereas double bond rotary motors are generally fueled by light. The rotation speed of the motor can also be tuned by careful molecular design. | 
|
| Molecular necklace | A class of mechanically interlocked molecules derived from catenanes where a large macrocycle backbone connects at least three small rings in the shape of a necklace (see image for example). A molecular necklace consisting of a large macrocycle threaded by n-1 rings (hence comprising n rings) is represented as [n]MN. The first molecular necklace was synthesized in 1992, featuring several α-cyclodextrins on a single polyethylene glycol chain backbone; the authors connected this to the idea of a "molecular abacus" proposed by Stoddart and coworkers around the same time. Several interesting applications have emerged for these molecules, such as antibacterial activity, desulfurization of fuels, and piezoelectricity. | 
|
| Molecular propeller | A molecule that can propel fluids when rotated, due to its special shape that is designed in analogy to macroscopic propellers (see schematic image on right). It has several molecular-scale blades attached at a certain pitch angle around the circumference of a nanoscale shaft. Propellers have been shown to have interesting properties, such as variations in pumping rates for hydrophilic and hydrophobic fluids. | 
|
| Molecular shuttle | A molecule capable of shuttling molecules or ions from one location to another. This is schematically depicted in the image on the right, where a ring (in green) can bind to either one of the yellow sites on the blue macrocyclic backbone. A common molecular shuttle consists of a rotaxane where the macrocycle can move between two sites or stations along the dumbbell backbone; controlling the properties of either site and by regulating conditions like pH can enable control over which site is selected for binding. This has led to novel applications in catalysis and drug delivery. | 
|
| Molecular switch | A molecule that can be reversibly shifted between two or more stable states in response to certain stimuli. This change of states influences the properties of the molecule according to the state it occupies at the moment. Unlike a molecular motor, any mechanical work done due to the motion in a switch is generally undone once the molecule returns to its original state unless it is part of a larger motor-like system. The image on the right shows a hydrazone-based switch that switches in response to pH changes. | 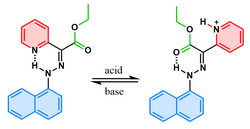
|
| Molecular tweezers | Host molecules capable of holding items between their two arms. The open cavity of the molecular tweezers binds items using non-covalent bonding including hydrogen bonding, metal coordination, hydrophobic forces, van der Waals forces, π interactions, or electrostatic effects. For instance, the image on the right depicts tweezers formed by corannulene pincers clasping a C60 fullerene molecule, termed "buckycatcher". Examples of molecular tweezers have been reported that are constructed from DNA and are considered DNA machines. | 
|
| Nanocar | Single-molecule vehicles that resemble macroscopic automobiles and are important for understanding how to control molecular diffusion on surfaces. The image on the right shows an example with wheels made of fullerene molecules. The first nanocars were synthesized by James M. Tour in 2005. They had an H-shaped chassis and 4 molecular wheels (fullerenes) attached to the four corners. In 2011, Feringa and co-workers synthesized the first motorized nanocar which had molecular motors attached to the chassis as rotating wheels. The authors were able to demonstrate directional motion of the nanocar on a copper surface by providing energy from a scanning tunneling microscope tip. Later, in 2017, the world's first-ever nanocar race took place in Toulouse. | 
|
Biological molecular machines
Many macromolecular machines are found within cells, often in the form of multi-protein complexes. Examples of biological machines include motor proteins such as myosin, which is responsible for muscle contraction, kinesin, which moves cargo inside cells away from the nucleus along microtubules, and dynein, which moves cargo inside cells towards the nucleus and produces the axonemal beating of motile cilia and flagella. "[I]n effect, the [motile cilium] is a nanomachine composed of perhaps over 600 proteins in molecular complexes, many of which also function independently as nanomachines ... Flexible linkers allow the mobile protein domains connected by them to recruit their binding partners and induce long-range allostery via protein domain dynamics." Other biological machines are responsible for energy production, for example ATP synthase which harnesses energy from proton gradients across membranes to drive a turbine-like motion used to synthesise ATP, the energy currency of a cell. Still other machines are responsible for gene expression, including DNA polymerases for replicating DNA, RNA polymerases for producing mRNA, the spliceosome for removing introns, and the ribosome for synthesising proteins. These machines and their nanoscale dynamics are far more complex than any molecular machines that have yet been artificially constructed.
Biological machines have potential applications in nanomedicine. For example, they could be used to identify and destroy cancer cells. Molecular nanotechnology is a speculative subfield of nanotechnology regarding the possibility of engineering molecular assemblers, biological machines which could re-order matter at a molecular or atomic scale.[citation needed] Nanomedicine would make use of these nanorobots, introduced into the body, to repair or detect damages and infections, but these are considered to be far beyond current capabilities.
Research and applications
Advances in this area are inhibited by the lack of synthetic methods. In this context, theoretical modeling has emerged as a pivotal tool to understand the self-assembly or -disassembly processes in these systems.
Possible applications have been demonstrated for AMMs, including those integrated into polymeric, liquid crystal, and crystalline systems for varied functions. Homogenous catalysis is a prominent example, especially in areas like asymmetric synthesis, utilizing noncovalent interactions and biomimetic allosteric catalysis. AMMs have been pivotal in the design of several stimuli-responsive smart materials, such as 2D and 3D self-assembled materials and nanoparticle-based systems, for versatile applications ranging from 3D printing to drug delivery.
AMMs are gradually moving from the conventional solution-phase chemistry to surfaces and interfaces. For instance, AMM-immobilized surfaces (AMMISs) are a novel class of functional materials consisting of AMMs attached to inorganic surfaces forming features like self-assembled monolayers; this gives rise to tunable properties such as fluorescence, aggregation and drug-release activity.
Most of these "applications" remain at the proof-of-concept level. Challenges in streamlining macroscale applications include autonomous operation, the complexity of the machines, stability in the synthesis of the machines and the working conditions.










