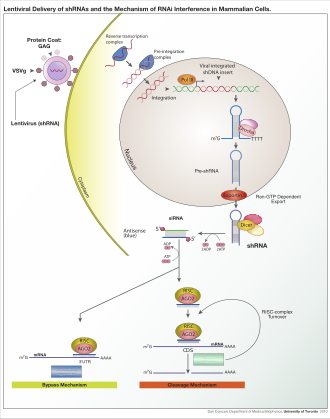
Lentiviral delivery of designed shRNA's and the mechanism of RNA interference in mammalian cells.
RNA interference (RNAi) is a biological process in which RNA molecules inhibit gene expression or translation, by neutralizing targeted mRNA molecules. Historically, RNA interference was known by other names, including co-suppression, post-transcriptional gene silencing (PTGS), and quelling. The detailed study of each of these seemingly different processes, elucidated that the identity of these phenomena were all actually RNAi. Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine for their work on RNA interference in the nematode worm Caenorhabditis elegans, which they published in 1998. Since the discovery of RNAi and its regulatory potentials, it has become evident that RNAi has immense potential in suppression of desired genes. RNAi is now known as precise, efficient, stable and better than antisense technology for gene suppression. However, antisense RNA produced intracellularly by an expression vector may be developed and find utility as novel therapeutic agents.
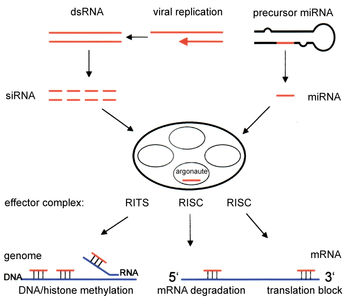
Two types of small ribonucleic acid (RNA) molecules – microRNA (miRNA) and small interfering RNA (siRNA) – are central to RNA interference. RNAs are the direct products of genes, and these small RNAs can direct enzyme complexes to degrade messenger RNA (mRNA) molecules and thus decrease their activity by preventing translation, via post-transcriptional gene silencing. Moreover, transcription can be inhibited via the pre-transcriptional silencing mechanism of RNA interference, through which an enzyme complex catalyzes DNA methylation at genomic positions complementary to complexed siRNA or miRNA. RNA interference has an important role in defending cells against parasitic nucleotide sequences – viruses and transposons. It also influences development.
The RNAi pathway is found in many eukaryotes, including animals, and is initiated by the enzyme Dicer, which cleaves long double-stranded RNA (dsRNA) molecules into short double-stranded fragments of ~21 nucleotide siRNAs. Each siRNA is unwound into two single-stranded RNAs (ssRNAs), the passenger strand and the guide strand. The passenger strand is degraded and the guide strand is incorporated into the RNA-induced silencing complex (RISC). The most well-studied outcome is post-transcriptional gene silencing, which occurs when the guide strand pairs with a complementary sequence in a messenger RNA molecule and induces cleavage by Argonaute 2 (Ago2), the catalytic component of the RISC. In some organisms, this process spreads systemically, despite the initially limited molar concentrations of siRNA.
RNAi is a valuable research tool, both in cell culture and in living organisms, because synthetic dsRNA introduced into cells can selectively and robustly induce suppression of specific genes of interest. RNAi may be used for large-scale screens that systematically shut down each gene in the cell, which can help to identify the components necessary for a particular cellular process or an event such as cell division. The pathway is also used as a practical tool in biotechnology, medicine and insecticides.[3]
Cellular mechanism
The dicer protein from Giardia intestinalis, which catalyzes the cleavage of dsRNA to siRNAs. The RNase domains are colored green, the PAZ domain yellow, the platform domain red, and the connector helix blue.[4]
RNAi is RNA-dependent gene silencing process that is controlled by the RNA-induced silencing complex (RISC) and is initiated by short double-stranded RNA molecules in a cell's cytoplasm, where they interact with the catalytic RISC component argonaute.[5] When the dsRNA is exogenous (coming from infection by a virus with an RNA genome or laboratory manipulations), the RNA is imported directly into the cytoplasm and cleaved to short fragments by Dicer. The initiating dsRNA can also be endogenous (originating in the cell), as in pre-microRNAs expressed from RNA-coding genes in the genome. The primary transcripts from such genes are first processed to form the characteristic stem-loop structure of pre-miRNA in the nucleus, then exported to the cytoplasm. Thus, the two dsRNA pathways, exogenous and endogenous, converge at the RISC.[6]
Exogenous dsRNA initiates RNAi by activating the ribonuclease protein Dicer,[7] which binds and cleaves double-stranded RNAs (dsRNAs) in plants, or short hairpin RNAs (shRNAs) in humans, to produce double-stranded fragments of 20–25 base pairs with a 2-nucleotide overhang at the 3' end.[8]
Bioinformatics studies on the genomes of multiple organisms suggest this length maximizes target-gene specificity and minimizes non-specific effects.[9] These short double-stranded fragments are called small interfering RNAs (siRNAs). These siRNAs are then separated into single strands and integrated into an active RISC, by RISC-Loading Complex (RLC). RLC includes Dicer-2 and R2D2, and is crucial to unite Ago2 and RISC.[10] TATA-binding protein-associated factor 11 (TAF11) assembles the RLC by facilitating Dcr-2-R2D2 tetramerization, which increases the binding affinity to siRNA by 10-fold. Association with TAF11 would convert the R2-D2-Initiator (RDI) complex into the RLC.[11] R2D2 carries tandem double-stranded RNA-binding domains to recognize the thermodynamically stable terminus of siRNA duplexes, whereas Dicer-2 the other less stable extremity. Loading is asymmetric: the MID domain of Ago2 recognizes the thermodynamically stable end of the siRNA. Therefore, the "passenger" (sense) strand whose 5′ end is discarded by MID is ejected, while the saved "guide" (antisense) strand cooperates with AGO to form the RISC.[10]
After integration into the RISC, siRNAs base-pair to their target mRNA and cleave it, thereby preventing it from being used as a translation template.[12] Differently from siRNA, a miRNA-loaded RISC complex scans cytoplasmic mRNAs for potential complementarity. Instead of destructive cleavage (by Ago2), miRNAs rather target the 3′ untranslated region (UTR) regions of mRNAs where they typically bind with imperfect complementarity, thus blocking the access of ribosomes for translation.[13]
Exogenous dsRNA is detected and bound by an effector protein, known as RDE-4 in C. elegans and R2D2 in Drosophila, that stimulates dicer activity.[14] The mechanism producing this length specificity is unknown and this protein only binds long dsRNAs.[14]
In C. elegans this initiation response is amplified through the synthesis of a population of 'secondary' siRNAs during which the dicer-produced initiating or 'primary' siRNAs are used as templates.[15] These 'secondary' siRNAs are structurally distinct from dicer-produced siRNAs and appear to be produced by an RNA-dependent RNA polymerase (RdRP).[16][17]
MicroRNA
MicroRNAs (miRNAs) are genomically encoded non-coding RNAs that help regulate gene expression, particularly during development.[18] The phenomenon of RNA interference, broadly defined, includes the endogenously induced gene silencing effects of miRNAs as well as silencing triggered by foreign dsRNA. Mature miRNAs are structurally similar to siRNAs produced from exogenous dsRNA, but before reaching maturity, miRNAs must first undergo extensive post-transcriptional modification. A miRNA is expressed from a much longer RNA-coding gene as a primary transcript known as a pri-miRNA which is processed, in the cell nucleus, to a 70-nucleotide stem-loop structure called a pre-miRNA by the microprocessor complex. This complex consists of an RNase III enzyme called Drosha and a dsRNA-binding protein DGCR8. The dsRNA portion of this pre-miRNA is bound and cleaved by Dicer to produce the mature miRNA molecule that can be integrated into the RISC complex; thus, miRNA and siRNA share the same downstream cellular machinery.[19] First, viral encoded miRNA was described in EBV.[20] Thereafter, an increasing number of microRNAs have been described in viruses. VIRmiRNA is a comprehensive catalogue covering viral microRNA, their targets and anti-viral miRNAs [21] (see also VIRmiRNA resource: http://crdd.osdd.net/servers/virmirna/).
siRNAs derived from long dsRNA precursors differ from miRNAs in that miRNAs, especially those in animals, typically have incomplete base pairing to a target and inhibit the translation of many different mRNAs with similar sequences. In contrast, siRNAs typically base-pair perfectly and induce mRNA cleavage only in a single, specific target.[22] In Drosophila and C. elegans, miRNA and siRNA are processed by distinct argonaute proteins and dicer enzymes.[23][24]
Three prime untranslated regions and microRNAs
Three prime untranslated regions (3'UTRs) of messenger RNAs (mRNAs) often contain regulatory sequences that post-transcriptionally cause RNA interference. Such 3'-UTRs often contain both binding sites for microRNAs (miRNAs) as well as for regulatory proteins. By binding to specific sites within the 3'-UTR, miRNAs can decrease gene expression of various mRNAs by either inhibiting translation or directly causing degradation of the transcript. The 3'-UTR also may have silencer regions that bind repressor proteins that inhibit the expression of a mRNA.The 3'-UTR often contains microRNA response elements (MREs). MREs are sequences to which miRNAs bind. These are prevalent motifs within 3'-UTRs. Among all regulatory motifs within the 3'-UTRs (e.g. including silencer regions), MREs make up about half of the motifs.
As of 2014, the miRBase web site,[25] an archive of miRNA sequences and annotations, listed 28,645 entries in 233 biologic species. Of these, 1,881 miRNAs were in annotated human miRNA loci. miRNAs were predicted to have an average of about four hundred target mRNAs (affecting expression of several hundred genes).[26] Friedman et al.[26] estimate that &45,000 miRNA target sites within human mRNA 3'UTRs are conserved above background levels, and & 60% of human protein-coding genes have been under selective pressure to maintain pairing to miRNAs.
Direct experiments show that a single miRNA can reduce the stability of hundreds of unique mRNAs.[27] Other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).[28][29]
The effects of miRNA dysregulation of gene expression seem to be important in cancer.[30] For instance, in gastrointestinal cancers, nine miRNAs have been identified as epigenetically altered and effective in down regulating DNA repair enzymes.[31]
The effects of miRNA dysregulation of gene expression also seem to be important in neuropsychiatric disorders, such as schizophrenia, bipolar disorder, major depression, Parkinson's disease, Alzheimer's disease and autism spectrum disorders.[32][33][34]
RISC activation and catalysis
Exogenous dsRNA is detected and bound by an effector protein, known as RDE-4 in C. elegans and R2D2 in Drosophila, that stimulates dicer activity.[14] This protein only binds long dsRNAs, but the mechanism producing this length specificity is unknown.[14] This RNA-binding protein then facilitates the transfer of cleaved siRNAs to the RISC complex.[35]In C. elegans this initiation response is amplified through the synthesis of a population of 'secondary' siRNAs during which the dicer-produced initiating or 'primary' siRNAs are used as templates.[15] These 'secondary' siRNAs are structurally distinct from dicer-produced siRNAs and appear to be produced by an RNA-dependent RNA polymerase (RdRP).[16][17]
small RNA
Biogenesis: primary miRNAs (pri-miRNAs) are transcribed in the nucleus
and fold back onto themselves as hairpins that are then trimmed in the
nucleus by a microprocessor complex to form a ~60-70nt hairpin pre-RNA. This pre-miRNA is transported through the nuclear pore complex (NPC) into the cytoplasm, where Dicer
further trims it to a ~20nt miRNA duplex (pre-siRNAs also enter the
pathway at this step). This duplex is then loaded into Ago to form the
“pre-RISC(RNA induced silencing complex)” and the passenger strand is
released to form active RISC.
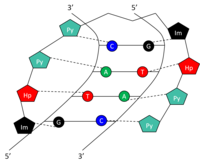
Left: A full-length argonaute protein from the archaea species Pyrococcus furiosus. Right: The PIWI domain of an argonaute protein in complex with double-stranded RNA.
The active components of an RNA-induced silencing complex (RISC) are endonucleases called argonaute proteins, which cleave the target mRNA strand complementary to their bound siRNA.[5] As the fragments produced by dicer are double-stranded, they could each in theory produce a functional siRNA. However, only one of the two strands, which is known as the guide strand, binds the argonaute protein and directs gene silencing. The other anti-guide strand or passenger strand is degraded during RISC activation.[36] Although it was first believed that an ATP-dependent helicase separated these two strands,[37] the process proved to be ATP-independent and performed directly by the protein components of RISC.[38][39] However, an in vitro kinetic analysis of RNAi in the presence and absence of ATP showed that ATP may be required to unwind and remove the cleaved mRNA strand from the RISC complex after catalysis.[40] The guide strand tends to be the one whose 5' end is less stably paired to its complement,[41] but strand selection is unaffected by the direction in which dicer cleaves the dsRNA before RISC incorporation.[42] Instead, the R2D2 protein may serve as the differentiating factor by binding the more-stable 5' end of the passenger strand.[43]
The structural basis for binding of RNA to the argonaute protein was examined by X-ray crystallography of the binding domain of an RNA-bound argonaute protein. Here, the phosphorylated 5' end of the RNA strand enters a conserved basic surface pocket and makes contacts through a divalent cation (an atom with two positive charges) such as magnesium and by aromatic stacking (a process that allows more than one atom to share an electron by passing it back and forth) between the 5' nucleotide in the siRNA and a conserved tyrosine residue. This site is thought to form a nucleation site for the binding of the siRNA to its mRNA target.[44] Analysis of the inhibitory effect of mismatches in either the 5’ or 3’ end of the guide strand has demonstrated that the 5’ end of the guide strand is likely responsible for matching and binding the target mRNA, while the 3’ end is responsible for physically arranging target mRNA into a cleavage-favorable RISC region.[40]
It is not understood how the activated RISC complex locates complementary mRNAs within the cell. Although the cleavage process has been proposed to be linked to translation, translation of the mRNA target is not essential for RNAi-mediated degradation.[45] Indeed, RNAi may be more effective against mRNA targets that are not translated.[46] Argonaute proteins are localized to specific regions in the cytoplasm called P-bodies (also cytoplasmic bodies or GW bodies), which are regions with high rates of mRNA decay;[47] miRNA activity is also clustered in P-bodies.[48] Disruption of P-bodies decreases the efficiency of RNA interference, suggesting that they are a critical site in the RNAi process.[49]
Transcriptional silencing
The enzyme dicer trims double stranded RNA, to form small interfering RNA or microRNA. These processed RNAs are incorporated into the RNA-induced silencing complex (RISC), which targets messenger RNA to prevent translation.[50]
Components of the RNAi pathway are used in many eukaryotes in the maintenance of the organization and structure of their genomes. Modification of histones and associated induction of heterochromatin formation serves to downregulate genes pre-transcriptionally;[51] this process is referred to as RNA-induced transcriptional silencing (RITS), and is carried out by a complex of proteins called the RITS complex. In fission yeast this complex contains argonaute, a chromodomain protein Chp1, and a protein called Tas3 of unknown function.[52] As a consequence, the induction and spread of heterochromatic regions requires the argonaute and RdRP proteins.[53] Indeed, deletion of these genes in the fission yeast S. pombe disrupts histone methylation and centromere formation,[54] causing slow or stalled anaphase during cell division.[55] In some cases, similar processes associated with histone modification have been observed to transcriptionally upregulate genes.[56]
The mechanism by which the RITS complex induces heterochromatin formation and organization is not well understood. Most studies have focused on the mating-type region in fission yeast, which may not be representative of activities in other genomic regions/organisms. In maintenance of existing heterochromatin regions, RITS forms a complex with siRNAs complementary to the local genes and stably binds local methylated histones, acting co-transcriptionally to degrade any nascent pre-mRNA transcripts that are initiated by RNA polymerase. The formation of such a heterochromatin region, though not its maintenance, is dicer-dependent, presumably because dicer is required to generate the initial complement of siRNAs that target subsequent transcripts.[57] Heterochromatin maintenance has been suggested to function as a self-reinforcing feedback loop, as new siRNAs are formed from the occasional nascent transcripts by RdRP for incorporation into local RITS complexes.[58] The relevance of observations from fission yeast mating-type regions and centromeres to mammals is not clear, as heterochromatin maintenance in mammalian cells may be independent of the components of the RNAi pathway.[59]
Crosstalk with RNA editing
The type of RNA editing that is most prevalent in higher eukaryotes converts adenosine nucleotides into inosine in dsRNAs via the enzyme adenosine deaminase (ADAR).[60] It was originally proposed in 2000 that the RNAi and A→I RNA editing pathways might compete for a common dsRNA substrate.[61] Some pre-miRNAs do undergo A→I RNA editing[62][63] and this mechanism may regulate the processing and expression of mature miRNAs.[63] Furthermore, at least one mammalian ADAR can sequester siRNAs from RNAi pathway components.[64] Further support for this model comes from studies on ADAR-null C. elegans strains indicating that A→I RNA editing may counteract RNAi silencing of endogenous genes and transgenes.[65]
Illustration of the major differences between plant and animal gene silencing. Natively expressed microRNA or exogenous small interfering RNA is processed by dicer and integrated into the RISC complex, which mediates gene silencing.[66]
Variation among organisms
Organisms vary in their ability to take up foreign dsRNA and use it in the RNAi pathway. The effects of RNA interference can be both systemic and heritable in plants and C. elegans, although not in Drosophila or mammals. In plants, RNAi is thought to propagate by the transfer of siRNAs between cells through plasmodesmata (channels in the cell walls that enable communication and transport).[37] Heritability comes from methylation of promoters targeted by RNAi; the new methylation pattern is copied in each new generation of the cell.[67] A broad general distinction between plants and animals lies in the targeting of endogenously produced miRNAs; in plants, miRNAs are usually perfectly or nearly perfectly complementary to their target genes and induce direct mRNA cleavage by RISC, while animals' miRNAs tend to be more divergent in sequence and induce translational repression.[66] This translational effect may be produced by inhibiting the interactions of translation initiation factors with the messenger RNA's polyadenine tail.[68]Some eukaryotic protozoa such as Leishmania major and Trypanosoma cruzi lack the RNAi pathway entirely.[69][70] Most or all of the components are also missing in some fungi, most notably the model organism Saccharomyces cerevisiae.[71] The presence of RNAi in other budding yeast species such as Saccharomyces castellii and Candida albicans, further demonstrates that inducing two RNAi-related proteins from S. castellii facilitates RNAi in S. cerevisiae.[72] That certain ascomycetes and basidiomycetes are missing RNA interference pathways indicates that proteins required for RNA silencing have been lost independently from many fungal lineages, possibly due to the evolution of a novel pathway with similar function, or to the lack of selective advantage in certain niches.[73]
Related prokaryotic systems
Gene expression in prokaryotes is influenced by an RNA-based system similar in some respects to RNAi. Here, RNA-encoding genes control mRNA abundance or translation by producing a complementary RNA that anneals to an mRNA. However these regulatory RNAs are not generally considered to be analogous to miRNAs because the dicer enzyme is not involved.[74] It has been suggested that CRISPR interference systems in prokaryotes are analogous to eukaryotic RNA interference systems, although none of the protein components are orthologous.[75]Biological functions
Immunity
RNA interference is a vital part of the immune response to viruses and other foreign genetic material, especially in plants where it may also prevent the self-propagation of transposons.[76] Plants such as Arabidopsis thaliana express multiple dicer homologs that are specialized to react differently when the plant is exposed to different viruses.[77] Even before the RNAi pathway was fully understood, it was known that induced gene silencing in plants could spread throughout the plant in a systemic effect and could be transferred from stock to scion plants via grafting.[78] This phenomenon has since been recognized as a feature of the plant adaptive immune system and allows the entire plant to respond to a virus after an initial localized encounter.[79] In response, many plant viruses have evolved elaborate mechanisms to suppress the RNAi response.[80] These include viral proteins that bind short double-stranded RNA fragments with single-stranded overhang ends, such as those produced by dicer.[81] Some plant genomes also express endogenous siRNAs in response to infection by specific types of bacteria.[82] These effects may be part of a generalized response to pathogens that downregulates any metabolic process in the host that aids the infection process.[83]Although animals generally express fewer variants of the dicer enzyme than plants, RNAi in some animals produces an antiviral response. In both juvenile and adult Drosophila, RNA interference is important in antiviral innate immunity and is active against pathogens such as Drosophila X virus.[84][85] A similar role in immunity may operate in C. elegans, as argonaute proteins are upregulated in response to viruses and worms that overexpress components of the RNAi pathway are resistant to viral infection.[86][87]
The role of RNA interference in mammalian innate immunity is poorly understood, and relatively little data is available. However, the existence of viruses that encode genes able to suppress the RNAi response in mammalian cells may be evidence in favour of an RNAi-dependent mammalian immune response,[88][89] although this hypothesis has been challenged as poorly substantiated.[90] Maillard et al.[91] and Li et al.[92] provide evidence for the existence of a functional antiviral RNAi pathway in mammalian cells. Other functions for RNAi in mammalian viruses also exist, such as miRNAs expressed by the herpes virus that may act as heterochromatin organization triggers to mediate viral latency.[93]
Downregulation of genes
Endogenously expressed miRNAs, including both intronic and intergenic miRNAs, are most important in translational repression[66] and in the regulation of development, especially on the timing of morphogenesis and the maintenance of undifferentiated or incompletely differentiated cell types such as stem cells.[94] The role of endogenously expressed miRNA in downregulating gene expression was first described in C. elegans in 1993.[95] In plants this function was discovered when the "JAW microRNA" of Arabidopsis was shown to be involved in the regulation of several genes that control plant shape.[96] In plants, the majority of genes regulated by miRNAs are transcription factors;[97] thus miRNA activity is particularly wide-ranging and regulates entire gene networks during development by modulating the expression of key regulatory genes, including transcription factors as well as F-box proteins.[98] In many organisms, including humans, miRNAs are linked to the formation of tumors and dysregulation of the cell cycle. Here, miRNAs can function as both oncogenes and tumor suppressors.[99]Upregulation of genes
RNA sequences (siRNA and miRNA) that are complementary to parts of a promoter can increase gene transcription, a phenomenon dubbed RNA activation. Part of the mechanism for how these RNA upregulate genes is known: dicer and argonaute are involved, possibly via histone demethylation.[100] miRNAs have been proposed to upregulate their target genes upon cell cycle arrest, via unknown mechanisms.[101]Evolution
Based on parsimony-based phylogenetic analysis, the most recent common ancestor of all eukaryotes most likely already possessed an early RNA interference pathway; the absence of the pathway in certain eukaryotes is thought to be a derived characteristic.[102] This ancestral RNAi system probably contained at least one dicer-like protein, one argonaute, one PIWI protein, and an RNA-dependent RNA polymerase that may also have played other cellular roles. A large-scale comparative genomics study likewise indicates that the eukaryotic crown group already possessed these components, which may then have had closer functional associations with generalized RNA degradation systems such as the exosome.[103] This study also suggests that the RNA-binding argonaute protein family, which is shared among eukaryotes, most archaea, and at least some bacteria (such as Aquifex aeolicus), is homologous to and originally evolved from components of the translation initiation system.The ancestral function of the RNAi system is generally agreed to have been immune defense against exogenous genetic elements such as transposons and viral genomes.[102][104] Related functions such as histone modification may have already been present in the ancestor of modern eukaryotes, although other functions such as regulation of development by miRNA are thought to have evolved later.[102]
RNA interference genes, as components of the antiviral innate immune system in many eukaryotes, are involved in an evolutionary arms race with viral genes. Some viruses have evolved mechanisms for suppressing the RNAi response in their host cells, particularly for plant viruses.[80] Studies of evolutionary rates in Drosophila have shown that genes in the RNAi pathway are subject to strong directional selection and are among the fastest-evolving genes in the Drosophila genome.[105]
Applications
Gene knockdown
The RNA interference pathway is often exploited in experimental biology to study the function of genes in cell culture and in vivo in model organisms.[5] Double-stranded RNA is synthesized with a sequence complementary to a gene of interest and introduced into a cell or organism, where it is recognized as exogenous genetic material and activates the RNAi pathway. Using this mechanism, researchers can cause a drastic decrease in the expression of a targeted gene. Studying the effects of this decrease can show the physiological role of the gene product. Since RNAi may not totally abolish expression of the gene, this technique is sometimes referred as a "knockdown", to distinguish it from "knockout" procedures in which expression of a gene is entirely eliminated.[106] In a recent study validation of RNAi silencing efficiency using gene array data showed 18.5% failure rate across 429 independent experiments.[107]Extensive efforts in computational biology have been directed toward the design of successful dsRNA reagents that maximize gene knockdown but minimize "off-target" effects. Off-target effects arise when an introduced RNA has a base sequence that can pair with and thus reduce the expression of multiple genes. Such problems occur more frequently when the dsRNA contains repetitive sequences. It has been estimated from studying the genomes of humans, C. elegans and S. pombe that about 10% of possible siRNAs have substantial off-target effects.[9] A multitude of software tools have been developed implementing algorithms for the design of general[108][109] mammal-specific,[110] and virus-specific[111] siRNAs that are automatically checked for possible cross-reactivity.
Depending on the organism and experimental system, the exogenous RNA may be a long strand designed to be cleaved by dicer, or short RNAs designed to serve as siRNA substrates. In most mammalian cells, shorter RNAs are used because long double-stranded RNA molecules induce the mammalian interferon response, a form of innate immunity that reacts nonspecifically to foreign genetic material.[112] Mouse oocytes and cells from early mouse embryos lack this reaction to exogenous dsRNA and are therefore a common model system for studying mammalian gene-knockdown effects.[113] Specialized laboratory techniques have also been developed to improve the utility of RNAi in mammalian systems by avoiding the direct introduction of siRNA, for example, by stable transfection with a plasmid encoding the appropriate sequence from which siRNAs can be transcribed,[114] or by more elaborate lentiviral vector systems allowing the inducible activation or deactivation of transcription, known as conditional RNAi.[115][116]
Functional genomics
A normal adult Drosophila fly, a common model organism used in RNAi experiments.
Most functional genomics applications of RNAi in animals have used C. elegans[117] and Drosophila,[118] as these are the common model organisms in which RNAi is most effective. C. elegans is particularly useful for RNAi research for two reasons: firstly, the effects of gene silencing are generally heritable, and secondly because delivery of the dsRNA is extremely simple. Through a mechanism whose details are poorly understood, bacteria such as E. coli that carry the desired dsRNA can be fed to the worms and will transfer their RNA payload to the worm via the intestinal tract. This "delivery by feeding" is just as effective at inducing gene silencing as more costly and time-consuming delivery methods, such as soaking the worms in dsRNA solution and injecting dsRNA into the gonads.[119] Although delivery is more difficult in most other organisms, efforts are also underway to undertake large-scale genomic screening applications in cell culture with mammalian cells.[120]
Approaches to the design of genome-wide RNAi libraries can require more sophistication than the design of a single siRNA for a defined set of experimental conditions. Artificial neural networks are frequently used to design siRNA libraries[121] and to predict their likely efficiency at gene knockdown.[122] Mass genomic screening is widely seen as a promising method for genome annotation and has triggered the development of high-throughput screening methods based on microarrays.[123][124] However, the utility of these screens and the ability of techniques developed on model organisms to generalize to even closely related species has been questioned, for example from C. elegans to related parasitic nematodes.[125][126]
Functional genomics using RNAi is a particularly attractive technique for genomic mapping and annotation in plants because many plants are polyploid, which presents substantial challenges for more traditional genetic engineering methods. For example, RNAi has been successfully used for functional genomics studies in bread wheat (which is hexaploid)[127] as well as more common plant model systems Arabidopsis and maize.[128]
Medicine
History of RNAi use in medicine
Timeline of the use of RNAi in medicine between 1996 and 2017
The first instance of RNA silencing in animals was documented in 1996, when Guo and Kemphues observed that, by introducing sense and antisense RNA to par-1 mRNA in Caenorhabditis elegans caused degradation of the par-1 message.[129] It was thought that this degradation was triggered by single stranded RNA (ssRNA), but two years later, in 1998, Fire and Mello discovered that this ability to silence the par-1 gene expression was actually triggered by double-stranded RNA (dsRNA).[129] They would eventually share the Nobel Prize in Physiology or Medicine for this discovery.[130] Just after Fire and Mello's ground-breaking discovery, Elbashir et al. discovered, by using synthetically made small interfering RNA (siRNA), it was possible to target the silencing of specific sequences in a gene, rather than silencing the entire gene.[131] Only a year later, McCaffrey and colleagues demonstrated that this sequence specific silencing had therapeutic applications by targeting a sequence from the Hepatitis C virus in transgenic mice.[132] Since then, multiple researchers have been attempting to expand the therapeutic applications of RNAi, specifically looking to target genes that cause various types of cancer.[133][134] Finally, in 2004, this new gene silencing technology entered a Phase I clinical trial in humans for wet age-related macular degeneration.[131] Six years later the first-in-human Phase I clinical trial was started, using a nanoparticle delivery system to target solid tumors.[135] Although most research is currently looking into the applications of RNAi in cancer treatment, the list of possible applications is extensive. RNAi could potentially be used to treat viruses,[136] bacterial diseases,[137] parasites,[138] maladaptive genetic mutations,[139] control drug consumption,[140] provide pain relief,[141] and even modulate sleep.[142]
Therapeutic applications
Viral infection
Antiviral treatment is one of the earliest proposed RNAi-based medical applications, and two different types have been developed. The first type is to target viral RNAs. Many studies have shown that targeting viral RNAs can suppress the replication of numerous viruses, including HIV,[143] HPV,[144] hepatitis A,[145] hepatitis B,[146] Influenza virus,[147] and Measles virus.[148] The other strategy is to block the initial viral entries by targeting the host cell genes. For example, suppression of chemokine receptors (CXCR4 and CCR5)on host cells can prevent HIV viral entry.[149]Cancer
While traditional chemotherapy can effectively kill cancer cells, lack of specificity for discriminating normal cells and cancer cells in these treatments usually cause severe side effects. Numerous studies have demonstrated that RNAi can provide a more specific approach to inhibit tumor growth by targeting cancer-related genes (i.e., oncogene).[150] It has also been proposed that RNAi can enhance the sensitivity of cancer cells to chemotherapeutic agents, providing a combinatorial therapeutic approach with chemotherapy.[151] Another potential RNAi-based treatment is to inhibit cell invasion and migration.[152]Neurological diseases
RNAi strategies also show potential for treating neurodegenerative diseases. Studies in cells and in mouse have shown that specifically targeting Amyloid beta-producing genes (e.g. BACE1 and APP) by RNAi can significantly reduced the amount of Aβ peptide which is correlated with the cause of Alzheimer's disease.[153][154][155] In addition, this silencing-based approaches also provide promising results in treatment of Parkinson's disease and Polyglutamine disease.[156][157][158]Difficulties in Therapeutic Application
To achieve the clinical potential of RNAi, siRNA must be efficiently transportated to the cells of target tissues. However, there are various barriers that must be fixed before it can be used clinically. For example, "Naked" siRNA is susceptible to several obstacles that reduce its therapeutic efficacy.[159] Additionally, once siRNA has entered the bloodstream, naked RNA can be degraded by serum nucleases and can stimulate the innate immune system.[160] Due to its size and highly polyanionic (containing negative charges at several sites) nature, unmodified siRNA molecules cannot readily enter the cells through the cell membrane. Therefore, artificial or nanoparticle encapsulated siRNA must be used. However, transporting siRNA across the cell membrane still has its own unique challenges. If siRNA is transferred across the cell membrane, unintended toxicities can occur if therapeutic doses are not optimized, and siRNAs can exhibit off-target effects (e.g. unintended downregulation of genes with partial sequence complementarity).[161] Even after entering the cells, repeated dosing is required since their effects are diluted at each cell division.Safety and Uses in Cancer treatment
Compared with chemotherapy or other anti-cancer drugs, there are a lot of advantages of siRNA drug.[162] SiRNA acts on the post-translational stage of gene expression, so it doesn’t modify or change DNA in a deleterious effect.[162] SiRNA can also be used to produced a specific response in a certain type of way, such as by downgrading suppression of gene expression.[162] In a single cancer cell, siRNA can cause dramatic suppression of gene expression with just several copies.[162] This happens by silencing cancer-promoting genes with RNAi, as well as targeting an mRNA sequence.[162]RNAi drugs treat cancer by silencing certain cancer promoting genes.[162] This is done by complementing the cancer genes with the RNAi, such as keeping the mRNA sequences in accordance with the RNAi drug.[162] Ideally, RNAi is should be injected and/or chemically modified so the RNAi can reach cancer cells more efficiently.[162] RNAi uptake and regulation is monitored by the kidneys.[162]
Stimulation of immune response
The human immune system is divided into two separate branches: the innate immune system and the adaptive immune system.[163] The innate immune system is the first defense against infection and responds to pathogens in a generic fashion.[163] On the other hand, the adaptive immune system, a system that was evolved later than the innate, is composed mainly of highly specialized B and T cells that are trained to react to specific portions of pathogenic molecules.[163]The challenge between old pathogens and new has helped create a system of guarded cells and particles that are called safe framework.[163] This framework has given humans an army systems that search out and destroy invader particles, such as pathogens, microscopic organisms, parasites, and infections.[163] The mammalian safe framework has developed to incorporate siRNA as a tool to indicate viral contamination, which has allowed siRNA is create an intense innate immune response.[163]
siRNA is controlled by the innate immune system, which can be divided into the acute inflammatory responses and antiviral responses.[163] The inflammatory response is created with signals from small signaling molecules, or cytokines.[163] These include interleukin-1 (IL-1), interleukin-6 (IL-6), interleukin-12 (IL-12) and tumor necrosis factor α (TNF-α).[163] The innate immune system generates inflammation and antiviral responses, which cause the release pattern recognition receptors (PRRs).[163] These receptors help in labeling which pathogens are viruses, fungi, or bacteria.[163] Moreover, the importance of siRNA and the innate immune system is to include more PRRs to help recognize different RNA structures.[163] This makes it more likely for the siRNA to cause an immunostimulant response in the event of the pathogen.[163]
Prospects as a Therapeutic Technique
Clinical Phase I and II studies of siRNA therapies conducted between 2015 and 2017 have demonstrated potent and durable gene knockdown in the liver, with some signs of clinical improvement and without unacceptable toxicity.[164] Two Phase III studies are in progress to treat familial neurodegenerative and cardiac syndromes caused by mutations in transthyretin (TTR).[165] Numerous publications have shown that in vivo delivery systems are very promising and are diverse in characteristics, allowing numerous applications. The nanoparticle delivery system shows the most promise yet this method presents additional challenges in the scale-up of the manufacturing process, such as the need for tightly controlled mixing processes to achieve consistent quality of the drug product.[166] The table below shows different drugs using RNA interference and what their phases and status is in clinical trials.[159]| Drug | Target | Delivery System | Disease | Phase | Status | Company | Identifier |
| ALN–VSP02 | KSP and VEGF | LNP | Solid tumours | I | Completed | Alnylam Pharmaceuticals | NCT01158079 |
| siRNA–EphA2–DOPC | EphA2 | LNP | Advanced cancers | I | Recruiting | MD Anderson Cancer Center | NCT01591356 |
| Atu027 | PKN3 | LNP | Solid tumours | I | Completed | Silence Therapeutics | NCT00938574 |
| TKM–080301 | PLK1 | LNP | Cancer | I | Recruiting | Tekmira Pharmaceutical | NCT01262235 |
| TKM–100201 | VP24, VP35, Zaire Ebola L-polymerase | LNP | Ebola-virus infection | I | Recruiting | Tekmira Pharmaceutical | NCT01518881 |
| ALN–RSV01 | RSV nucleocapsid | Naked siRNA | Respiratory syncytial virus infections | II | Completed | Alnylam Pharmaceuticals | NCT00658086 |
| PRO-040201 | ApoB | LNP | Hypercholesterolaemia | I | Terminated | Tekmira Pharmaceutical | NCT00927459 |
| ALN–PCS02 | PCSK9 | LNP | Hypercholesterolaemia | I | Completed | Alnylam Pharmaceuticals | NCT01437059 |
| ALN–TTR02 | TTR | LNP | Transthyretin-mediated amyloidosis | II | Recruiting | Alnylam Pharmaceuticals | NCT01617967 |
| CALAA-01 | RRM2 | Cyclodextrin NP | Solid tumours | I | Active | Calando Pharmaceuticals | NCT00689065 |
| TD101 | K6a (N171K mutation) | Naked siRNA | Pachyonychia congenita | I | Completed | Pachyonychia Congenita Project | NCT00716014 |
| AGN211745 | VEGFR1 | Naked siRNA | Age-related macular degeneration, choroidal neovascularization | II | Terminated | Allergan | NCT00395057 |
| QPI-1007 | CASP2 | Naked siRNA | Optic atrophy, non-arteritic anterior ischaemic optic neuropathy | I | Completed | Quark Pharmaceuticals | NCT01064505 |
| I5NP | p53 | Naked siRNA | Kidney injury, acute renal failure | I | Completed | Quark Pharmaceuticals | NCT00554359 |
| Delayed graft function, complications of kidney transplant | I, II | Recruiting | Quark Pharmaceuticals | NCT00802347 | |||
| PF-655 (PF-04523655) | RTP801 (Proprietary target) | Naked siRNA | Choroidal neovascularization, diabetic retinopathy, diabetic macular oedema | II | Active | Quark Pharmaceuticals | NCT01445899 |
| siG12D LODER | KRAS | LODER polymer | Pancreatic cancer | II | Recruiting | Silenseed | NCT01676259 |
| Bevasiranib | VEGF | Naked siRNA | Diabetic macular oedema, macular degeneration | II | Completed | Opko Health | NCT00306904 |
| SYL1001 | TRPV1 | Naked siRNA | Ocular pain, dry-eye syndrome | I, II | Recruiting | Sylentis | NCT01776658 |
| SYL040012 | ADRB2 | Naked siRNA | Ocular hypertension, open-angle glaucoma | II | Recruiting | Sylentis | NCT01739244 |
| CEQ508 | CTNNB1 | Escherichia coli-carrying shRNA | Familial adenomatous polyposis | I, II | Recruiting | Marina Biotech | Unknown |
| RXi-109 | CTGF | Self-delivering RNAi compound | Cicatrix scar prevention | I | Recruiting | RXi Pharmaceuticals | NCT01780077 |
| ALN–TTRsc | TTR | siRNA–GalNAc conjugate | Transthyretin-mediated amyloidosis | I | Recruiting | Alnylam Pharmaceuticals | NCT01814839 |
| ARC-520 | Conserved regions of HBV | DPC | HBV | I | Recruiting | Arrowhead Research | NCT01872065 |
Biotechnology
RNA interference has been used for applications in biotechnology and is nearing commercialization in others. RNAi has developed many novel crops such as nicotine free tobacco, decaffeinated coffee, nutrient fortified and hypoallergenic crops. The genetically engineered Arctic apples received FDA approval in 2015.[167] The apples were produced by RNAi suppression of PPO (polyphenol oxidase) gene making apple varieties that will not undergo browning after being sliced. PPO-silenced apples are unable to convert chlorogenic acid into quinone product.[1]There are several opportunities for the applications of RNAi in crop science for its improvement such as stress tolerance and enhanced nutritional level. RNAi will prove its potential for inhibition of photorespiration to enhance the productivity of C3 plants. This knockdown technology may be useful in inducing early flowering, delayed ripening, delayed senescence, breaking dormancy, stress-free plants, overcoming self-sterility, etc.[1]
Foods
RNAi has been used to genetically engineer plants to produce lower levels of natural plant toxins. Such techniques take advantage of the stable and heritable RNAi phenotype in plant stocks. Cotton seeds are rich in dietary protein but naturally contain the toxic terpenoid product gossypol, making them unsuitable for human consumption. RNAi has been used to produce cotton stocks whose seeds contain reduced levels of delta-cadinene synthase, a key enzyme in gossypol production, without affecting the enzyme's production in other parts of the plant, where gossypol is itself important in preventing damage from plant pests.[168] Similar efforts have been directed toward the reduction of the cyanogenic natural product linamarin in cassava plants.[169]No plant products that use RNAi-based genetic engineering have yet exited the experimental stage. Development efforts have successfully reduced the levels of allergens in tomato plants[170] and fortification of plants such as tomatoes with dietary antioxidants.[171] Previous commercial products, including the Flavr Savr tomato and two cultivars of ringspot-resistant papaya, were originally developed using antisense technology but likely exploited the RNAi pathway.[172][173]
Other crops
Another effort decreased the precursors of likely carcinogens in tobacco plants.[174] Other plant traits that have been engineered in the laboratory include the production of non-narcotic natural products by the opium poppy[175] and resistance to common plant viruses.[176]Insecticide
RNAi is under development as an insecticide, employing multiple approaches, including genetic engineering and topical application.[3] Cells in the midgut of some insects take up the dsRNA molecules in the process referred to as environmental RNAi.[177] In some insects the effect is systemic as the signal spreads throughout the insect's body (referred to as systemic RNAi).[178]RNAi technology is shown to be safe for consumption by mammals, including humans.[179]
RNAi has varying effects in different species of Lepidoptera (butterflies and moths).[180] Possibly because their saliva and gut juice is better at breaking down RNA, the cotton bollworm, the beet armyworm and the Asiatic rice borer have so far not been proven susceptible to RNAi by feeding.[3]
To develop resistance to RNAi, the western corn rootworm would have to change the genetic sequence of its Snf7 gene at multiple sites. Combining multiple strategies, such as engineering the protein Cry, derived from a bacterium called Bacillus thuringiensis (Bt), and RNAi in one plant delay the onset of resistance.[3][181]
Transgenic plants
Transgenic crops have been made to express dsRNA, carefully chosen to silence crucial genes in target pests. These dsRNAs are designed to affect only insects that express specific gene sequences. As a proof of principle, in 2009 a study showed RNAs that could kill any one of four fruit fly species while not harming the other three.[3]In 2012 Syngenta bought Belgian RNAi firm Devgen for $522 million and Monsanto paid $29.2 million for the exclusive rights to intellectual property from Alnylam Pharmaceuticals. The International Potato Center in Lima, Peru is looking for genes to target in the sweet potato weevil, a beetle whose larvae ravage sweet potatoes globally. Other researchers are trying to silence genes in ants, caterpillars and pollen beetles. Monsanto will likely be first to market, with a transgenic corn seed that expresses dsRNA based on gene Snf7 from the western corn rootworm, a beetle whose larvae annually cause one billion dollars in damage in the United States alone. A 2012 paper showed that silencing Snf7 stunts larval growth, killing them within days. In 2013 the same team showed that the RNA affects very few other species.[3]
Topical
Alternatively dsRNA can be supplied without genetic engineering. One approach is to add them to irrigation water. The molecules are absorbed into the plants' vascular system and poison insects feeding on them. Another approach involves spraying dsRNA like a conventional pesticide. This would allow faster adaptation to resistance. Such approaches would require low cost sources of dsRNAs that do not currently exist.[3]Genome-scale screening
Genome-scale RNAi research relies on high-throughput screening (HTS) technology. RNAi HTS technology allows genome-wide loss-of-function screening and is broadly used in the identification of genes associated with specific phenotypes. This technology has been hailed as the second genomics wave, following the first genomics wave of gene expression microarray and single nucleotide polymorphism discovery platforms.[182] One major advantage of genome-scale RNAi screening is its ability to simultaneously interrogate thousands of genes. With the ability to generate a large amount of data per experiment, genome-scale RNAi screening has led to an explosion of data generation rates. Exploiting such large data sets is a fundamental challenge, requiring suitable statistics/bioinformatics methods. The basic process of cell-based RNAi screening includes the choice of an RNAi library, robust and stable cell types, transfection with RNAi agents, treatment/incubation, signal detection, analysis and identification of important genes or therapeutical targets.[183]History
Example petunia plants in which genes for pigmentation are silenced by RNAi. The left plant is wild-type; the right plants contain transgenes
that induce suppression of both transgene and endogenous gene
expression, giving rise to the unpigmented white areas of the flower.[184]
The process of RNAi was referred to as "co-suppression" and "quelling" when observed prior to the knowledge of an RNA-related mechanism. The discovery of RNAi was preceded first by observations of transcriptional inhibition by antisense RNA expressed in transgenic plants,[185] and more directly by reports of unexpected outcomes in experiments performed by plant scientists in the United States and the Netherlands in the early 1990s.[186] In an attempt to alter flower colors in petunias, researchers introduced additional copies of a gene encoding chalcone synthase, a key enzyme for flower pigmentation into petunia plants of normally pink or violet flower color. The overexpressed gene was expected to result in darker flowers, but instead caused some flowers to have less visible purple pigment, sometimes in variegated patterns, indicating that the activity of chalcone synthase had been substantially decreased or became suppressed in a context-specific manner. This would later be explained as the result of the transgene being inserted adjacent to promoters in the opposite direction in various positions throughout the genomes of some transformants, thus leading to expression of antisense transcripts and gene silencing when these promoters are active. Another early observation of RNAi was came from a study of the fungus Neurospora crassa,[187] although it was not immediately recognized as related. Further investigation of the phenomenon in plants indicated that the downregulation was due to post-transcriptional inhibition of gene expression via an increased rate of mRNA degradation.[188] This phenomenon was called co-suppression of gene expression, but the molecular mechanism remained unknown.[189]
Not long after, plant virologists working on improving plant resistance to viral diseases observed a similar unexpected phenomenon. While it was known that plants expressing virus-specific proteins showed enhanced tolerance or resistance to viral infection, it was not expected that plants carrying only short, non-coding regions of viral RNA sequences would show similar levels of protection. Researchers believed that viral RNA produced by transgenes could also inhibit viral replication.[190] The reverse experiment, in which short sequences of plant genes were introduced into viruses, showed that the targeted gene was suppressed in an infected plant.[191] This phenomenon was labeled "virus-induced gene silencing" (VIGS), and the set of such phenomena were collectively called post transcriptional gene silencing.[192]
After these initial observations in plants, laboratories searched for this phenomenon in other organisms.[193][194] Craig C. Mello and Andrew Fire's 1998 Nature paper reported a potent gene silencing effect after injecting double stranded RNA into C. elegans.[195] In investigating the regulation of muscle protein production, they observed that neither mRNA nor antisense RNA injections had an effect on protein production, but double-stranded RNA successfully silenced the targeted gene. As a result of this work, they coined the term RNAi. This discovery represented the first identification of the causative agent for the phenomenon. Fire and Mello were awarded the 2006 Nobel Prize in Physiology or Medicine.