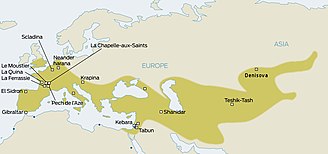
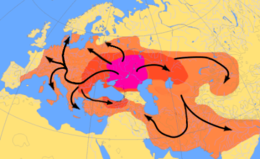
Ancient pathogen genomics is a scientific field related to the study of pathogen genomes recovered from ancient human, plant or animal remains. Ancient pathogens are microorganisms, now extinct, that in the past centuries caused several epidemics and deaths worldwide. Their genome, to which we referred as ancient DNA (aDNA), is isolated from the burial's remains (bones and teeth) of victims of the pandemics caused by these pathogens.
The analysis of the genomic features of ancient pathogen genomes allows researchers to understand the evolution of modern microbial strains that can hypothetically generate new pandemics or outbreaks. The analysis of aDNA is carried out by bioinformatic tools and molecular biology techniques to compare ancient pathogens with the modern descendants. The comparison also provides phylogenetic information of these strains.
Reconstructing ancient pathogen genomes through NGS technologies
Pathogen DNA detection in ancient remains can be achieved with laboratory or computational methods. In both cases, the procedure starts with the extraction of DNA from ancient specimens. The laboratory methods are based on the construction of NGS libraries and the subsequent capture-based screening. Computational tools are used to map the reads obtained by NGS against a single- or multi-genome reference (targeted approach); alternatively, metagenomic profiling or taxonomic assignment of shotgun NGS reads methods can be applied (broad approach).
Isolating ancient DNA
The limited preservation and thus low abundance, the highly fragmented and damaged state and the presence of modern DNA contamination and environmental DNA background makes the retrieval of ancient DNA (aDNA) a challenging procedure.
In order to efficiently recover aDNA, DNA is generally isolated from tissues that contain a high quantity of aDNA, like bone and teeth, which are abundant in archaeological record. The preservation of pathogens across different anatomical elements is very variable according to the type of pathogen and its tissue tropism, its route of entry into the body and the resulting disease. Pathogens that cause chronic infections in their hosts typically produce diagnostic bone changes as opposed to acute blood-borne infections. Therefore, for that infections that have caused the death of the host in the acute phase, the preferred sampling material is the inner chamber of the teeth since this is a tissue that is highly vascularized during life.
aDNA is characterised by damages that are accumulated over the course of time: the evaluation of DNA 'damage pattern' through computational tools is useful to authenticate ancient pathogen DNA since the same pattern is not found in modern contaminants.
The most represented chemical damage that affects the DNA post-mortem is the hydrolytic deamination of cytosines, converting them in uracils, which are then read as thymines. Due to this reaction, ancient DNA contains an unexpected proportion of cytosine to thymine transitions, in particular at the ends of the molecules. Other common DNA modifications, besides the deamination of cytosine into thymine (this occurs when cytosines were methylated), is the presence of abasic sites and single-strand breaks.
aDNA is extensively fragmented (most of the fragments are less than 100 base pairs long): this tendency can be used as a quantitative measure of authenticity, as modern contaminant molecules are expected to be longer. To exploit this characteristic feature of ancient DNA, improved silica-based extraction protocols with modified volume and composition of the DNA-binding buffer were introduced.
Construction of DNA libraries
In order to be sequenced with second generation sequencing methods, template molecules have to be modified through ligation of adaptors. Both the steps of library construction and the PCR amplification that follows are subject to errors. In particular, adaptor binding biases can occur and the relative efficacy of PCR enzymes in amplifying the construct can be variable.
There are three most common types of aDNA libraries. The double-stranded DNA library uses double-stranded DNA templates and firstly requires a step for the repair of the ends of aDNA fragments. Then, fragments are ligated to double-stranded adaptors and the resultant nicks are filled in. This method has some limitations, like the presence of a fraction of constructs that do not contain both the different adaptors and the possible formation of adaptor dimers.
To overcome this latter problem, a method for the construction of an A-tailed library was developed. In this method, aDNA is end-repaired and then an adenine residue is added to the 3' ends of the strands, which can facilitate the ligation of the template with adaptors that contain a tailor of thymine. Furthermore, the use of these T-tailed adaptors prevents the formation of adaptor dimers. The type of adaptor that is typically used is double-stranded and has a Y shape, which means that it has a region located at the T-tailed end where it is complementary and a region at the other end where it is non-complementary. The use of this type of adaptors allows to generate a template of aDNA flanked by different non-complementary adaptor sequences at each end that are useful for the unidirectional sequencing.
Another strategy is based on the use of single-stranded DNA libraries. In this method, DNA is first denatured to generate a single strand through heat and then ligated to a single-stranded biotinylated adaptor. The DNA strand is then used as a template by a DNA-polymerase which produces the complementary strand. Subsequently, a second adaptor is ligated at the 3' end of the complementary strand and the full construct is amplified through PCR and then sequenced. The purification step is performed using streptavidin-coated paramagnetic beads which allow minimising the DNA loss during this phase of the procedure.
Enriching libraries for aDNA
Different methods (called enrichment methods) have been developed to improve accessibility to endogenous DNA in ancient remains. These approaches can mainly be divided into three types: those used during library construction, by preferentially incorporating aDNA fragments characterised by the high level of damage, those applied after library construction, by separating exogenous and endogenous fractions through annealing to pre-defined sets of probes (in solution or on microarrays), or those based on targeted digestion of environmental microbial DNA using restriction enzymes and primer extension capture (PEC).
Selective uracil enrichment
During the construction of the library, the ssDNA fragments are bound through a biotinylated adaptor to streptavidin-coated beads. In the polymerase extension step, the DNA strand complementary to the original template is generated. In this kind of enrichment, the constructs undergo phosphorylation at the 5' end, to enable the ligation of a non-phosphorylated adaptor (ligation between the 3' end of the adaptor and the 5' end of the newly synthesized strand). DNA is then treated with uracil DNA glycosylase (UDG) and endonuclease VIII (USER mix): UDG generates abasic sites at cytosine that were deaminated into uracils post-mortem, endo VIII cuts at the resulting abasic site. This cleavage generates new 3' termini, which are then dephosphorylated, resulting in 3'OH ends that can be used as starting points for a new step of extension. This results in the elongation of the damaged strand, from the damaged region towards the bounded bead: while the new DNA molecule is synthesised, the original fragment is displaced. As a result, the dsDNA molecules newly formed no longer contain the adaptor bound to the beads, leaving in the supernatant a dsDNA library of the strands that originally harboured deaminated cytosines, available for further amplification and sequencing. The undamaged DNA template fraction remains attached to the paramagnetic beads.
Extension-free target enrichment in solution
This approach is based on in solution target-probe hybridization to screen for only a single microorganism, after the construction of the library. It is a species-specific assay that requires heat denaturation of DNA libraries and the construction of a probe DNA library using long-range PCR if fresh DNA material from closely related species is available, or through custom design and synthesis of oligonucleotides. This method is useful when the microorganism to target is known, for example, when the hypothesis exists for the causative agent of an epidemic or in presence of skeletal lesions in the studied individuals.
Solid-phase target enrichment
Another enriching strategy applied after constructing the library is the direct application of microarrays. They are applied for a wide laboratory-based pathogen screening that searches simultaneously for various pathogenic microorganisms. This kind of approach is favourable for those pathogens that leave no physical skeletal evidence and whose presence cannot be easily hypothesized a priori. The probes are designed to represent conserved or unique regions from a range of pathogenic viruses, parasites or bacteria.
Since microarrays contain sequences derived from modern strains of ancient pathogens, the limits of this method are the poor detection of the most divergent genomic regions and the omission of regions with important genomic rearrangements or unknown additional plasmids.
Whole-genome enrichment
The whole-genome in-solution capture (WISC) allows the characterization of the entire genome sequence of ancient individuals. This technique is based on the use of a genome-wide biotinylated RNA probe library generated through in vitro transcription of fresh modern DNA extracts from species closely related to the target aDNA sample. The heat-denatured aDNA library is then annealed to the RNA probes. To improve stringency and reduce enrichment for highly repetitive regions, low-complexity DNA and adaptor-blocking RNA oligonucleotides are added. The library fraction of interest in then recovered through elution from streptavidin-coated paramagnetic beads (to which the RNA probes are bound).
Computational analysis
The analysis of sequence data obtained by NGS relies on the same computational approaches used for modern DNA, with some peculiarities. A widely used tool to align reads from aDNA against reference genomes is the PALEOMIX package, which can quantify DNA damage levels through mapDamage2 and perform phylogenomic and metagenomic analyses. It is important to consider that the alignment will always exhibit substantial fractions of nucleotides mismatched that do not result from sequencing errors or polymorphisms but from the presence of damaged bases. For this reason, the acceptance threshold for read-to-reference edit distance should be chosen according to the phylogenetic distance to the reference genome. Probabilistic aligners that take into account the damage pattern of aDNA have been developed to improve alignments.
MALT
Studies of the ancient DNA of pathogens is restricted to skeletal collections that change their appearance as a result of infections. A pathogen linked to a known epidemiological context is identified through screening without prior knowledge of its presence. Methods include broad-spectrum molecular approaches focused on pathogen detection via fluorescence hybridization-based microarray technology, identification via DNA enrichment of certain microbial regions or computational screening of non-enriched sequence data against human microbiome data sets. These approaches offer improvements but remain biased in the bacterial taxa used for species-level assignments.
MEGAN alignment tool (MALT) is a new program for the fast alignment and taxonomic assignment method to the identification of ancient DNA. MALT is similar to BLAST as it computes local alignments between highly conserved sequences and references. MALT can also calculate semi-global alignments where reads are aligned end-to-end. All references, complete bacterial genomes, are contained in a database called National Center for Biotechnology Information (NCBI) RefSeq. MALT consists of two programs: malt-build and malt-run. Malt-build is used to construct an index for the given database of reference sequences. Instead, malt-run is used to align a set of query sequences against the reference database. The program then computes the bit-score and the expected value (E-value) of the alignment and decides whether to keep or discard the alignment depending on user-specified thresholds for the bit-score, the E-value or the per cent identity. The bit-score is the requires size of a sequence database in which the current match could be found just by chance. The higher the bit-score, the better the sequence similarity. E-value is the number of expected hits of similar quality (score) that could be found just by chance. The smaller is the E-value, the better is the match.
MALT allows the screening of non-enriched sequence data in the search for unknown candidate bacterial pathogens that are involved in past disease outbreaks and for the exclusion of the environmental bacterial background. MALT is very important because it offers the advantage of genome-level screening without selection of a particular target organism, avoiding errors that are common to other screening approaches. To authenticate the candidate taxonomic assignments complete alignments are needed, but the target DNA is often present in a low amount so a small number of a marked region may not be sufficient for identification. This approach can detect only bacterial DNA and viral DNA, so it is not possible to identify other infectious agents that may be present in a population. This method is useful for studies dealing with the identification of pathogens responsible for ancient and modern disease, especially in cases for which candidate organisms are not known a priori.
Applications
Ancient pathogen genomics as a tool against future epidemics
One interesting application of the different sequencing techniques available nowadays is the investigation of historical disease outbreaks to provide an answer to important and long-standing questions in epidemiology, pathogen evolution and also human history.
So, much effort is spent to find more and more information about the aetiology of infectious diseases of historical importance, such as plague and the cocoliztli epidemic, to describe the geographic spread of viruses and to try defining the pathogenic mechanism of these infectious agents that are actually active elements of the evolutionary process. Today Y.pestis and S. enterica seem to be so far from us and no more dangerous at all, but scientists are still interested in the long-term tracing of genetic adaptation of these bacteria and accurate quantification of rates of their evolutionary change. This is because they can extract from this knowledge of the past the right ideas to develop a strategy against future epidemics.
Being perfectly aware of the fact that bacteria and viruses are one of most variable elements in nature, prone to unlimited mutational events, and taking for granted that it is impossible to manage all the external factors that can influence the development of a pathogenic virus, nobody is talking about defeating a new possible outbreak of plague or any other infective agent of the past: here the aim is to define a strategy, a "guideline", to be more prepared when a new dangerous pathogen will come. The contribution of the environment in infections is to be defined and factors such as human migration, climate change, overcrowding in cities or animal domestication are some of the major causes that contribute to the emergence and spread of disease. Of course, these factors are unpredictable and this is a reason why researchers are trying to bring relevant information from the past, that can be useful, today and tomorrow. While they continue to develop strategies to defeat emerging threats using diagnostic, molecular and advanced tools, they are still looking back at how ancient pathogens have evolved and adapted through historical events. The more it's known about the genomic basis of virulence in historical diseases, the more it can be understood about the emergence and re-emergence of infectious diseases today and in the future.
Ancient infections and human evolution
The analyses of phylogenic relationships between the human host and viral pathogens suggest that many diseases have been coevolving with humans for millennia, since the very start of human history in Africa.
In particular, the long-term interaction with pathogens is considered a selection that can be very strong since not all the individuals could survive in touch with all infectious agents that they had met over the years: the natural selection by pathogens is implicated in the evolution of species. This interaction has been already used to track human population movements and to reconstruct human migration flows within and out of Africa.
A pretty new application and interpretation of this feature is using aDNA to better understand human evolution. Many tropical infections probably played a significant role in the human evolutionary process. The correlation between humans and viruses can be understood if it is seen as a "fight" that continues for millennia and that is not still won by anyone: when viruses have changed their features in order to be infective for the other "fighters", humans had to find a strategy to increase their fitness and survived among changes.
In this continuous challenge through the years, next to infective diseases and other illnesses afflicting modern human society, cancer recently represents one of the most enigmatical ailments. Scientists are investigating if neoplastic diseases are restricted to postindustrial human society or if their origins can be found further back in time, maybe into prehistory. The difficulty is that cancer, lethal and fast, leaves very few indications in skeletons in those cases that succumb to death shortly, and even no signs of existence at all, in the case of extraskeletal tumours. Anyway, the knowledge about the aetiology of cancer is incomplete and microorganisms are taking their part with the role of their infection: migration movements in the past could have brought with them viruses, so possible reservoir of tropical disease as well as predisposition to cancer. For this reason, molecular analytical techniques are applied to archaeological remains to study hominin evolution, but also to improve the research in understanding the epidemiology and aetiology of tumours. Information derived from the aDNA can be used to anchor pathogen mutations and reconstruct back from the presence of microorganisms the evolutionary process, it can be useful to develop new vaccines or to discover possible future pathogenic threats.
Past pandemics are much more than just ancient history
What happened in the past is not all history, there is something hidden that can still drive human genetic diversity and natural selection, something that went in contact with humankind hundreds of years ago but that can still have an impact on global human health. Since epidemics are one of the most frequent phenomena that have affected and potentially devastated human populations, it is important to detect, prevent and control potential infective agents. After all, archaeologists, geneticists, and medical scientists are concerned in exploring the influences of pathogens that can contribute, threatening or improving, human health and longevity.
Evolution and phylogenesis of Yersinia pestis
Yersinia pestis is a gram-negative bacterium and belongs to the family of Enterobatteriaceae. Its closest relatives are Yersinia pseudotuberculosis and Yersinia enterocolitica, which are environmental species.
They all possess the plasmid pCD1, which encodes for a type III secretory system. Among chromosomal protein-coding genes, their nucleotide genomic identity rates 97%. They are different in terms of their virulence potential and transmission mechanisms.
Y. pestis is not a human-adapted bacteria. Its main reservoirs are rodents (like marmots, mice, great gerbils, voles and prairie dogs) and it is transmitted to humans by the flea. One of the most studied vectors of this pathogen is Xenopsylla cheopis.
After the bite of an infected flea, the bacteria enter into the host organism and travel to the closest lymph node, where bacteria replicate causing the large swellings called buboes. Bacteria can also disseminate into the bloodstream (causing septicaemia) and to the lung (causing pneumonia). The pulmonary disease has a direct human-to-human transmission.
It has been determined that Y. pestis became so dangerous because of the acquisition of ymt (yersina murine toxin). This gene is present on the pMT1 plasmid and allowed the survival of the bacterium in the flea vector and facilitated colonization of the midgut in arthropod, giving rise to the past millennium large-scale pandemics.
Early evolution and divergence from Yersinia pseudotuberculosis
Y. pestis is distinguishable from the other two species because of its pathogenicity and transmission mechanism. These differences are given by two plasmids: pPCP1, that confer to the bacterium its invasive properties in humans and pMT1, which is involved in flea colonisation (along with some loss of function on bacterial chromosomal genes).
Samples dated on the Late Neolithic and Bronze Age allowed identifying a first genetic divergence between Y. pseudotuberculosis and Y. pestis ancestors. The characteristics that confer to Y. pestis its virulence were absent in these strains: they lack of ymt, a gene necessary to the colonization of the vector; also, they presented an active form of genes required for biofilm formation (inactive in the pathogen Y. pestis) and an active flagellin gene, that is an inducer of immune response (is a pseudogene in Y. pestis).
The comparison of a draft of the genome and the two plasmids (pCD1 and pMT1) with samples of Black Death victims (1348-1349) in the East Smithfield burial ground underlined a very high genetic conservation of the sequence: only 97 single-nucleotide differences over 660 years.
Y. pestis microevolution
The London 6330 individual strain owns mutations absent in other isolates of the same period (1348-1350): the reason may be either the presence of multiple strains circulating in Europe at the same time or the microevolution of one single strain during the pandemic.
Three major outbreaks of plague
There are three pandemic outbreaks of Y. Pestis:
- The first is known as the Plague of Justinian, it first occurred in Egypt in 541-543 and then spread to Constantinople and neighbouring regions. It had outbreaks in Europe until 750 CE. Phylogenetic analysis showed that both genomes belong to a lineage that is extinct today and is closely related to stains from modern-day China, which suggest the possibility of an East Asian origin of the first pandemic.
- The second pandemic is known as the Black Death or as Great Pestilence. It occurred in 1346-1352 in Europe and had a lot of resurgences of plague, it continued until the 18th century. It could be possible that in this pandemic there were two different strains of Y. pestis that entered the continent through different pulses.
- The third pandemic started in China in 1860. It has a fast spread to other countries, due to the use of railways and steamships.
The strains associated with the Justinian Plague appear on a novel branch, which is phylogenetically distinct from the second and the third plague pandemics. The first strain of Y. pestis found during the second outbreak survives and give rise to modern branch 1 strains associated with the third pandemic outbreak.
The first plague bacteria and the second and third plague strain have a common ancestor.
Linkage between 2nd and 3rd pandemics
In a recent study, genomes of Y. pestis from three samples resumed in Barcelona (deceased between 1300-1420), Ellwangen (between 1486-1627) and Bolgar city (between 1298-1388). The date of death of the individuals analysed was determined thanks to radiocarbon dates; the last one was confirmed by the presence of a coin produced only after the year 1362. Of 223 samples from 178 individuals, only one for each site had a suitable amount of DNA and was finally selected for the whole genome sequencing of the bacillus (through a genome-capture assay, using as a draft Y. pseudotuberculosis genome and pMT1 and pCD1 from Y. pestis).
The alignment with a Y. pestis phylogeny tree created with previously know ancient genomes revealed an increase genetic diversity outside of China in comparison to what was previously thought; all the three new genomes mapped in Branch 1 and possess two SNPs associated to the Black Death (all the genomes of Y. pestis dated to the Black Death map in Branch 1). The Barcelona strain has no differences with the London strain; the two individuals from which the genome was obtained died of plague with a distance of some months (spring and autumn 1348) underlining the presence in Europe of a single wave of plague with low genetic diversity. The Ellwangen strain maps in a sub-branch of Branch 1 and is ancestral to a previously sequenced strain (L'Observance). it descends from the one circulating in London and Barcelona during the Black Death but also have additional mutations. Is therefore considered a lineage diverged from Branch 1 before the 16th century (Ellwangen outbreak) and with no known modern descendants.
In comparison with isolates from the Black Death, the Bolgar city strain presents:
- p3 and p4, shared by the "London individual 6330";
- p6, shared with all modern Branch 1 strains;
- p7, unique of this strain;
The Bolgar City strain possesses SNPs associated to the Black Death and can be an evidence of a movement of plague towards east; These findings give credit to one of the models that try to explain the linkage between 2nd and 3rd pandemic: in this scenario, there was a single exit of plague to Europe (causing the Black Death) that after a radiation event, travelled eastward to establish in former soviet union and Asia, from which it spread in the 18th century to give raise to the 3rd pandemic.
Another hypothesis is that the 3rd pandemic's lineage may have been generated by a pre-existing genetic variability in Y. pestis strains in China: this hypothesis is actually supported by the correlation between following waves of the pandemic in Europe and climatic fluctuations that would have allowed its spread in the continent. This model can't explain the genetic diversity of the Black Death (four different lineages at least, that would have required the introduction from Asia of four different strains).
Again, there are two models that try to explain the multiple plague outbreaks in Europe following the black death:
- Repeated introduction of plague from Asia. This scenario is compatible with the 2nd theory discussed before that sees a genetic variability of Y. pestis in China;
- Presence in Europe of a reservoir (now extinct) that caused continue outbreaks until the 18th century;
Both models can be valid and nowadays we're not able to demonstrate one over the other. However, the Ellwangen strain genome sequenced in this study may be considered a proof of the second hypothesis due to the geographical position of the city that tends to exclude the possibility of an introduction of plague from eastward.
Modern Y. pestis strains
Sequencing of Y. pestis genomes allowed to discover a variation event preceding Black Death that gave rise to many strains that circulate today.
Salmonella enterica genomes analysis
During the 16th an epidemic occurred in Mexico and it caused high mortality in Indigenous populations of the Americas. This high mortality has been the consequence of the influence of the demographic collapse of many indigenous populations. This epidemic has been called "cocoliztli" by the native Aztec because of the symptoms, in particular, high fever and bleeding.
This pestilence is considered one of the worst epidemics in the history of Mexico and the cause of this outbreaks is remained a mystery for over 500 years.
A group of scientists from Harvard and Max-Planck Institute published a study in the journal of Nature ecology and Evolution, and they suggest Salmonella enterica as a good candidate for the strong epidemic in Mexico during the 16th century. Many studies suggest that this bacterium has been introduced in the Indigenous populations by Europeans.
The group of scientists analyzed the aDNA extracted from the teeth of 24 skeletons buried in a cemetery in the city of Teposcolula-Yucundaa and they found in 10 of the 24 skeletons aDNA traces of Salmonella enterica. Also, to demonstrate that the bacterium was introduced in Mexico by the Europeans, they analyzed five individuals that were buried before the influx of Europeans. The results revealed that there was no evidence of Salmonella enterica in the pre-contact era.
Analysis of Salmonella enterica genomes
The scientists performed the extraction of the aDNA from the teeth of 24 indigenous individuals' remains from the contact era epidemic cemetery and of 5 individuals buried in the pre-contact era cemetery. The extraction was performed according to the protocol for aDNA extraction. The group of researchers examined, in parallel, also a soil sample of the cemeteries to get an overview of the environment microorganisms that could have penetrated the samples.
After the extraction, the genomes were sequenced using Illumina genome analyzer. Then, using a bioinformatic tool, called MALT, the researchers performed an analysis of metagenomic sequences data. This program lets the researchers align the extracted sequences with a reference without specifying a precise target. The researchers performed MALT run two times: one using the complete bacterial genomes that were available through NCBI (National Center for Biotechnology Information) RefSeq as a reference, and the second run was carried out using the full NCBI Nucleotide database to screen for viral DNA.
The result of the screening process was positive for the presence of Salmonella enterica DNA in 10 sequences up to 24 collected from the samples and three tooth sample had a high amount of reads assigned to S. enterica. In particular, the major S. enterica strain present in the samples is the S. Paratyphi C. This strain causes enteric fever in human individuals. In the pre-contact era samples, they did not find any evidence of S. enterica, supporting the hypothesis that S. enterica was not a local bacteria.
A further analysis was carried out to identify the classical pattern of damage of aDNA in the three positive tooth samples and this was conducted by mapping the data sets to the S. Paratyphy C genome reference. The results were positive and supported the thesis of S.enterica as the cause of cocolitzli.
To go in-depth with the analyses and confirm the thesis, the researchers conducted further experiments and computational analysis. They performed a whole-genome target array and in-solution hybridization capture using probes that include the modern S. enterica genome differences and using S. Paratyphi C as reference. The hybridization was successful for the ten positive samples, while the other samples resulted negative for the ancient DNA.