The last universal common ancestor (LUCA) is the hypothesized latest common ancestral cell population from which all subsequent life forms descend, including Bacteria, Archaea, and Eukarya. The cell had a lipid bilayer; it possessed the genetic code and ribosomes which translated from DNA or RNA to proteins. Although the timing of the LUCA cannot be definitively constrained, most studies suggest that the LUCA existed by 3.5 billion years ago, and possibly as early as 4.3 billion years ago or earlier. The nature of this point or stage of divergence remains a topic of research.
All earlier forms of life preceding this divergence and all extant organisms are generally thought to share common ancestry. On the basis of a formal statistical test, this theory of a universal common ancestry (UCA) is supported in preference to competing multiple-ancestry hypotheses. The first universal common ancestor (FUCA) is a hypothetical non-cellular ancestor to LUCA and other now-extinct sister lineages.
Whether the genesis of viruses falls before or after the LUCA–as well as the diversity of extant viruses and their hosts–remains a subject of investigation.
While no fossil evidence of the LUCA exists, the detailed biochemical similarity of all current life (divided into the three domains) makes its existence widely accepted by biochemists. Its characteristics can be inferred from shared features of modern genomes. These genes describe a complex life form with many co-adapted features, including transcription and translation mechanisms to convert information from DNA to mRNA to proteins.
Historical background

A phylogenetic tree directly portrays the idea of evolution by descent from a single ancestor. An early tree of life was sketched by Jean-Baptiste Lamarck in his Philosophie zoologique in 1809. Charles Darwin more famously proposed the theory of universal common descent through an evolutionary process in his book On the Origin of Species in 1859: "Therefore I should infer from analogy that probably all the organic beings which have ever lived on this earth have descended from some one primordial form, into which life was first breathed." The last sentence of the book begins with a restatement of the hypothesis:
There is grandeur in this view of life, with its several powers, having been originally breathed into a few forms or into one ...
By 1871, in another letter to Hooker, Darwin speculated on the natural origin of life itself, writing that life might have begun in a "warm little pond, with all sorts of ammonia and phosphoric salts, light, heat, electricity, etc., present", an early expression of abiogenesis.
The term "last universal common ancestor" or "LUCA" was first used in the 1990s for such a primordial organism.
Inferring LUCA's features
Biochemical mechanisms
While the anatomy of LUCA cannot be reconstructed with certainty, its biochemical mechanisms can be deduced and described in some detail, based on properties shared by currently living organisms as well as genetic analysis.
LUCA certainly had genes and a genetic code. Its genetic material was most likely DNA, so that it lived after the RNA world. The DNA was kept double-stranded by an enzyme, DNA polymerase, which recognises the structure and directionality of DNA. The integrity of the DNA was maintained by a group of repair enzymes including DNA topoisomerase. If the genetic code was based on dual-stranded DNA, it was expressed by copying the information to single-stranded RNA. The RNA was produced by a DNA-dependent RNA polymerase using nucleotides similar to those of DNA. It had multiple DNA-binding proteins, such as histone-fold proteins. The genetic code was expressed into proteins. These were assembled from 20 free amino acids by translation of a messenger RNA via a mechanism of ribosomes, transfer RNAs, and a group of related proteins.
Although LUCA was likely not capable of sexual interaction, gene functions were present that promoted the transfer of DNA between individuals of the population to facilitate genetic recombination. Homologous gene products that promote genetic recombination are present in bacteria, archaea and eukaryota, such as the RecA protein in bacteria, the RadA protein in archaea, and the Rad51 and Dmc1 proteins in eukaryota.
LUCA's functionality, and evidence for the early evolution of membrane-dependent biological systems, together suggest that LUCA was a cell with membranes. It contained a water-based cytoplasm enclosed by a lipid bilayer membrane; it reproduced by cell division. It tended to exclude sodium and concentrate potassium by means of specific ion transporters (or ion pumps). The cell multiplied by duplicating all its contents followed by cellular division. The cell used chemiosmosis to produce energy. It also reduced CO2 and oxidized H2 (methanogenesis or acetogenesis) via acetyl-thioesters.
By phylogenetic bracketing, analysis of its offspring groups, LUCA appears to have been a small, single-celled organism. It likely had a ring-shaped coil of DNA floating freely within the cell. Morphologically, it would likely not have stood out within a mixed population of small modern-day bacteria. The originator of the three-domain system, Carl Woese, stated that in its genetic machinery, the LUCA would have been a "simpler, more rudimentary entity than the individual ancestors that spawned the three [domains] (and their descendants)".
Because bacteria and archaea differ in their structure of phospholipids and cell wall, ion pumping, most proteins involved in DNA replication, and glycolysis, it is inferred that LUCA had a permeable membrane without an ion pump. The emergence of Na+/H+ antiporters likely led to the later evolution of impermeable membranes in eukaryotes, archaea, and bacteria. This would accord with LUCA's having made use of the natural geochemical proton gradient in its environment across a leaky membrane to provide it with energy. Cell walls, too, would have evolved later. Although LUCA likely had DNA, it is unknown if it could replicate DNA: as Weiss et al write, it "might just have been a chemically stable repository for RNA-based replication". It is likely that LUCA's permeable membrane was composed of archaeal lipids (isoprenoids) and bacterial lipids (fatty acids). Isoprenoids would have helped to stabilize LUCA's membrane in the surrounding extreme habitat.
LUCA's genome was likely similar in size to that of modern prokaryotes, encoding around 2,600 proteins, based on statistical inference using the probabilistic gene- and species-tree reconciliation algorithm ALE. It may have been an acetogen, respiring anaerobically, and may have had an early CAS-based anti-viral immune system. The inferred metabolic features are consistent with the early Earth hydrothermal systems with high concentrations of CO2 and H2.
An anaerobic thermophile
An alternative to the search for "universal" traits is to use genome analysis to identify phylogenetically ancient genes. This gives a picture of a LUCA that could live in a geochemically harsh environment and is like modern prokaryotes. Analysis of biochemical pathways implies the same sort of chemistry as does phylogenetic analysis.

In 2016, Madeline C. Weiss and colleagues genetically analyzed 6.1 million protein-coding genes and 286,514 protein clusters from sequenced prokaryotic genomes representing many phylogenetic trees, and identified 355 protein clusters that were probably common to the LUCA. The results of their analysis are highly specific, though debated. They depict LUCA as "anaerobic, CO2-fixing, H2-dependent with a Wood–Ljungdahl pathway (the reductive acetyl-coenzyme A pathway), N2-fixing and thermophilic. LUCA's biochemistry was replete with FeS clusters and radical reaction mechanisms." The cofactors indicate "dependence upon transition metals, flavins, S-adenosyl methionine, coenzyme A, ferredoxin, molybdopterin, corrins and selenium. Its genetic code required nucleoside modifications and S-adenosylmethionine-dependent methylations." They show that methanogens and clostridia were basal, near the root of the phylogenetic tree, in the 355 protein lineages examined, and that the LUCA may therefore have inhabited an anaerobic hydrothermal vent setting in a geochemically active environment rich in H2, CO2, and iron, where ocean water interacted with hot magma beneath the ocean floor. It is inferred that LUCA grew from H2 and CO2 via the reverse incomplete Krebs cycle. Other metabolic pathways inferred in LUCA are the pentose phosphate pathway, glycolysis, and gluconeogenesis. Even if phylogenetic evidence may point to a hydrothermal vent environment for a thermophilic LUCA, this does not constitute evidence that the origin of life took place at a hydrothermal vent since mass extinctions may have removed previously existing branches of life.

Weiss and colleagues write that "Experiments ... demonstrate that ... acetyl-CoA pathway [chemicals used in anaerobic respiration] formate, methanol, acetyl moieties, and even pyruvate arise spontaneously ... from CO2, native metals, and water", a combination present in hydrothermal vents.
An experiment shows that Zn2+, Cr3+, and Fe can promote 6 of the 11 reactions of an ancient anabolic pathway called the reverse Krebs cycle in acidic conditions which implies that LUCA might have inhabited either hydrothermal vents or acidic metal-rich hydrothermal fields.
Undersampled protein families
Some other researchers have challenged Weiss et al.'s 2016
conclusions. Sarah Berkemer and Shawn McGlynn argue that Weiss et al.
undersampled the families of proteins, so that the phylogenetic trees
were not complete and failed to describe the evolution of proteins
correctly. There are two risks in attempting to attribute LUCA's
environment from near-universal gene distribution (as in Weiss et al.
2016). On the one hand, it risks misattributing convergence
or horizontal gene transfer events to vertical descent; on the other
hand, it risks misattributing potential LUCA gene families as horizontal
gene transfer events. A phylogenomic and geochemical analysis of a set
of proteins that probably traced to the LUCA show that it had K+-dependent GTPases and the ionic composition and concentration of its intracellular fluid was seemingly high K+/Na+ ratio, NH+
4, Fe2+, CO2+, Ni2+, Mg2+, Mn2+, Zn2+, pyrophosphate, and PO3−
4 which would imply a terrestrial hot spring habitat. It possibly had a phosphate-based metabolism. Further, these proteins were unrelated to autotrophy (the ability of an organism to create its own organic matter), suggesting that the LUCA had a Heterotrophic lifestyle (consuming organic matter) and that its growth was dependent on organic matter produced by the physical environment.
The presence of the energy-handling enzymes CODH/acetyl-coenzyme A synthase in LUCA could be compatible with being an autotroph and with life as a mixotroph or heterotroph. Weiss et al. in 2018 replied that no enzyme defines a trophic lifestyle, and that heterotrophs evolved from autotrophs.
A 2024 study directly estimated the order in which amino acids were added into the genetic code from early protein domain sequences. A total of 969 protein domains were classified as present in LUCA, including 101 domain sequences that dated back to the even-older pre-LUCA communities. 88% of the protein domains annotated as LUCA or pre-LUCA were confirmed by Moody et al. 2024, by being associated with proteins that are more than 50% likely to be present in LUCA. It found that amino acids that bind metals, and those that contain sulphur, came early in the genetic code. The study suggests that sulphur metabolism and catalysis involving metals were important elements of life at the time of LUCA.
Possibly a mesophile
Several lines of evidence suggest that LUCA was non-thermophilic. The content of G + C nucleotide pairs (compared to the occurrence of A + T pairs) can indicate an organism's thermal optimum as they are more thermally stable due to an additional hydrogen bond. As a result, they occur more frequently in the rRNA of thermophiles; however, this is not seen in LUCA's reconstructed rRNA.
The identification of thermophilic genes in the LUCA has been challenged, as they may instead represent genes that evolved later in archaea or bacteria, then migrated between these via horizontal gene transfer, as in Woese's 1998 hypothesis. For instance, the thermophile-specific topoisomerase, reverse gyrase, was initially attributed to LUCA before an exhaustive phylogenetic study revealed a more recent origin of this enzyme followed by extensive horizontal gene transfer. LUCA could have been a mesophile that fixed CO2 and relied on H2, and lived close to hydrothermal vents.
Further evidence that LUCA was mesophilic comes from the amino acid composition of its proteins. The abundance of I, V, Y, W, R, E, and L amino acids (denoted IVYWREL) in an organism's proteins is correlated with its optimal growth temperature. According to phylogenetic analysis, the IVYWREL content of LUCA's proteins suggests its ideal temperature was below 50°C.
Evidence that bacteria and archaea both independently underwent phases of increased and subsequently decreased thermo-tolerance suggests a dramatic post-LUCA climate shift that affected both populations, and would explain the seeming genetic pervasiveness of thermo-tolerant genetics.
Age
Studies from 2000 to 2018 have suggested an increasingly ancient time for the LUCA. In 2000, estimates of the LUCA's age ranged from 3.8 to 3.5 bya (billion years ago) in the Paleoarchean, a few hundred million years after the earliest fossil evidence of life, for which candidates range in age from 4.28 to 3.48 bya. This placed the origin of the first forms of life shortly after the Late Heavy Bombardment which was thought to have repeatedly sterilized Earth's surface. However, a 2018 study by Holly Betts and colleagues applied a molecular clock model to the genomic and fossil record (102 species, 29 common protein-coding genes, mostly ribosomal), concluding that LUCA preceded the Late Heavy Bombardment (making the LUCA over 3.9 bya). A 2022 study suggested an age of around 4.2–3.6 bya for the LUCA. A 2024 study suggested that the LUCA lived around 4.2 bya (with a confidence interval of 4.33–4.09 bya). This 2024 dating implies that the time gap between the origin of life and LUCA is only about 100-200 million years, as the first habitable environments likely formed around 4.3 or 4.4 bya.
Root of the tree of life

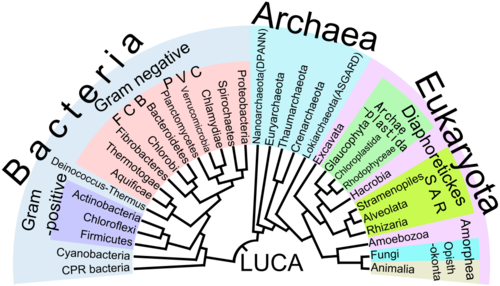
In 1990, a novel concept of the tree of life was presented, dividing the living world into three stems, classified as the domains Bacteria, Archaea, Eukarya. It is the first tree founded exclusively on molecular phylogenetics, and which includes the evolution of microorganisms. It has been called a "universal phylogenetic tree in rooted form". This tree and its rooting became the subject of debate.
In the meantime, numerous modifications of this tree, mainly concerning the role and importance of horizontal gene transfer for its rooting and early ramifications have been suggested. Since heredity occurs both vertically and horizontally, the tree of life may have been more weblike or netlike in its early phase and more treelike when it grew three-stemmed. Presumably horizontal gene transfer has decreased with growing cell stability.
A modified version of the tree, based on several molecular studies, has its root between a monophyletic domain Bacteria and a clade formed by Archaea and Eukaryota. A small minority of studies place the root in the domain bacteria, in the phylum Bacillota, or state that the phylum Chloroflexota (formerly Chloroflexi) is basal to a clade with Archaea and Eukaryotes and the rest of bacteria (as proposed by Thomas Cavalier-Smith). Metagenomic analyses recover a two-domain system with the domains Archaea and Bacteria; in this view of the tree of life, Eukaryotes are derived from Archaea. With the later gene pool of LUCA's descendants, sharing a common framework of the AT/GC rule and the standard twenty amino acids, horizontal gene transfer would have become feasible and could have been common.
The nature of LUCA remains disputed. In 1994, on the basis of primordial metabolism (as discussed by Wächtershäuser), Otto Kandler proposed a successive divergence of the three domains of life from a multiphenotypical population of pre-cells, reached by gradual evolutionary improvements (cellularization). The phenotypically diverse pre-cells of this population were metabolising, self-reproducing entities exhibiting frequent mutual exchange of genetic information. Thus, in this scenario there was no "first cell". It may explain the unity and, at the same time, the partition into three lines (the three domains) of life. Kandler's pre-cell theory is supported by Wächtershäuser. In 1998, Carl Woese, based on the RNA world concept, proposed that no individual organism could be considered a LUCA, and that the genetic heritage of all modern organisms derived through horizontal gene transfer among an ancient community of organisms. Other authors concur that there was a "complex collective genome" at the time of the LUCA, and that horizontal gene transfer was important in the evolution of later groups; Nicolas Glansdorff states that LUCA "was in a metabolically and morphologically heterogeneous community, constantly shuffling around genetic material" and "remained an evolutionary entity, though loosely defined and constantly changing, as long as this promiscuity lasted."
The theory of a universal common ancestry of life is widely accepted. In 2010, based on "the vast array of molecular sequences now available from all domains of life", D. L. Theobald published a "formal test" of universal common ancestry (UCA). This deals with the common descent of all extant terrestrial organisms, each being a genealogical descendant of a single species from the distant past. His formal test favoured the existence of a universal common ancestry over a wide class of alternative hypotheses that included horizontal gene transfer. Basic biochemical principles imply that all organisms do have a common ancestry.
A proposed non-cellular ancestor to LUCA is the first universal common ancestor (FUCA). FUCA would therefore be the ancestor to every modern cell as well as to ancient, now-extinct cellular lineages not descending from LUCA. FUCA is assumed to have had descendants other than LUCA, none of which have modern descendants. Some genes of these ancient now-extinct cell lineages are thought to have been horizontally transferred into the genome of early descendants of LUCA.
LUCA and viruses
The origin of viruses remains disputed. Since viruses need host cells for their replication, it is likely that they emerged after the formation of cells. Viruses may even have multiple origins and different types of viruses may have evolved independently over the history of life. There are different hypotheses for the origins of viruses, for instance an early viral origin from the RNA world or a later viral origin from selfish DNA.
Based on how viruses are currently distributed across the bacteria and archaea, the LUCA is suspected of having been prey to multiple viruses, ancestral to those that now have those two domains as their hosts. Furthermore, extensive virus evolution seems to have preceded the LUCA, since the jelly-roll structure of capsid proteins is shared by RNA and DNA viruses across all three domains of life. LUCA's viruses were probably mainly dsDNA viruses in the groups called Duplodnaviria and Varidnaviria. Two other single-stranded DNA virus groups within the Monodnaviria, the Microviridae and the Tubulavirales, likely infected the last bacterial common ancestor. The last archaeal common ancestor was probably host to spindle-shaped viruses. All of these could well have affected the LUCA, in which case each must since have been lost in the host domain where it is no longer extant. By contrast, RNA viruses do not appear to have been important parasites of LUCA, even though straightforward thinking might have envisaged viruses as beginning with RNA viruses directly derived from an RNA world. Instead, by the time the LUCA lived, RNA viruses had probably already been out-competed by DNA viruses.
LUCA might have been the ancestor of some viruses, with one of its descendants being the archaic virocell ancestor, the ancestor to large-to-medium-sized DNA viruses. The distribution pattern of DNA viruses in the tree of life also supports the hypothesis that DNA viruses originated from RNA-based LUCA. Viruses could equally have evolved before LUCA but after the first universal common ancestor (FUCA), according to the reduction hypothesis, where giant viruses evolved from primordial cells that became parasitic.









