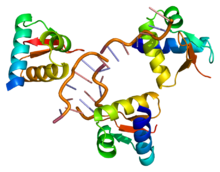
The
horseshoe shaped ribonuclease inhibitor (shown as wireframe) forms a
protein–protein interaction with the ribonuclease protein. The contacts
between the two proteins are shown as coloured patches.
Protein–protein interactions (PPIs) are the physical contacts of high specificity established between two or more protein molecules as a result of biochemical events steered by electrostatic forces including the hydrophobic effect.
Many are physical contacts with molecular associations between chains
that occur in a cell or in a living organism in a specific biomolecular
context.
Proteins rarely act alone as their functions tend to be
regulated. Many molecular processes within a cell are carried out by
molecular machines that are built from a large number of protein
components organized by their PPIs. These interactions make up the
so-called interactomics of the organism, while aberrant PPIs are the basis of multiple aggregation-related diseases, such as Creutzfeldt–Jakob, Alzheimer's diseases, and may lead to cancer.
PPIs have been studied from different perspectives: biochemistry, quantum chemistry, molecular dynamics, signal transduction, among others. All this information enables the creation of large protein interaction networks – similar to metabolic or genetic/epigenetic networks – that empower the current knowledge on biochemical cascades and molecular etiology of disease, as well as the discovery of putative protein targets of therapeutic interest.
Examples
Electron transfer proteins
In many metabolic reactions, a protein that acts as an electron carrier binds to an enzyme that acts its reductase. After it receives an electron, it dissociates and then binds to the next enzyme that acts its oxidase
(i.e. an acceptor of the electron). These interactions between proteins
are dependent on highly specific binding between proteins to ensure
efficient electron transfer. Examples: mitochondrial oxidative
phosphorylation chain system components cytochrome c-reductase / cytochrome c / cytochrome c oxidase; microsomal and mitochondrial P450 systems.
In the case of the mitochondrial P450 systems, the specific residues involved in the binding of the electron transfer protein adrenodoxin
to its reductase were identified as two basic Arg residues on the
surface of the reductase and two acidic Asp residues on the adrenodoxin.
More recent work on the phylogeny of the reductase has shown that these
residues involved in protein-protein interactions have been conserved
throughout the evolution of this enzyme.
Signal transduction
The
activity of the cell is regulated by extracellular signals. Signal
propagation inside and/or along the interior of cells depends on PPIs
between the various signaling molecules. The recruitment of signaling
pathways through PPIs is called signal transduction and plays a fundamental role in many biological processes and in many diseases including Parkinson's disease and cancer.
Membrane transport
A protein may be carrying another protein (for example, from cytoplasm to nucleus or vice versa in the case of the nuclear pore importins).
Cell metabolism
In many biosynthetic processes enzymes interact with each other to produce small compounds or other macromolecules.
Muscle contraction
Physiology of muscle contraction involves several interactions. Myosin filaments act as molecular motors and by binding to actin enables filament sliding. Furthermore, members of the skeletal muscle lipid droplet-associated proteins family associate with other proteins, as activator of adipose triglyceride lipase and its coactivator comparative gene identification-58, to regulate lipolysis in skeletal muscle.
Types
To describe the types of protein–protein interactions (PPIs) it is
important to consider that proteins can interact in a "transient" way
(to produce some specific effect in a short time) or to interact with
other proteins in a "stable" way to build multiprotein complexes that
are molecular machines within the living systems. A protein complex
assembly can result in the formation of homo-oligomeric or hetero-oligomeric complexes.
In addition to the conventional complexes, as enzyme-inhibitor and
antibody-antigen, interactions can also be established between
domain-domain and domain-peptide. Another important distinction to
identify protein-protein interactions is the way they have been
determined, since there are techniques that measure direct physical
interactions between protein pairs, named “binary” methods, while there
are other techniques that measure physical interactions among groups of
proteins, without pairwise determination of protein partners, named
“co-complex” methods.
Homo-oligomers vs. hetero-oligomers
Homo-oligomers are macromolecular complexes constituted by only one type of protein subunit. Protein subunits assembly is guided by the establishment of non-covalent interactions in the quaternary structure of the protein. Disruption of homo-oligomers in order to return to the initial individual monomers often requires denaturation of the complex. Several enzymes, carrier proteins,
scaffolding proteins, and transcriptional regulatory factors carry out
their functions as homo-oligomers.
Distinct protein subunits interact in hetero-oligomers, which are
essential to control several cellular functions. The importance of the
communication between heterologous proteins is even more evident during
cell signaling events and such interactions are only possible due to
structural domains within the proteins (as described below).
Stable interactions vs. transient interactions
Stable
interactions involve proteins that interact for a long time, taking
part of permanent complexes as subunits, in order to carry out
structural or functional roles. These are usually the case of
homo-oligomers (e.g. cytochrome c), and some hetero-oligomeric proteins, as the subunits of ATPase. On the other hand, a protein may interact briefly and in a reversible manner with other proteins in only certain cellular contexts – cell type, cell cycle stage, external factors, presence of other binding proteins, etc. – as it happens with most of the proteins involved in biochemical cascades. These are called transient interactions. For example, some G protein-coupled receptors only transiently bind to Gi/o proteins when they are activated by extracellular ligands, while some Gq-coupled receptors, such as muscarinic receptor M3, pre-couple with Gq proteins prior to the receptor-ligand binding. Interactions between intrinsically disordered protein regions to globular protein domains (i.e. MoRFs) are transient interactions.
Covalent vs. non-covalent
Covalent interactions are those with the strongest association and are formed by disulphide bonds or electron sharing. Although being rare, these interactions are determinant in some posttranslational modifications, as ubiquitination and SUMOylation. Non-covalent bonds are usually established during transient interactions by the combination of weaker bonds, such as hydrogen bonds, ionic interactions, Van der Waals forces, or hydrophobic bonds.
Role of water
Water molecules play a significant role in the interactions between proteins.
The crystal structures of complexes, obtained at high resolution from
different but homologous proteins, have shown that some interface water
molecules are conserved between homologous complexes. The majority of
the interface water molecules make hydrogen bonds with both partners of
each complex. Some interface amino acid residues or atomic groups of one
protein partner engage in both direct and water mediated interactions
with the other protein partner. Doubly indirect interactions, mediated
by two water molecules, are more numerous in the homologous complexes of
low affinity.
Carefully conducted mutagenesis experiments, e.g. changing a tyrosine
residue into a phenylalanine, have shown that water mediated
interactions can contribute to the energy of interaction. Thus, water molecules may facilitate the interactions and cross-recognitions between proteins.
Structure
Crystal structure of modified Gramicidin S horizontally determined by X-ray crystallography
NMR structure of cytochrome C illustrating its dynamics in solution
The molecular structures of many protein complexes have been unlocked by the technique of X-ray crystallography. The first structure to be solved by this method was that of sperm whale myoglobin by Sir John Cowdery Kendrew.
In this technique the angles and intensities of a beam of X-rays
diffracted by crystalline atoms are detected in a film, thus producing a
three-dimensional picture of the density of electrons within the
crystal.
Later, nuclear magnetic resonance
also started to be applied with the aim of unravelling the molecular
structure of protein complexes. One of the first examples was the
structure of calmodulin-binding domains bound to calmodulin.
This technique is based on the study of magnetic properties of atomic
nuclei, thus determining physical and chemical properties of the
correspondent atoms or the molecules. Nuclear magnetic resonance is
advantageous for characterizing weak PPIs.
Domains
Proteins hold structural domains that allow their interaction with and bind to specific sequences on other proteins:
- Src homology 2 (SH2) domain
- SH2 domains are structurally composed by three-stranded twisted beta sheet sandwiched flanked by two alpha-helices. The existence of a deep binding pocket with high affinity for phosphotyrosine, but not for phosphoserine or phosphothreonine, is essential for the recognition of tyrosine phosphorylated proteins, mainly autophosphorylated growth factor receptors. Growth factor receptor binding proteins and phospholipase Cγ are examples of proteins that have SH2 domains.
- Src homology 3 (SH3) domain
- Structurally, SH3 domains are constituted by a beta barrel formed by two orthogonal beta sheets and three anti-parallel beta strands. These domains recognize proline enriched sequences, as polyproline type II helical structure (PXXP motifs)[verification needed] in cell signaling proteins like protein tyrosine kinases and the growth factor receptor bound protein 2 (Grb2).
- Phosphotyrosine-binding (PTB) domain
- PTB domains interact with sequences that contain a phosphotyrosine group. These domains can be found in the insulin receptor substrate.
- LIM domain
- LIM domains were initially identified in three homeodomain transcription factors (lin11, is11, and mec3). In addition to this homeodomain proteins and other proteins involved in development, LIM domains have also been identified in non-homeodomain proteins with relevant roles in cellular differentiation, association with cytoskeleton and senescence. These domains contain a tandem cysteine-rich Zn2+-finger motif and embrace the consensus sequence CX2CX16-23HX2CX2CX2CX16-21CX2C/H/D. LIM domains bind to PDZ domains, bHLH transcription factors, and other LIM domains.
- Sterile alpha motif (SAM) domain
- SAM domains are composed by five helices forming a compact package with a conserved hydrophobic core. These domains, which can be found in the Eph receptor and the stromal interaction molecule (STIM) for example, bind to non-SAM domain-containing proteins and they also appear to have the ability to bind RNA.
- PDZ domain
- PDZ domains were first identified in three guanylate kinases: PSD-95, DlgA and ZO-1. These domains recognize carboxy-terminal tri-peptide motifs (S/TXV), other PDZ domains or LIM domains and bind them through a short peptide sequence that has a C-terminal hydrophobic residue. Some of the proteins identified as having PDZ domains are scaffolding proteins or seem to be involved in ion receptor assembling and receptor-enzyme complexes formation.
- FERM domain
- FERM domains contain basic residues capable of binding PtdIns(4,5)P2. Talin and focal adhesion kinase (FAK) are two of the proteins that present FERM domains.
- Calponin homology (CH) domain
- CH domains are mainly present in cytoskeletal proteins as parvin.
- Pleckstrin homology domain
- Pleckstrin homology domains bind to phosphoinositides and acid domains in signaling proteins.
- WW domain
- WW domains bind to proline enriched sequences.
- WSxWS motif
- Found in cytokine receptors
Properties of the interface
The
study of the molecular structure can give fine details about the
interface that enables the interaction between proteins. When
characterizing PPI interfaces it is important to take into account the
type of complex.
Parameters evaluated include size (measured in absolute dimensions Å2 or in solvent-accessible surface area (SASA)),
shape, complementarity between surfaces, residue interface
propensities, hydrophobicity, segmentation and secondary structure, and
conformational changes on complex formation.
The great majority of PPI interfaces reflects the composition of
protein surfaces, rather than the protein cores, in spite of being
frequently enriched in hydrophobic residues, particularly in aromatic
residues. PPI interfaces are dynamic and frequently planar, although they can be globular and protruding as well. Based on three structures – insulin dimer, trypsin-pancreatic trypsin inhibitor complex, and oxyhaemoglobin – Cyrus Chothia and Joel Janin found that between 1,130 and 1,720 Å2 of surface area was removed from contact with water indicating that hydrophobicity is a major factor of stabilization of PPIs. Later studies refined the buried surface area of the majority of interactions to 1,600±350 Å2. However, much larger interaction interfaces were also observed and were associated with significant changes in conformation of one of the interaction partners. PPIs interfaces exhibit both shape and electrostatic complementarity.
Regulation
- Protein concentration, which in turn are affected by expression levels and degradation rates;
- Protein affinity for proteins or other binding ligands;
- Ligands concentrations (substrates, ions, etc.);
- Presence of other proteins, nucleic acids, and ions;
- Electric fields around proteins;
- Occurrence of covalent modifications.
Experimental Methods
There are a multitude of methods to detect them. Each of the approaches has its own strengths and weaknesses, especially with regard to the sensitivity and specificity of the method. The most conventional and widely used high-throughput methods are yeast two-hybrid screening and affinity purification coupled to mass spectrometry.
Principles of yeast and mammalian two-hybrid systems
Yeast two-hybrid screening
This system was firstly described in 1989 by Fields and Song using Saccharomyces cerevisiae as biological model. Yeast two hybrid allows the identification of pairwise PPIs (binary method) in vivo,
in which the two proteins are tested for biophysically direct
interaction. The Y2H is based on the functional reconstitution of the
yeast transcription factor Gal4 and subsequent activation of a selective
reporter such as His3. To test two proteins for interaction, two
protein expression constructs are made: one protein (X) is fused to the
Gal4 DNA-binding domain (DB) and a second protein (Y) is fused to the
Gal4 activation domain (AD). In the assay, yeast cells are transformed
with these constructs. Transcription of reporter genes does not occur
unless bait (DB-X) and prey (AD-Y) interact with each other and form a
functional Gal4 transcription factor. Thus, the interaction between
proteins can be inferred by the presence of the products resultant of
the reporter gene expression.
In cases in which the reporter gene expresses enzymes that allow the
yeast to synthesize essential amino acids or nucleotides, yeast growth
under selective media conditions indicates that the two proteins tested
are interacting.
Despite its usefulness, the yeast two-hybrid system has
limitations. It uses yeast as main host system, which can be a problem
when studying proteins that contain mammalian-specific
post-translational modifications. The number of PPIs identified is
usually low because of a high false negative rate; and, understates membrane proteins, for example.
In initial studies that utilized Y2H, proper controls for false
positives (e.g. when DB-X activates the reporter gene without the
presence of AD-Y) were frequently not done, leading to a higher than
normal false positive rate. An empirical framework must be implemented
to control for these false positives.
Limitations in lower coverage of membrane proteins have been overcoming
by the emergence of yeast two-hybrid variants, such as the membrane
yeast two-hybrid (MYTH) and the split-ubiquitin system, which are not limited to interactions that occur in the nucleus; and, the bacterial two-hybrid system, performed in bacteria.
Principle of tandem affinity purification
Affinity purification coupled to mass spectrometry
Affinity purification coupled to mass spectrometry mostly detects
stable interactions and thus better indicates functional in vivo PPIs. This method starts by purification of the tagged protein, which is expressed in the cell usually at in vivo
concentrations, and its interacting proteins (affinity purification).
One of the most advantageous and widely used method to purify proteins
with very low contaminating background is the tandem affinity purification,
developed by Bertrand Seraphin and Matthias Mann and respective
colleagues. PPIs can then be quantitatively and qualitatively analysed
by mass spectrometry using different methods: chemical incorporation,
biological or metabolic incorporation (SILAC), and label-free methods.
Nucleic acid programmable protein array
This
system was first developed by LaBaer and colleagues in 2004 by using in
vitro transcription and translation system. They use DNA template
encoding the gene of interest fused with GST protein, and it was
immobilized in the solid surface. Anti-GST antibody and biotinylated
plasmid DNA were bounded in aminopropyltriethoxysilane (APTES)-coated
slide. BSA can improve the binding efficiency of DNA. Biotinylated
plasmid DNA was bound by avidin. New protein was synthesized by using
cell-free expression system i.e. rabbit reticulocyte lysate (RRL), and
then the new protein was captured through anti-GST antibody bounded on
the slide. To test protein-protein interaction, the targeted protein
cDNA and query protein cDNA were immobilized in a same coated slide. By
using in vitro transcription and translation system, targeted and query
protein was synthesized by the same extract. The targeted protein was
bound to array by antibody coated in the slide and query protein was
used to probe the array. The query protein was tagged with hemagglutinin
(HA) epitope. Thus, the interaction between the two proteins was
visualized with the antibody against HA.
Other potential methods
Diverse techniques to identify PPIs have been emerging along with technology progression. These include co-immunoprecipitation, protein microarrays, analytical ultracentrifugation, light scattering, fluorescence spectroscopy,
luminescence-based mammalian interactome mapping (LUMIER),
resonance-energy transfer systems, mammalian protein–protein interaction
trap, electro-switchable biosurfaces, protein-fragment complementation assay, as well as real-time label-free measurements by surface plasmon resonance, and calorimetry.
Computational methods
Computational Prediction of Protein-Protein Interactions
The
experimental detection and characterization of PPIs is labor intensive
and time-consuming. However, many PPIs can be also predicted
computationally, usually using experimental data as a starting point.
However, methods have also been developed that allow the prediction of
PPI de novo, that is without prior evidence for these interactions.
Genomic Context Methods
The Rosetta Stone or Domain Fusion method is based on the hypothesis that interacting proteins are sometimes fused into a single protein in another genome.
Therefore, we can predict if two proteins may be interacting by
determining if they each have non-overalaping sequence similarity to a
region of a single protein sequence in another genome.
The Conserved Neighborhood method
is based on the hypothesis that if genes encoding two proteins are
neighbors on a chromosome in many genomes, then they are likely
functionally related (and possibly physically interacting).
The Phylogenetic Profile method
is based on the hypothesis that if two or more proteins are
concurrently present or absent across several genomes, then they are
likely functionally related.
Therefore, potentially interacting proteins can be identified by
determining the presence or absence of genes across many genomes and
selecting those genes which are always present or absent together.
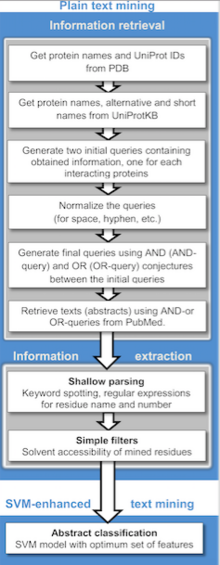
Text mining methods
Text mining protocol.
Publicly available information from biomedical documents is readily
accessible through the internet and is becoming a powerful resource for
collecting known protein-protein interactions (PPIs), PPI prediction and
protein docking. Text mining is much less costly and time-consuming
compared to other high-throughput techniques. Currently, text mining
methods generally detect binary relations between interacting proteins
from individual sentences using rule/pattern-based information
extraction and machine learning approaches.
A wide variety of text mining applications for PPI extraction and/or
prediction are available for public use, as well as repositories which
often store manually validated and/or computationally predicted PPIs.
Text mining can be implemented in two stages: information retrieval, where texts containing names of either or both interacting proteins are retrieved and information extraction, where targeted information (interacting proteins, implicated residues, interaction types, etc.) is extracted.
There are also studies using phylogenetic profiling, basing their
functionalities on the theory that proteins involved in common pathways
co-evolve in a correlated fashion across species. Some more complex
text mining methodologies use advanced Natural Language Processing
(NLP) techniques and build knowledge networks (for example, considering
gene names as nodes and verbs as edges). Other developments involve
kernel methods to predict protein interactions.
Machine learning methods
These
methods use machine learning to distinguish how interacting protein
pairs differ from non-interacting protein pairs in terms of pairwise
features such as cellular colocalization, gene co-expression, how
closely located on a DNA are the genes that encode the two proteins, and
so on. Random Forest has been found to be most-effective machine learning method for protein interaction prediction. Such methods have been applied for discovering protein interactions on human interactome, specifically the interactome of Membrane proteins and the interactome of Schizophrenia-associated proteins.
Databases
Large
scale identification of PPIs generated hundreds of thousands of
interactions, which were collected together in specialized biological databases that are continuously updated in order to provide complete interactomes. The first of these databases was the Database of Interacting Proteins (DIP).
Since that time, the number of public databases has been increasing.
Databases can be subdivided into primary databases, meta-databases, and
prediction databases.
Primary databases collect information about published PPIs proven to exist via small-scale or large-scale experimental methods. Examples: DIP, Biomolecular Interaction Network Database (BIND), Biological General Repository for Interaction Datasets (BioGRID),
Human Protein Reference Database (HPRD), IntAct Molecular Interaction
Database, Molecular Interactions Database (MINT), MIPS Protein
Interaction Resource on Yeast (MIPS-MPact), and MIPS Mammalian
Protein–Protein Interaction Database (MIPS-MPPI).
Meta-databases normally result from the integration of
primary databases information, but can also collect some original data.
Examples: Agile Protein Interactomes Dataserver (APID), The Microbial Protein Interaction Database (MPIDB), Protein Interaction Network Analysis (PINA) platform, (GPS-Prot), and Wiki-Pi.
Prediction databases include many PPIs that are predicted
using several techniques (main article). Examples: Human Protein–Protein
Interaction Prediction Database (PIPs), Interlogous Interaction Database (I2D), Known and Predicted Protein–Protein Interactions (STRING-db), and Unified Human Interactive (UniHI).
The aforementioned computational methods all depend on source
databases whose data can be extrapolated to predict novel
protein-protein interactions. Coverage differs greatly between
databases. In general, primary databases have the fewest total protein
interactions recorded as they do not integrate data from multiple other
databases, while prediction databases have the most because they include
other forms of evidence in addition to experimental. For example, the
primary database IntAct has 572,063 interactions, the meta-database APID has 678,000 interactions, and the predictive database STRING has 25,914,693 interactions.
However, it is important to note that some of the interactions in the
STRING database are only predicted by computational methods such as
Genomic Context and not experimentally verified.
Interaction networks
Schizophrenia PPI.
Information found in PPIs databases supports the construction of
interaction networks. Although the PPI network of a given query protein
can be represented in textbooks, diagrams of whole cell PPIs are frankly
complex and difficult to generate.
One example of a manually produced molecular interaction map is the Kurt Kohn's 1999 map of cell cycle control.
Drawing on Kohn's map, Schwikowski et al. in 2000 published a paper on
PPIs in yeast, linking 1,548 interacting proteins determined by
two-hybrid screening. They used a layered graph drawing method to find
an initial placement of the nodes and then improved the layout using a
force-based algorithm.
Bioinformatic tools have been developed to simplify the difficult
task of visualizing molecular interaction networks and complement them
with other types of data. For instance, Cytoscape is an open-source software widely used and lots of plugins are currently available. Pajek software is advantageous for the visualization and analysis of very large networks.
Identification of functional modules in PPI networks is an
important challenge in bioinformatics. Functional modules means a set of
proteins that are highly connected to each other in PPI network. It is almost similar problem as community detection in social networks. There are some methods such as Jactive modules and MoBaS. Jactive modules integrate PPI network and gene expression data where as MoBaS integrate PPI network and Genome Wide association Studies.
The awareness of the major roles of PPIs in numerous
physiological and pathological processes has been driving the challenge
of unravel many interactomes. Examples of published interactomes are the
thyroid specific DREAM interactome and the PP1α interactome in human brain.
Protein-protein relationships are often the result of multiple
types of interactions or are deduced from different approaches,
including co-localization, direct interaction, suppressive genetic
interaction, additive genetic interaction, physical association, and
other associations.
Signed interaction networks
The
protein protein interactions are displayed in a signed network that
describes what type of interactions that are taking place
Protein–protein interactions often result in one of the interacting
proteins either being 'activated' or 'repressed'. Such effects can be
indicated in a PPI network by "signs" (e.g. "activation" or
"inhibition"). Although such attributes have been added to networks for a
long time, Vinayagam et al. (2014) coined the term Signed network
for them. Signed networks are often expressed by labeling the
interaction as either positive or negative. A positive interaction is
one where the interaction results in one of the proteins being
activated. Conversely a negative interaction indicates that one of the
proteins being inactivated.
Protein–protein interaction networks are often constructed as a
result of lab experiments such as yeast two hybrid screens or 'affinity
purification and subsequent mass spectrometry techniques.
However these methods do not provide the layer of information needed in
order to determine what type of interaction is present in order to be
able to attribute signs to the network diagrams.
RNA interference screens
RNA interference
(RNAi) screens (repression of individual proteins between transcription
and translation) are one method that can be utilized in the process of
providing signs to the protein-protein interactions. Individual proteins
are repressed and the resulting phenotypes are analyzed. A correlating
phenotypic relationship (i.e. where the inhibition of either of two
proteins results in the same phenotype) indicates a positive, or
activating relationship. Phenotypes that do not correlate (i.e. where
the inhibition of either of two proteins results in two different
phenotypes) indicate a negative or inactivating relationship. If protein
A is dependent on protein B for activation then the inhibition of
either protein A or B will result in a cell losing the service that is
provided by protein A and the phenotypes will be the same for the
inhibition of either A or B. If, however, protein A is inactivated by
protein B then the phenotypes will differ depending on which protein is
inhibited (inhibit protein B and it can no longer inactivate protein A
leaving A active however inactivate A and there is nothing for B to
activate since A is inactive and the phenotype changes). Multiple RNAi
screens need to be performed in order to reliably appoint a sign to a
given protein-protein interaction. Vinayagam et al. who devised this
technique state that a minimum of nine RNAi screens are required with confidence increasing as one carries out more screens.
As therapeutic targets
Modulation of PPI is challenging and is receiving increasing attention by the scientific community. Several properties of PPI such as allosteric sites and hotspots, have been incorporated into drug-design strategies.
The relevance of PPI as putative therapeutic targets for the
development of new treatments is particularly evident in cancer, with
several ongoing clinical trials within this area.
The consensus among these promising targets is, nonetheless, denoted in
the already available drugs on the market to treat a multitude of
diseases. Examples are Tirobifan, inhibitor of the glycoprotein
IIb/IIIa, used as a cardiovascular drug, and Maraviroc, inhibitor of the
CCR5-gp120 interaction, used as anti-HIV drug.
Recently,
Amit Jaiswal and others were able to develop 30 peptides using
protein–protein interaction studies to inhibit telomerase recruitment
towards telomeres.